Abstract
B cell-acute lymphoblastic leukemia with t(5;14)(q31;q32) is a rare malignancy accounting for less than 1% of cases of acute lymphoblastic leukemia. Patients often present with eosinophilia and normal or increased blast counts; thus, the diagnosis may be easily confused for an alternative entity if t(5;14)(q31;q32) is not identified. We present a case of B-ALL with t(5;14)(q31;q32) where the diagnosis was especially challenging given the patient’s multiple risk factors for eosinophilia and normal chromosome analysis and fluorescence in situ hybridization studies.The diagnosis was ultimately made with the identification of t(5;14)(q31;q32) by next generation sequencing. The patient received chemotherapy and is in clinical remission. This case adds to the limited body of literature of this rare entity and illustrates the value of next generation sequencing in the evaluation of patients with eosinophilia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
B cell-acute lymphoblastic leukemia (B-ALL) with t(5;14)(q31;q32); IGH::IL3 is a distinct entity in the World Health Organization classification of tumors of hematopoietic and lymphoid tissues [1]. This ALL subtype is rare, accounting for less than 1% of cases. No specific blast count is required for the diagnosis of this subtype, and patients may present with prominent eosinophilia and variable blast counts, making the detection of t(5;14) important to distinguishing this subtype from benign and malignant disorders of eosinophilia [2]. It is thus important that sensitive methods for the detection of this translocation be utilized so that a diagnosis of ALL with t(5;14) is not missed.
This report presents a case of B-ALL with t(5;14)(q31;q32) where the diagnosis was challenging given the patient’s multiple risk factors for eosinophilia as well as normal chromosome analysis and fluorescent in situ hybridization (FISH) studies. Next generation sequencing (NGS) was critical to making the correct diagnosis. This case adds to the limited literature on the pathology and clinical behavior of this entity and highlights the value of NGS in the diagnosis of disorders of eosinophilia.
Clinical history
A 48-year-old male hog farmer from rural Missouri who had immigrated from Central America approximately 20 years prior, and who had a past medical history of hypertension and hyperlipidemia, presented to the emergency department with 2–3 weeks of chest pain and 1–2 days of right upper extremity weakness. The patient’s EKG findings and an elevated serum troponin were suggestive of non-ST elevated myocardial infarction (NSTEMI). Cardiac catheterization showed patent coronary arteries. Magnetic resonance imaging of the brain demonstrated scattered bilateral cerebral and cerebellar subacute infarcts. His complete blood count showed leukocytosis (white blood cell count (WBC) 107.59 × 109/L) and thrombocytopenia (platelets 134 × 109/L), without anemia (hemoglobin 16.0 g/dl, hematocrit 45%). A WBC differential identified marked eosinophilia (eosinophils 70.01 × 109/L, 66%). The remainder of the differential count for the peripheral blood was segmented neutrophils 12%, bands 0%, metamyelocytes 4%, promyelocytes 1%, lymphocytes 10%, monocytes 4%, and basophils 3%. No blasts were identified in the peripheral blood. The patient denied history of allergies, medication usage, or recent travel. Infectious causes of eosinophilia were ruled out. Serum tryptase level was within the reference range. The patient was treated empirically with high dose steroids and ivermectin and showed a remarkable clinical response, including a drastic decline of his white blood cell count. Due to the lack of explanation for the eosinophilia seen at presentation, a bone marrow biopsy was performed.
Materials and methods
Flow cytometry
Flow cytometry was performed on the bone marrow aspirate at University Hospital (Columbia, MO). Data acquisition was completed using a BD FACSCanto II 8-color cytometer, utilizing BD FACSDiva Software v8.0.1 for analysis.
Chromosome analysis
Chromosome analysis was performed on the bone marrow aspirate at Mayo Clinic Laboratories (Rochester, MN). Cells were cultured without mitogens and underwent G-banding. A total of 20 metaphases were examined.
Fluorescence in situ hybridization
An ALL FISH panel was performed on the bone marrow aspirate at Mayo Clinic Laboratories (Rochester, MN). The panel included an IGH break-apart probe for identifying rearrangement of 14q32.
Next generation sequencing
Next generation sequencing (FoundationOne Heme panel) was performed on the bone marrow aspirate at Foundation One (Cambridge, MA). FoundationOne Heme includes DNA sequencing of 400 genes and RNA sequencing of 200 genes.
Results
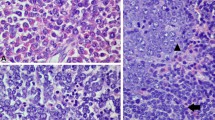
The bone marrow aspirate was hypercellular and contained morphologically abnormal eosinophils with vacuolated cytoplasm (Fig. 1a). Erythroid and myeloid cells were present in decreased numbers but showed normal maturation. Blasts were not increased by morphologic assessment of the aspirate (1.4%). The bone marrow biopsy demonstrated hypercellularity with increased numbers of eosinophils (Fig. 1b). Blasts were positive for CD34 and TdT, and had diffuse distribution and minimal clustering (Fig. 1c and d). Flow cytometry of the bone marrow aspirate demonstrated hypereosinophilia and 3.4% precursor B-cell lymphoblasts with expression of CD34, CD19, and aberrant bright CD10; moderate cTdT; and dim to moderate HLA-DR. The lymphoblasts were negative for CD45, CD13, CD33, CD11b, CD15, CD16, CD20, CD5, CD23, CD117, CD36, CD64, CD61, CD14, cMPO, cCD22, cCD3, and all related T-cell antigens (Fig. 2). The CSF was negative for blasts. Cytogenetic studies revealed a normal male karyotype. A fluorescence in situ hybridization (FISH) study, which included an IGH break-apart probe for 14q32, was negative. FISH probes for genes associated with eosinophilia such as PDGFRA and PGDFRB were not used, as it was expected that alterations in such genes would be identified by next generation sequencing (NGS). NGS utilizing the FoundationOne Heme panel detected the presence of an IGH::IL3 rearrangement (t(5;14)(q31;q32)). The patient was treated with hyper-CVAD. Bone marrow examination following 2 cycles of treatment was negative for leukemia. The patient has completed a total of 4 cycles of chemotherapy and remains clinically in remission.
Bone marrow aspirate with morphologically abnormal eosinophils containing vacuolated cytoplasm (a). Hematoxylin and eosin stain of bone marrow demonstrating hypercellularity with increased numbers of eosinophils (b). Blasts are positive for CD34 by immunohistochemical stain (c). Blasts were positive for TdT by immunohistochemical stain (d)
Discussion
B cell-acute lymphoblastic leukemia (B-ALL) with t(5;14)(q31;q32); IGH::IL3 is an exceedingly rare leukemic subtype. The eosinophilia often present at the time of diagnosis is due to an increase in IL3 production secondary to the IGH::IL3 translocation present in leukemic blasts [3, 4]. IL-3 is known to lead to the maturation, release, and activation of eosinophils [5]. Blast counts may be normal or increased, and no specific blast count is required for the diagnosis [1].
Patients frequently present with neurologic involvement, including transient ischemic attack (TIA), acute stroke, seizure, and decreased consciousness [2, 6]. In addition to the upper extremity weakness attributable to multiple infarcts, the patient in this case presented with chest pain, EKG changes, and troponin elevation with patent coronary arteries. It has been reported in the literature that acute eosinophilic myocarditis can mimic myocardial infarction [7]. No myocardial biopsy was performed in this patient; thus, it is unknown if the cardiac symptoms were actually due to eosinophilic myocarditis. The chest pain resolved with administration of steroids.
Given that patients with this subtype of ALL can initially present with eosinophilia and variable blast counts, the disease process may be incorrectly characterized as reactive eosinophilia, idiopathic eosinophilic syndrome, or other disorders of eosinophilia, and as a result the diagnosis may be delayed [8]. It is thus critical that the presence of the IGH::IL3 translocation be identified. Conventional cytogenetics may identify this translocation; however, the sensitivity is limited by the number of metaphases examined. Other case reports have demonstrated that chromosome analysis has failed to identify the presence of the translocation in cases with both normal and increased blast counts [2, 9,10,11]. FISH may identify the translocation by using an IGH break apart probe, but further characterization is required to identify the specific translocation [11]. FISH panels often do not include a probe specific to this translocation due to its rarity. As with conventional cytogenetics, the sensitivity of FISH is limited by the number of cells examined. In addition, the sensitivity of FISH is also limited by the positive signal threshold required to distinguish true positive from false positive signal.
In this case, chromosome analysis and a FISH panel which included an IGH break apart probe were normal, and only NGS was able to identify the translocation. This is comparable to other cases reported in the literature where NGS, but not chromosome analysis or FISH, was able to identify the translocation [10, 11]. Recognition of t(5;14)(q31;q32) may be particularly challenging because the translocation is only present in leukemic blasts and not eosinophils; thus, the eosinophilia may mask the presence of the translocation [3]. Future studies may explore if IL-3 RNA expression levels could provide insight into the presence of this translocation in cases with low blast counts or in the detection of minimal residual disease.
This report is unique in that the diagnosis was particularly challenging given that the patient had multiple risk factors for eosinophilia including occupation as a hog farmer (parasite risk), time spent in central America (parasite risk), and residence in Missouri (endemic fungal infection risk).
This case provides support for the use of NGS in the workup of eosinophilia. Depending on the panel used, NGS has the benefit of being able to identify rare translocations such as t(5;14)(q31;q32), as well as the more common alterations seen in other neoplasms with accompanying eosinophilia. By incorporating NGS into the standard workup of eosinophilia, we may discover that the incidence of B-ALL t(5;14)(q31;q32) is greater than realized.
References
Swerdlow SH, International Agency for Research on Cancer (2008) WHO classification of tumours of haematopoietic and lymphoid tissues, 4th edn. Internat. Agency for Research on Cancer, Lyon
Fournier B, Balducci E, Duployez N et al (2019) B-ALL With t(5;14)(q31;q32); IGH-IL3 rearrangement and eosinophilia: a comprehensive analysis of a peculiar IGH-rearranged B-ALL. Front Oncol 9:1374. https://doi.org/10.3389/fonc.2019.01374
Kobayashi K, Mizuta S, Yamane N et al (2019) Paraneoplastic hypereosinophilic syndrome associated with IL3-IgH positive acute lymphoblastic leukemia. Pediatr Blood Cancer 66:e27449. https://doi.org/10.1002/pbc.27449
Grimaldi JC, Meeker TC (1989) The t(5;14) chromosomal translocation in a case of acute lymphocytic leukemia joins the interleukin-3 gene to the immunoglobulin heavy chain gene. Blood 73:2081–2085
Esnault S, Kelly EA (2016) Essential mechanisms of differential activation of eosinophils by IL-3 compared to GM-CSF and IL-5. Crit Rev Immunol 36:429–444. https://doi.org/10.1615/CritRevImmunol.2017020172
Toboso DG, Campos CB (2017) Peripheral eosinophilia as the first manifestation of B-cell acute lymphoblastic leukemia with t(5;14)(q31;q32). Blood 130:380. https://doi.org/10.1182/blood-2016-12-754812
Thambidorai SK, Korlakunta HL, Arouni AJ et al (2009) Acute eosinophilic myocarditis mimicking myocardial infarction. Tex Heart Inst J 36:355–357
Bomken S, Haigh S, Bown N et al (2015) Cutaneous B-lymphoblastic lymphoma with IL3/IgH translocation presenting with hypereosinophilia and acute endocarditis. Pediatr Blood Cancer 62:1055–1057. https://doi.org/10.1002/pbc.25318
Derrieux C, Freynet N, Frayfer J et al (2018) A case of B-cell precursor acute lymphoblastic leukemia with IL3-IGH rearrangement revealed by thromboembolism and marked eosinophilia. Leuk Lymphoma 59:2489–2492. https://doi.org/10.1080/10428194.2018.1430796
Yu H, Wertheim G, Shankar S et al (2016) Marked eosinophilia masking B lymphoblastic leukemia. Am J Hematol 91:543–544. https://doi.org/10.1002/ajh.24266
Guenzel AJ, Smadbeck JB, Golden CL et al (2021) Clinical utility of next generation sequencing to detect IGH/IL3 rearrangements [t(5;14)(q31.1;q32.1)] in B-lymphoblastic leukemia/lymphoma. Ann Diagn Pathol 53:151761. https://doi.org/10.1016/j.anndiagpath.2021.151761
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
For this type of study, formal consent is not required. The work conformed to the ethical guidelines set forth by our institution and was acknowledged by the Institutional Review Board.
Informed consent
For this type of study, informed consent is not required.
Consent for publication
For this type of study, consent for publication is not required.
Competing interests
R.H has received research funding and honoraria from Roche and has been a speaker for the company.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chelsky, Z.L., Saleh, M., Laziuk, K. et al. The value of next generation sequencing in the diagnosis of a rare cause of eosinophilia: B cell-acute lymphoblastic leukemia with t(5;14)(q31;q32). J Hematopathol 15, 239–243 (2022). https://doi.org/10.1007/s12308-022-00521-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12308-022-00521-8