Abstract
To explore how molecules became signs I will ask: “What sort of process is necessary and sufficient to treat a molecule as a sign?” This requires focusing on the interpreting system and its interpretive competence. To avoid assuming any properties that need to be explained I develop what I consider to be a simplest possible molecular model system which only assumes known physics and chemistry but nevertheless exemplifies the interpretive properties of interest. Three progressively more complex variants of this model of interpretive competence are developed that roughly parallel an icon-index-symbol hierarchic scaffolding logic. The implication of this analysis is a reversal of the current dogma of molecular and evolutionary biology which treats molecules like DNA and RNA as the original sources of biological information. Instead I argue that the structural characteristics of these molecules have provided semiotic affordances that the interpretive dynamics of viruses and cells have taken advantage of. These molecules are not the source of biological information but are instead semiotic artifacts onto which dynamical functional constraints have been progressively offloaded during the course of evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
When Erwin Schrödinger (1944) pondered What is Life? from a physicist’s point of view he focused on two conundrums: how organisms maintain themselves in a far from equilibrium thermodynamic state and how they store and pass on the information that determines their organization. In his metaphor of an aperiodic crystal as the carrier of this information he both foreshadowed Claude Shannon’s (1948) analysis of information storage and transmission and Watson and Crick’s (1953) discovery of the double helix structure of the DNA molecule. So by 1958 when Francis Crick (1958) first articulated what he called the “central dogma” of molecular biology (i.e. that information in the cell flows from DNA to RNA to protein structure and not the reverse) it was taken for granted that that DNA and RNA molecules were “carriers” of information. By scientific rhetorical fiat it had become legitimate to treat molecules as able to provide information “about” other molecules. By the mid 1970s Richard Dawkins (1976) could safely assume this as fact and follow the idea to its logical implications for evolutionary theory in his popular book The Selfish Gene. By describing a sequence of nucleotides in a DNA molecule as information and DNA replication as the essential defining feature of life, information was reduced to pattern and interpretation was reduced to copying. What may have initially been a metaphor became difficult to disentangle from the chemistry.
In this way the concept of biological information lost its aboutness but became safe for use in a materialistic science that had no place for what seemed like a nonphysical property. This also made the concept of biological information consistent with the engineering conception of communication described by Claude Shannon (1948) in the introduction to his famous “Mathematical theory of Communication.” In the introductory paragraph he says that “The fundamental problem of communication is that of reproducing at one point either exactly or approximately a message selected at another point.”
Notice the near identity with Dawkins’ conception of replication. Both approaches only consider the properties of the communication medium itself and ignore all referential and functional properties. Shannon acknowledges this when he follows this by immediately pointing out that “Frequently the messages have meaning; that is they refer to or are correlated according to some system with certain physical or conceptual entities. These semantic aspects of communication are irrelevant to the engineering problem.” As the information theorist Robert Fano once remarked, when discussing Shannon’s theory:
“I didn’t like the term Information Theory. Claude didn’t like it either. You see, the term ‘information theory’ suggests that it is a theory about information – but it’s not. It’s the transmission of information, not information. Lots of people just didn’t understand this … information is always about something. It is information provided by something, about something.” Interview with R. Fano (2001)
But Dawkins makes no such distinction. Unlike Shannon’s “engineering problem,” however, the “biological problem” cannot be adequately addressed with out taking into account the function of molecular information. A physical pattern by itself is not about anything. The sequence of nucleotides in a DNA molecule is just a molecular structure considered outside the context of a living cell. For this structure to be about something there must be a process that interprets it. And not just any process will do.
So, is replication such a process?
The Centrality of Interpretation
Shannon’s analysis demonstrates that replication characterizes the communication or transmission of information, irrespective of any considerations of meaning or use. This is a conception of information in terms of intrinsic properties alone. But this use of the concept of information begs the question: In what sense are the intrinsic properties of a communication medium able to be about anything? This question has a semiotic counterpart: In what way do the properties of a sign vehicle determine its reference? Does similarity in form determine iconicity? Does regular correlation determine indexicality? Clearly this is too simple. Determination is not operative here, since there are unlimited classes of similarity and correlational relationships in the world. Though it is a common shorthand to treat portraits as icons and thermometers as indices, this has more to do with what they were created for and what a community assumes is their “proper” interpretation. But when an art critique recognizes the style of a particular portrait and infers from it who it was painted by, it is an index, and when a thermometer reminds someone of a drinking straw, it is an icon. This demonstrates that if we equate semiotic properties with sign vehicle properties or with the multitude of different uses that are possible, we are forced to say that portraits and thermometers are at the same time each both icons and indices.
This leads to a principle that will frame the remainder of this essay. Perhaps it could be (ironically) described as the central dogma of semiotics. It can be stated as follows:
Any property of a physical medium can serve as a sign vehicle of any type (icon, index, or symbol) referring to any object of reference for whatever function or purpose because these properties are generated by and entirely dependent upon the form of the particular interpretive process that it is incorporated into.
Thus, we should not ask what it is about some sign vehicle that makes it an icon index, or symbol. These are not sign vehicle intrinsic properties. Intrinsic properties are not what make something semiotic. Sign vehicle properties aren’t irrelevant, of course. But intrinsic properties are merely semiotic affordances (to borrow a concept from ecological psychology). They may or may not be utilized for any semiotic purpose. Often the semiotically relevant property of a sign vehicle is only one of its many attributes, and not necessarily the one most salient. What matters is how the relevant property is incorporated into an interpretive process, because being interpreted is what matters.
This does not mean that Shannon’s analysis of the mathematics of communication is irrelevant for biosemiotic analysis. Indeed, semiosis must be consistent with the constraints on communication, storage, and rectification that Shannon’s theory specifies. It’s just that semiotic properties involve something more: interpretation.
So when in the title of this essay I ask “How molecules became signs?” I am actually asking “What form of molecular process is necessary and sufficient to interpret some property of a molecule as providing information about other molecular properties?” In Peircean terms, this amounts to asking what sort of molecular system is competent to produce the interpretants that can bring this re-presented property into useful relation with that system?
In many respects, this question focuses on an attribute of semiosis that Peirce assiduously avoided: talk of interpreters. In his care to avoid the fallacy of psychologism—i.e. not falling into the trap of attributing semiotic processes to some unanalyzed homunculus—Peirce bracketed any description of how interpretation is physically implemented and instead focused on its logical structure.
In an age when neuroscience was in its early infancy and molecular biology was not even imaginable, it is not surprising that he avoided speculating about what sorts of dynamical systems were competent to be interpreters. Because of the vast complexity of brains and despite remarkable advances in neuroscience, it may still be premature to speculate about the neural implementation of mental semiosis. On the other hand, there are reasons to be more hopeful that insights into the physical implementation of interpretation might be obtained within molecular biology.
Origin of Information?
Ironically, I suggest that one of the most enigmatic unsolved mysteries in biology can provide the best place to look for insight into the physical implementation of interpretation. I am referring to the mystery of the origin of life. Why should this unlikely subject offer a privileged view of the issue? First, because it arose by accident, not design, the first life-forms almost certainly were constituted by quite simple molecular processes. Second, despite its simplicity, this molecular complex must have locally inverted one of the most ubiquitous regularities of the universe: the second law of thermodynamics. Though living functions act to compensate for this increase of entropy internally, organisms accomplish this by doing work that ultimately “exports” entropy to the environment at a rate higher than if they were just dissipating heat as they fell to equilibrium. So the origin of life problem brings together three seemingly incommensurate properties. It involves an extremely simple spontaneously produced molecule system that persists far from thermodynamic equilibrium (unlike almost all other chemical processes), and selectively interacts with its immediate environment in ways that support the persistence of these processes. This latter disposition is what demands a simple form of interpretive competence. To persist and even reproduce its unstable far from equilibrium condition this tiny first step toward life required an ability to re-presentFootnote 1 itself in ever new substrates ultimately borrowed from its environment. In other words, it was adapted to its environment.
It is precisely in this origins of life context that the eliminativist perspective on biological information is alive and well, and squarely in the mainstream. It is currently recapitulated in the dominant scenario for explaining the origins of Life: the RNA-World hypothesis. This approach was originally motivated by the discovery that RNA molecules could serve both as replication templates for copying its structure and as catalysts potentially able to facilitate this copying (though to date neither of these essential steps of the process have been demonstrated). The problem with a “naked replicator” approach, as Fano (above) recognized, is that replication isn’t about anything, nor does it contribute to anything except increasing numbers of similar objects. And although there can be something analogous to “selection” eliminating modified sequences that fail to replicate, the “external” environment does all the work. Replicating molecules are passive artifacts. They don’t actively adapt to their environment, and so their structure does not contain or acquire information about the environment and they do not have any intrinsic disposition to correct “errors” because the very concept of error has no intrinsic meaning. There just is what gets copied and what doesn’t, and whether something gets copied or not is only interpretable as success or failure from an external observer’s point of view. Nevertheless, the RNA-World hypothesis does have one thing going for it: its simplicity.
Some of the most significant advances in science have been based on the analysis of idealized simple model systems. A good model system captures the essential features of the problem without obscuring the critical assumptions in unanalyzed complexities. Examples include: Boltzmann’s molecule in a box, Maxwell’s demon, Bohr’s atom, Turing’s machine. A good model should include no unknown or undescribed processes, insure that all operations are physically realistic, include no opaque (black box) properties, and provide unambiguous exemplification of the properties of interest. It is precisely because of its simplicity that the weaknesses of the naked replicator approach are easily recognized.
A different Simple Model System
To investigate how a physical process could come to treat a molecule as information about something else I will employ a different but equally simple model system, but one that makes fundamentally different assumptions about the nature of information than do replicator models.
The model I will use for this purpose is a hypothetical but physically realizable minimally complex molecular process. I first introduced this sort of molecular model in a 2006 paper and have modified it slightly in the years since to ensure that it is both empirically realizable and adequate to its explanatory purpose.
It is modeled after virus structure. In this respect it is not an idealization, just an as yet physically unrealized chemical system. It can be described as a non-parasitic virus that can reproduce autonomously. In this regard it is an autogenic virus, able to autonomously generate copies of itself. A simple virus, like the polio virus, consists of a container or “capsid” shell typically made of protein molecules that assemble themselves into facets of a polyhedral structure that encloses an RNA or DNA molecule. When incorporated into a host cell the viral RNA or DNA commandeers the cell’s systems to make more capsid molecules and more copies of the viral RNA or DNA. Since viral replication requires these complex protein synthesis and polynucleotide synthesis processes, and the molecular machinery to do this involves dozens of molecules arranged in complex structures, viruses replicate parasitically. So a non-parasitic virus would need to use a different and much simpler molecular process to reproduce its parts.
One candidate process is reciprocal catalysis. The simplest form of reciprocal catalysis occurs when one catalytic reaction produces a product that catalyzes a second reaction which produces a product that catalyzes the first, and so on. When provided with appropriate substrate molecules this circular network of catalytic reactions becomes a chain reaction that can rapidly produce large numbers of catalyst molecules. Reciprocal catalysis can involve multiple steps, so long as the circle of reactions is closed, though as we’ll see below, increasing the numbers of interacting molecules is problematic.
Viral capsids self-assemble (as do cell membranes, microtubules, and many other complex molecular structures within cells). Self-assembly is essentially a variant of the process of crystalization. Because of the way that the regular geometries and affinities of these molecules cause them to associate with one another they can spontaneously form into sheets, polyhedrons, or tubes.
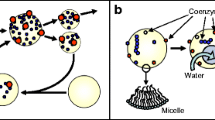
These two processes—reciprocal catalysis and self-assembly (depicted in Fig. 1)—are chemically complementary to one another because they each tend to produce conditions that are necessary for the other to occur. So reciprocal catalysis produces high locally asymmetric concentrations of a small number of molecular species while self-assembly requires persistently high local concentrations of a single species of component molecules. Likewise, self-assembly produces constraint on molecular diffusion while reciprocal catalysis requires limited diffusion of interdependent catalysts in order to occur. In this way reciprocal catalysis and self-assembly are molecular processes that each produce the boundary conditions that are critical for supporting each other.
Two molecular processes that are widely represented in essentially all life forms: reciprocal catalysis (left) and self-assembly (right). The example of self-assembly depicts the construction of a polyhedral viral capsid (shell) in which a complex of 3 proteins (1, 2, 3) form a symmetric facet of an icosahedral capsid
These process can become coupled and their reciprocal relationships linked if one of the molecular side products generated in a reciprocal catalytic process tends to self-assemble into a closed structure. In this case capsid formation will tend occur most effectively where reciprocal catalysis occurs. But this increases the probability that capsids will tend to grow to enclose a sample of the reciprocal catalysts that both produce one another and capsid-forming molecules.
As a result, catalysts that reciprocally depend on one another to be produced will tend to be co-localized, and prevented from diffusing away from one another. While contained, catalysis will quickly cease when substrates are used up, but in the case that the capsid is subsequently damaged and spills its contents, more catalysts and capsid molecules will be synthesized if there are additional substrate molecules nearby. So damage that causes an otherwise inert capsid to spill its catalytic contents into an environment with available substrates will initiate a process that effectively repairs the damage and reconstitutes its inert form. Moreover, depending on the extent of the damage, the distribution of catalytic contents, and the concentrations of substrate molecules the process could potentially produce a second copy of the original from the excess catalyst and capsid molecules that are generated. This makes possible self-repair and even self-reproduction. I will call such an autogenic virus an “autogen” for short (two variants of autogens along with a reaction diagram are shown in Fig. 2).

modified by catalyst f to produce catalyst c plus side product n and catalyst c modifies substrate molecule d to produce catalyst f and side product g which tends to self assemble onto a capsid and will thereby tend to encapsulate catalysts c and f and prevent them from diffusing away from one another
Depictions of two hypothetical forms of simple autogenetic (i.e. self-reproducing) viruses with polyhedral (left panel) and tubular (middle panel) capsid structure. The chemical logic of simple autogenesis is depicted in the diagram in the right panel. Molecule a is
This constitutes what can be described as an autogenic work cycle. A work cycle consists of a linked sequence of thermodynamic processes that involve transfer of work into and out of a system … and that eventually returns the system to its initial state (paraphrased from Wikipedia). A familiar example is provided by a motor. It is designed to operate continuously when supplied with a constant or periodic throughput of work that changes its configuration through a series of states until the system returns to its initial state. In this way it is able to repeat this cycle again and again. For example, an internal combustion engine uses exploding gasses to move it through a series of configurations so that eventually it expels the exploded gasses and is ready for new fuel and air to be taken in and exploded. The power or (endergonic = “ingoing” + “work”) phase and the exhaust and relaxation (exergonic = “outgoing” + ”work”) phase are matched so that energy doesn’t continually build up within the system.
An autogenic work cycle is similarly composed of two phases distinguished by their difference in chemistry and thermodynamic directionality (see Fig. 3). Catalysis lowers the threshold that must be exceeded in order to initiate a chemical process but once this threshold is crossed an energy gradient difference from reactant to product drives the reaction. Thus the process is endergonic. In contrast, self-assembly (and crystallization in general) enables molecules in a higher energy state in solution to precipitate out of solution into a lattice that absorbs and dissipates this kinetic energy (i.e. of motion, rotation, and vibration) and so spontaneously proceeds from a higher to lower energy state. Thus the process is exergonic.
Diagrams depicting the steps of the autogenic work cycle (left panel) highlighting the reciprocity of the codependent self-organizing dynamics (reciprocal catalysis and self-assembling capsid) and a simplification that highlights the way that each of these processes provides the critical boundary constraints that supports the other (right panel)
So, analogous to the two phase cyclic dynamics of an internal combustion engine, the energy that drives the autogenic cycle is provided by energy released by catalysis. This energy—liberated from chemical bonds of the substrate molecules—is the source of work that produces additional catalysts as well as capsid molecules. Self-assembly in turn accumulates the capsid molecules thereby produced and in the process dissipates this energy in the form of heat and an increase in surrounding entropy. But unlike an engine in which the work produced by its changes of state is directed externally to alter some extrinsic state of things, the autogenic work is directed inward, so to speak, to regenerate the very conditions that drive these changes.
This produces a higher order work cycle in which the entire molecular system cycles from disrupted to reconstructed, dynamic to inert, and open to closed. When returned to the reconstructed inert phase the system has been returned to an initial state from which the cycle can again be repeated. At this point the work of self-reconstruction has produced a far-from-equilibrium structure with a relatively high threshold required to dissipate it (in the form of capsid damage). And yet when loss of integrity due to extrinsic damage is sufficient to initiate change toward equilibrium the re-initiation of catalysis and self-assembly works against this.
It is in this way that each of these self-organizing processes produces the extrinsic boundary conditions that the other requires. As a result the critical boundary conditions are internalized and constantly available to channel the work necessary to maintain and reproduce these same constraints. The two self-organizing dynamics are in this sense co-dependent. Each is in effect the permissive environment for the other and in this sense each “contains” the other. This creates an intrinsic source of causal dispositions so that external influences and fixed properties no longer determine its behavior. An autogen is therefore self-individuated by this intrinsic co-dependent dynamical disposition, irrespective of whether it is enclosed or partially dispersed.
Autogens are not only able to self-repair, but because of their cycling from open to closed organization they will also tend to acquire and exchange molecules with their environment. Captured molecules that incidentally share catalytic inter-reactivity with autogen catalysts or capsid molecules will tend to be incorporated and replicated. This will create variant autogen lineages. Those captured molecules that don’t interact with autogen-intrinsic molecules or impede the process without being lethal will tend to get crowded out and eventually passively expelled into the environment in successive reproductions because they are not replicated. This provides a capacity to correct error and to evolve.
So autogenesis provides what amounts to a constraint production and preservation ratchet. During the dynamical phase new components are produced but because of their co-dependent relationships to one another the constraints that provide the reciprocal boundary conditions are also produced as the probability of occurrence of the component self-organizing processes increases. Together these reciprocal and recursive relationships would make autogenic viruses minimally evolvable.
Constraint, Work, and Information
This exemplifies an important inter-dependency between constraint, work, and information that Stuart Kauffman and colleagues (2008) described in a paper titled “Propagating Organization: An Enquiry.” They point out that “… it takes constraints on the release of energy for work to happen, but work for the constraints themselves to come into existence.” In autogenic terms, the co-localized system of constraints that is preserved passively in the inert phase is regenerated and re-co-localized in the dynamic phase. And co-localization itself is one of the critical constraints that is preserved and replicated.
This circular relationship between constraint and work is exemplified in autogenic self-propagation and self-repair (see Fig. 4). The analogy to viral genetics shows why information is based on constraint. Both the reciprocity of chemical boundary constraints and the constraint on their linkage due to sharing a common molecule are preserved from one cycle to the next despite complete substrate replacement. This preservation of constraints both provides a record and a source of instruction for organizing the work required to preserve this same capacity. The critical property of constraint that makes this possible is its substrate transferability. This enables constraints to preserve a trace of past instantiations and past work—i.e. reference—demonstrating that these constraints are analogous to the genetic information of a virus.
This diagram depicts two cycles of damage and self-repair in which integrity is temporarily lost but the intrinsic constraints distributed in co-localized molecules enables the recruitment of energy and new substrates from the environment to reconstitute autogenic integrity. As a result the constraints embodied in the first inert autogen are maintained throughout. They remain continuously present despite old molecular substrates being replaced by newly synthesized ones. In this way information in the form of these constraints is inherited by future materially independent replicas and “instructs” their formation
To summarize the argument so far: there are 5 holistic properties that even a simple autogenic system exhibits that are not reducible to the physical–chemical properties of its components and are emergent from the intrinsic dispositions of the whole integrated system. They are 1. individuation (it intrinsically maintains an unambiguous self/non-self distinction); 2. autonomy (it intrinsically embodies and maintains its own boundary conditions via component processes that reciprocally produce the external boundary conditions for each other); 3. recursive self-maintenanceFootnote 2 (it repairs and replicates the critical boundary conditions that are required to repair and replicate these same critical boundary conditions); 4. normativity (it is disposed to produce these results but can fail); and 5. interpretive competence (by being able to re-present its own boundary conditions in new instantiations it intrinsically re-presents and reproduces its own conditions of existence).
How can we characterize this most basic and simple interpretive competence in semiotic terms? The point of this model system is to establish what can be considered the ground of interpretive competence. In this respect it is effectively a “zeroth” level semiotic process. As such it “interprets” the most basic semiotic distinction; i.e. between self and non self. Thus disruption of integrity is a sign of non self and the dynamics that ensues and reconstitutes the stable state is the generation of an interpretant which actively reconstructs this self / non self distinction. So a cycle of autogenic disruption and self-repair treats every form of disruption as indistinguishable from each other—i.e. as iconic—because the system can only produce one form of interpretant. In this respect, iconism is the most basic semiotic operation because it marks the limit of what can be interpretively distinguished.
As G. Spencer Brown (1969) puts it: “That which cannot be distinguished must be confused.” Or as Abraham Maslow’s (1966) famous aphorism suggests: “If the only tool you have is a hammer, you will treat everything like a nail.”
All semiosis must therefore originate from and terminate with iconism in this most generic sense. It marks the point where no more developed interpretant can be generated. Importantly, this treats iconism not as a feature of a sign vehicle but rather as a function of interpretive in-distinction. This again reiterates what in the introduction I proposed as the central dogma of semiotics: that semiotic properties are not identified with sign vehicle properties but rather with how these properties provide affordances for an agent’s interpretive competence. This shift of emphasis becomes especially important for biosemiotic analysis because it helps to disambiguate the use of the icon-index-symbol terminology, originally derived from phenomenological reflection, from analogous uses at many levels of the biological hierarchy. In what follows, I will therefore focus on the way that different interpretive processes make use of different affordances provided by the available semiotic media (in this case molecular properties).
An Autogenic Analogue to Indexical Interpretation
From this most basic form of self-re-presentation two canonical complications of interpretive dynamics can be derived. An additional capacity beyond self-interpretation involves the ability to interpret different environmental conditions with respect to their relevance to the recursive self-maintenance of the interpreting system. This can be provided by incorporating a subordinate similarity plus correlation-dependent disposition into the basic autogenic process.
This slightly more complex interpretive capacity is exemplified by an autogen (see Fig. 5) that is selectively sensitive to its environment because 1. the capsid surface has structures (epitopes) onto which potential substrate molecules will tend to bind, and 2. capsid integrity is made increasingly fragile as the number of surface-bound substrates increases. This will make containment more likely to fail and release catalysts in reproductively supportive conditions. Moreover, the threshold level at which capsid integrity becomes unstable is a variable that is subject to a form of natural selection. So, over time, autogenic lineages more likely to break open when the concentration of external substrates is optimal for successful reconstruction and reproduction will tend to replace those whose sensitivity is less well correlated with successful self-reconstitution.
This slightly more complicated autogenic system that is capable of assaying its environment is depicted on the left (as a tubular autogen) and its reaction network structure is diagrammed on the right. This complication of the basic autogenic model system involves 1. a capsid surface with protruding features (epitopes) shaped in such a way that substrate molecules (indicated by molecules of type d in the diagram) will tend to bind to them; and 2. capsid molecule cohesion that is weakened in proportion to the number of surface-bound substrates, to the point that containment can be more easily be disrupted when surface binding is extensive. Notice that substrate binding is mediated by a similarity (isometry) relation and containment is weakened in correlation with the concentration of substrates in the environment (i.e. with respect to their physical contiguity and quantity)
Although conceived to apply to radically different domains of semiosis (i.e. molecular and mental), I think that a correspondence can be discerned between the phases of this molecular interpretive process and Peirce’s ten part taxonomy of semiotic relationships which he developed in the period from 1904 to 1909.Footnote 3 Thus, I would describe the sign vehicle (representamen) as the change in fragility of the capsid that causes it to rupture; the immediate interpretant as the disposition to change from inert to dynamical state that the sign initiates; the dynamical interpretant as the work that accomplishes autogenic reconstruction; the final interpretant as the system’s total disposition (or habit) to initiate self-regeneration in response to these conditions; the immediate object as the potential suitability of the environment with respect to this habit that this process signifies; and the dynamical object as the actual physical state of the environment.
Though the exegetical legitimacy of this comparison is irrelevant to the explanatory adequacy of this molecular process, the parallels suggest that similar principles may apply across very different levels of semiotic processes. So to develop the analogy further, in semiotic terms, the number of substrates bound to the autogenic capsid effectively indicates the presence or absence of extrinsic supportive conditions for persistence and reproduction of this same interpretive capacity. In this respect the interpretive process provides normative information about the environment that can potentially benefit the perpetuation of this same interpretive capacity.
This analogy is instructive in another sense. It demonstrates that the competence to interpret immediate conditions to be about correlated conditions is dependent on the more basic interpretive competence to re-present self. It is the self-correcting, self re-presenting capacity of simple autogenesis that enables the correlation between changes in capsid fragility to be about the value of the environment for that self and its interpretive capacity. To put this in semiotic terms, it suggests that indexical interpretive competence (grounded on correlational affordance) depends on more basic iconic interpretive competence (grounded on isomorphic affordance). As we will see below, this pattern of nested dependency in which different levels of semiosis are hierarchically constructed can be recursively iterated level upon level.
An Energetic Interlude
So far the account of the origins of biological information that I have presented does not involve either DNA or RNA. Instead, it has demonstrated that the constraints constituting a recursively self-maintaining molecular system provide the mnemonic, instructional, and normative attributes that we identify with biological information. But, as the title promises, it is the purpose of this model system approach to go one step further; to eventually explain how a molecule like DNA could come to be used as a source of information about the relationships among other molecules.
In order to accomplish this I will offer a somewhat more speculative scenario, that invokes a bit of currently uncharacterized chemistry (though it is also a critical missing step in the RNA-World and all other nucleic acid based scenarios). A hint as to why nucleic acids have become the primary carriers of information in living systems was originally suggested by the physicist Freeman Dyson (1985) in his book Origins of Life. Dyson suggested that, given their complexity, it is unlikely that nucleic acids could have developed fully blown prior to other metabolic processes. Instead he proposed a two stage process in which the building blocks of RNA originally served an energetic function and only later became repurposed (exapted) for information-bearing, information-preserving, and information-replicating. He based this on the fact that some nucleotides can serve dual roles. Besides being the building blocks of RNA and DNA molecules, nucleotides are also some of the principle molecules for acquiring, storing, and transporting chemical energy within a cell. This prompts the question “Why this curious chemical coincidence?".
Applying Dyson’s insight to the autogenic approach sketched above, this dual functional logic suggests a two phase autogenic evolutionary scenario for the origin of RNA.
Consider the following enhancement of simple autogenesis. If another of the side products produced by autogenic reciprocal catalysis is a molecule like the nucleotides ATP and GDP that can acquire and give up energy carried in pyrophosphate bonds, the availability of this generic free energy could potentially facilitate more effective catalysis and drive otherwise energetically unfavorable reactions. This could provide a sort of energy-assisted autogenesis which would tend to out-perform spontaneous autogenesis and be favored by natural selection. This could also enable a wider variety of potential substrate molecules to be useful, because the energy to drive reciprocal catalysis would not need to be derived from substrate lysis. The logic of this hypothetical energy-assisted autogenesis is diagramed in Fig. 6.
An energy-assisted autogenic reaction network. The diagram shows a side-product (n) of one of the catalytic steps that is a nucleotide-like molecule that is able to capture energy in the form of phosphate molecules (p) and transfer the energy to facilitate other catalytic reactions. On the right side of the diagram nucleotides stripped of their high-energy phosphates are shown linked into a polymer
But the availability of high-energy molecules is only useful during dynamic endergonic processes and can be disruptive of exergonic reactions and stable molecular structures. So energetic phosphates could cause potential damage during the inert phase of autogenesis. To be preserved safely and intact so they can be available when again catalysis is required they need to be somehow stored in an nonreactive form.
Nonreactive nucleotide-based molecules are of course well-known. They are DNA and RNA molecules. In these nucleotide polymers the phosphate residues serve as the links between adjacent sugars and so are nonreactive. By linking them into a polymer with phosphates unexposed, they can be effectively “stored” for later use via depolymerization. In this evolutionary scenario, then, the initial function of polynucleotide molecules is presumed to be energetic, and only later in evolution do they become recruited for their informational functions.
From Storage to Template to Information
The capacity to transfer constraints from one physical medium to another quite different one makes possible the transfer of the holistically embodied dynamical constraints of autogenesis onto a different sort of material substrate such as a nucleotide polymer.
This provides a means to overcome a critical limitation on the evolvability of autogenic interpretation. This limitation arises due to the threat of combinatorial catastrophe. A molecular combinatorial catastrophe can arise for autogenesis when the number of interdependent molecular interactions required to produce successful autogenic repair or reproduction increases. As the number of molecular species that need to interact increases linearly, the number of possible cross-reactions that could occur between members of the set increases geometrically. This is a problem because only a small fraction of these interactions will be supportive of autogenesis. The proliferation of alternative interaction possibilities will therefore compete with supportive interactions—using up critical components and wasting free energy. This will decrease efficiency and impede reproduction. So autogenic systems like the ones described above have limited evolvability, making autogenic evolution improbable beyond very simple forms. So unless non-supportive reactions can be selectively suppressed, autogenesis cannot lead to more complex forms of life.
But looked at from the perspective of living organisms, the suppression of all but a tiny fraction of possible chemical reactions is one way to view the function of the template molecules of life, the nucleic acids RNA and DNA and their roles in orchestrating cellular chemistry. In simple terms nucleic acids limit the kinds of proteins that are present in the cell, which in turn strongly biases the types of chemical reactions that tend to take place. Death of the cell or organism allows the myriad of previously suppressed chemical reactions to be re-expressed. So, although we generally tend to conceive of DNA-based synthesis of proteins as a generative process, it can also be considered to be the principle constraining influence that keeps deleterious reactions at bay.
To develop this scenario to show how these polynucleotide molecules could evolve to serve semiotic as well as energetic functions it is necessary to recognize that all five of the major nucleotide molecules (adenine, thymine, guanine, cytosine, and uracil) are capable of carrying and transferring phosphates. Among other related molecules, they could each have played slightly different energy transfer roles in early autogenic evolution due to their different purine or pyrimidine correlated nitrogenous bases, which affects the bonding affinities of these nucleotides. The phosphates are, however, attached to the opposite end of the nucleotide (to the ribose sugar) and so this difference in base minimally affects phosphate interactions. This results in the lack of any preferred phosphate bonding affinity between nucleotides during polymerization (a critical property for their informational role in living cells). As a result, diverse nucleotides will tend to form polymers of random order. And yet, although the sequence pattern of nucleotides is arbitrary, the specific nucleotide sequence produces a slightly different three dimensional conformation of the polymer at that location.
But as a relatively inert linear molecule, the structural properties of nucleotide polymers make them ideal to serve as templates. This is because conformation differences along the length of the molecule caused by the local nucleotide sequence provides a heterogeneous linear surface onto which other molecules can weakly bind. These structural differences will determine corresponding differences in how other molecules will tend to attach to the polymer due to their shape and charge complementarities. Since there will be both catalysts and polynucleotides within the inert autogen capsid, free catalysts will tend to associate with free nucleotide polymers with respect to these structural complementarities. The attached catalysts will therefore tend to be arranged into distinct sequences along the length of an extended nucleotide.
The spatial correlation relationships between catalysts aligned along a nucleic acid polymer will thereby tend to constrain the probability of particular catalytic interactions, increasing some and suppressing others. In this way the structural constraints of the template molecule can bias and constrain the interaction probabilities of the catalysts (see Fig. 7).
Three steps demonstrating how a nucleic acid template can regulate the reaction constraints of a complex autogenic process. The left panel depicts the weak binding of a protein (catalyst molecule) to a section of the nucleic acid double helix by virtue of their complementary shapes. The central panel abstractly depicts the binding of three catalysts on a nucleic acid template so that their tendency to interact with one another is determined by their relative proximity (blue = low probability, red = high probability). The right panel depicts the autogenically useful interactions as solid lines and the suppressed potentially disadvantageous interactions as dotted lines
This can lead to sequence-specific selection, since the order of nucleotides can affect the probabilities of catalyst interactions. Sequences that constrain catalyst interaction probabilities closer to the optimal interaction network will be selectively retained because of higher reproduction and repair rates, and the nucleotide sequences that correspond to this will be more likely preserved and replicated. In this way the template molecule can, in effect, offload some fraction of system dynamical constraints onto a structure that is not directly incorporated into or modified by the dynamics.
Whereas previous to the availability of the template molecule interaction constraints were entirely the result of specific chemical affinities with respect to one another, the availability of analogous biases provided by a template renders the intrinsic interaction affinities of the catalyst redundant and dispensable. Spontaneous degradation of these intrinsic interaction constraints can thus take place without loss of specificity. The result is that dynamical constraints previously provided by chemical interaction probabilities are transferred to the structure of an individual molecule. They are displaced from one substrate property and onto a very different substrate and its properties.
Because it is supported by template structure and not by any catalyst-intrinsic interaction tendencies this shifts the source of interaction constraints from catalyst properties to template properties. Since the template is not transformed by chemical reactions it can serve as a more stable source of memory and instruction allowing catalysts to be replaced by other kinds of molecules with chemical properties that might have superior catalytic capacity irrespective of their interaction specificity. The template is also subject to quite different chemical and physical influences than is the rest of the system. But the informational codependence between template and dynamics means that template modifications will have consequences for the dynamical organization of the whole system. Thus continuity of constraint across the change in molecular substrate can bring otherwise dynamically unrelated and independent physical–chemical properties into interaction with one another in ways that exploit their possible synergies.
Translating this into Peircean terminology again, two levels of semiosis can be distinguished in a system relying on a template—one offloaded from and nested within the other. First there is template interpretation, in which the template pattern can be considered the representamen (sign vehicle). The order of binding of the catalysts on the template can be considered an immediate interpretant. Their subsequently constrained interaction pattern can be considered a dynamical interpretant. And the habit that links these into a synergistic system can be considered a final interpretant.
Referential Displacement
In semiotic terms we could describe the result as creating a code-like relationship (though distinct from the so-called genetic code). It is code-like because it is based on a component-to-component mapping between the elements in two sets of otherwise unrelated substrates. In comparison, Marcello Barbieri (2015) attributes the code-like nature of the relationship of DNA sequences to amino acid sequences (the genetic code) to what he describes as “adaptors;” molecules that provide a physical link between distinct paired types of molecules. The paradigm example of a molecular adaptor is tRNA which physically links a particular amino acid to a distinct three nucleotide anticodon.
Although the genetic translation process is far more complex than the template-assisted autogenesis described here, there is an underlying abstract similarity in the way that the code-like (“arbitrary”) mapping in both is dependent on a particular combination of isometry and correlational relationships. In living cells distinct tRNA molecules become aligned with respect to mRNA template structure by virtue of codon-anticodon matching; i.e. isomorphism. And the correlation between a specific tRNA anticodon and the amino acid that is attached to that tRNA molecule enables correlational relationships between adjacent mRNA sequences to constrain corresponding correlational relationships between the amino acids constituting a protein. Analogously, the physical linkage between template and catalyst in template assisted autogenesis is also due to isomorphic similarity. This determines that structural correlations of template structure are additionally correlated with catalyst interaction constraints.
The substrate transferability of constraints thereby fractionates the previously holistic system of dynamical constraints, displacing some onto a comparatively inert substrate. As a result the structure of the molecular template literally re-presents the topology of the dynamical network of interactions that functions to re-present and re-produce itself. The result is what might be described as recursive self-representation; i.e. self-representation of self-representation. The circularity implied by this description refers to the way a part of a system is able to re-present the critical constraints of the whole system of which it is a part.
The template serves as both a record and a means to instruct the dynamics that reproduces the whole. This segregation of dynamical constraints and material structural constraints was originally described by the system theorist Howard Pattee as early as 1968 (and further developed in Pattee, 1969, 2001, 2006, and many others). It was held up as the defining property of living processes.Footnote 4 Offloading interaction constraints onto a static structure enables that structure to reliably re-present and preserve those critical constraints irrespective of any potentially degrading effects of dynamical interactions. This is because the correlated structural and dynamical constraints are embodied in otherwise unrelated physical properties linked only due to the functioning of the whole.
In summary: these variations on the autogenic model system exemplify a three tiered interpretive logic by which referential and instructional information can be derived and evolved. First there is simple autogenesis which is entirely determined by holistically embodied isomorphic (similarity) constraints distributed in its many components that preserve their own codependence despite damage and substrate replacement. Second there is context sensitive autogenesis which is determined by an augmentation of simple autogenesis in which the capsid surface presents structures with forms that are similar to the forms of useful substrates facilitating their binding to the surface where binding weakens capsid integrity. And third there is template-mediated autogenesis in which catalyst interaction constraints become offloaded onto a molecular structure. Offloading is afforded because complementary structural similarities between catalysts and regions of the template molecule facilitate catalyst binding in a particular order that by virtue of their positional correlations biases their interaction probabilities. In this way modifications of the structure of the template molecules can indirectly suppress potentially non beneficial interactions in favor of those that are conducive to autogenic repair and reproduction. The offloading of interaction constraints onto a physically separate and distinct structure preserves referential continuity while linking it to unrelated sign vehicle properties that can be harnessed for distinct semiotic functions, including semiotic recursion.
As noted above, it takes constraint on dynamics to perform physical work, but it takes physical work to produce new constraints. So separation of the source of constraint from the dynamics enables the dynamical interpretive process to re-interpret itself iteratively over time; i.e. to be recursive. This is the key to open-ended evolvability.
The Structure of Biosemiotic Scaffolding
This three-tiered structure of interpretive processes is general. Thus diplaced affordance (in which the information-bearing medium is segregated from the constrained dynamical medium) is made possible by the way that coupled isomorphic (similarity) and correlative (contiguous) affordances can mediate the displacement of constraints from one physical substrate to another. This provides a bridge that maintains continuity of information despite discontinuity of substrate. Since this change in substrate provides new isomorphic and correlational affordances, interpretive processes that take advantage of these properties simultaneously reinterpret the lower order interpretive processes. This enables what can be described as interpretive recursion, making it possible to evolve level upon level of interpretive complexity.
Semiotic scaffolding logic was introduced by Hoffmeyer (2007) and further developed in Hoffmeyer (2014a, 2014b, 2015) and subsequently explored by many other biosemioticians.Footnote 5 Semiotic scaffolding is well exemplified by the regulatory logic of molecular genetics. As discussed above, the genetic “code” enables the transfer of constraint from one kind of molecular substrate to another. Thus the sequence properties of DNA molecules inform the three dimensional interaction properties of proteins via the mediation of isomorphic and correlative relations between DNA and RNA molecules. In this way continuity of reference is maintained despite change in sign vehicle (molecular substrate). The displacement of constraint from one semiotic medium to another quite different one is what enables the scaffolding by which simple molecular semiosis can be recursively iterated level upon level. As discussed above, the local nucleotide sequence of a DNA molecule affects the twist of the DNA double helix at that locus. This facilitates selective binding of proteins able to alter adjacent gene expression. In this way protein structure specified by DNA sequences can act to promote, inhibit, or regulate the transcription of many other DNA sequences and the protein structures they determine (see Fig. 8). Thus, displacement of functional constraints onto a different molecular medium (i.e. from nucleotide sequence to protein structure) opens the door to recursive information dynamics.
A diagram of the “genetic code” depicting how constraint in the form of a nucleotide sequence is re-presented in the order of amino acids constituting a synthesized protein and how a protein can bind to the DNA template in a position determined by a specific nucleotide sequence at that position. The information continuity between DNA and protein is mediated by a combination of similarity and correlation (contiguity) relationships that preserves constraint (via iconism between nucleotide sequence and amino acid sequence) while transferring it to an otherwise unrelated molecular form. The material difference in sign vehicle makes possible recursive control of gene expression by virtue of the structural match between the 3 dimensional structure of a protein and the 3 dimensional difference in the local “twist” of the DNA molecule. Thus sequence constraints are transferred to 3 dimensional structural constraints which regulate the expression of sequence constraints, and so forth
By enabling recursive regulation of large suites of genes from a single locus, this regulatory logic provides the ground for semiotic scaffolding and the emergence of progressively higher levels of interpretive competence. The coordinated expression of large suites of genes can have large-scale phenotypic effects, both due to cell-intrinsic regulation and regulation of gene expression by whole suites of other cells. Thus semiotic constraint is progressively transferred from molecules to cells to tissues to body structure. With each higher level of displacement to a new level of substrate, a higher order form of recursion emerges. This is enhanced by the effect of gene duplication. In particular, the duplication and degeneration of regulatory genes creates the possibility of higher order displacement and interpretive affordance by virtue of similarity of gene expression despite differences in substrate correlations. For example the evolutionary duplication and variation of homeobox genes has been critical for determining the homologous anterior–posterior segmental morphologies of animal bodies and a similar family of genes is responsible for the theme and variation morphology of flowering plants (see Fig. 9).
The recursive relationship between gene sequence and protein structure that makes recursive gene regulation possible is the basis for higher order epigenetic recursive processes to evolve. Regulatory genes can thus influence the expression of whole suites of other genes by virtue of isomorphic binding sites (top left). Duplicated variants of regulatory genes which have slightly variant binding specificity can produce differences in the timing and location of expression, as well as differences in which genes are and are not expressed (exemplified by Hox gene duplications in different animal lineages; bottom left). This additionally allows a yet higher order recursion, as cells in different locales in the developing body can influence the expression of genes of cells in other locales. The result of this recursive iteration is that the theme and variation logic of regulatory gene information can be re-expressed in the quite different 3 dimensional structure of multi celled body plans (right panels). Arrows marked 1 in these images indicate theme and variation within an organism. Arrows marked 2 indicate theme and variation between different lineages
Conclusions
The sequence of hypothetical molecular models discussed here falls well short of explaining the origins of the “genetic code.” Indeed, it posits an evolutionary sequence that assumes that protein-like molecules are present long before nucleic acids (possibly arising from the prebiotic formation of hydrogen cyanide polymers; see Das et al. (2019) for a current review). This inverts the currently popular view that replicating molecules intrinsically constitute biological information. This popular assumption has implicitly reduced the concept of information to pattern replication without reference. As a result it begs the question of the origin of functional significance.
The logic of the autogenic approach, though not able to directly account for the evolution of the DNA-to-amino acid “code,” provides something more basic. It provides a “proof of principle” of a sort, showing step-by-chemically-realistic-step how a molecule like RNA or DNA could acquire the property of recording and instructing the dynamical molecular relationships that constitute and maintain the molecular system of which it is a part. In short, it explains how a molecule can become about other molecules. Importantly, this analysis inverts the logic that treats RNA and DNA replication as intrinsically informational and instead shows how the information-bearing function of nucleic acids is due to their ability to embody constraints inherited from the codependent dynamics of an open molecular` system able to repair itself. This may point the way to an alternative strategy for exploring the origin of the genetic code. Rather than thinking of the problem from an information molecule first perspective (how nucleic acid structure came to inform protein dynamics), it might be instructive to ask the question the other way around (how protein dynamics came to be reflected in nucleic acid structure). In other words, it might make sense to invert the order of Crick’s central dogma when considering the evolution of the genetic code.
Data Availability
Not applicable.
Code availability
Not applicable.
Notes
I will use this hyphenated version of the term representation in order to avoid any implicit psychologism and instead to highlight the more basic sense of being presented again in some other form.
Recursive self-maintenance (i.e. the self-maintenance of self-maintenance) is a term introduced by the philosopher and cognitive scientist Mark Bickhard to distinguish the structure of living self-maintenance from the self-perpetuating dynamics characteristic of non-living self-organized processes. His example of the latter is a candle flame that generates sufficient heat to vaporize wax that fuels the flame to vaporize additional wax. Such a system is self-maintaining, but is not organized to additionally maintain this capacity to maintain itself. See Bickhard (1993) for an early account.
See for example Peirce’s discussions in CP 4.536, 8.314, 333, 343; EP 2:404–9; and SS 111.
Pattee often referred to the passive constraint-bearing medium as a “symbol”—by which he meant a generic sign vehicle—and emphasized that although these “symbols” function to constrain the dynamics, their different physical form allows them to be manipulated and modified independently of the dynamic processes they inform. He anticipates the autogenic approach when he says, “Boundary conditions formed by local structures are often called constraints. Informational structures such as symbol vehicles are a special type of constraint.” From Pattee (2006),
For example Volume 8 Issue 2 of Biosemiotics edited by Jesper Hoffmeyer (2015) was a special issue dedicated to a discussion of the concept of semiotic scaffolding, and included articles by more than a dozen scholars discussing its relevance across many fields.
References
Barbieri, M. (2015). Code Biology: A New Science of Life. NJ: Springer.
Bickhard, M. H. (1993). Representational content in humans and machines. Journal of Experimental and Theoretical Artificial Intelligence, 5, 285–333.
Crick, F. H. (1958). On Protein Synthesis. In F. K. Sanders (Ed.), Symposia of the Society for Experimental Biology, Number XII: The Biological Replication of Macromolecules (pp. 138–163). Cambridge University Press.
Das, T., Ghule, S., & Vanka, K. (2019). Insights into the origin of life: Did it begin from HCN and H2O? ACS Central Science, 5(9), 1532–1540.
Dawkins, R. (1976). The Selfish Gene. Oxford University Press.
Deacon, T. (2006). Reciprocal linkage between self-organizing processes is sufficient for self-reproduction and evolvability. Biological Theory, 1(2), 136–149.
Dyson, F. (1985). Origins of Life. Cambridge University Press
Fano, Robert (2001). Interview in Aftab, Cheung, Kim, Thakkar, Yeddanapudi (2001) Information Theory & The Digital Revolution 6.933 Project History, Massachusetts Institute of Technology. https://www.web.mit.edu/6.933/www/Fall2001/Shannon2.pdf,p9. Accessed 24 Sept 2021
Gibson, J. J. (1950). The Perception of the Visual World. Houghton Mifflin.
Gilbert, W. (1986). The RNA world. Nature, 319, 618.
Hoffmeyer, J. (2015) Introduction: Semiotic scaffolding. Biosemiotics 8 (2) 153–158 & J. Hoffmeyer (Ed.). Special issue. (pp. 159–360)
Hoffmeyer, J. (2007). Semiotic scaffolding of living systems. In M. Barbieri (Ed.), Introduction to Biosemiotics: The New Biological Synthesis (pp. 149–166). Springer.
Hoffmeyer, J. (2014a). Semiotic scaffolding: A biosemiotic link between sema and soma. In K. R. Cabell & J. Valsiner (Eds.), The catalyzing mind: Beyond models of causality (pp. 95–110). Springer.
Hoffmeyer, J. (2014b). The semiome: From genetic to semiotic scaffolding. Semiotica, 198, 11–31.
Joyce, G. F. (1989). RNA evolution and the origins of life. Nature, 338, 217–224.
Joyce, G. F. (1991). The rise and fall of the RNA world. The New Biologist, 3, 399–407.
Kauffman, S., Logan, R. K., Este, R., Goebel, R., Hobill, D., & Smulevich, I. (2007). Propagating organization: An inquiry. Biology and Philosophy, 23, 27–45.
Maslow, A. (1966). The Psychology of Science: A Reconnaissance. Harper Collins
Pattee, H. H. (1969). How Does a Molecule Become a Message? A. Lang (Ed.) Communication in Development, Developmental Biology Supplement. (pp. 1–16). Academic Press, Cambridge.
Pattee, H. H. (1968). The Physical Basis of Coding and Reliability in Biological Evolution. In C. H. Waddington (Ed.), Towards a Theoretical Biology, 1, Prolegomena (pp. 67–93). Edinburgh University Press.
Pattee, H. H. (2001). The physics of symbols: Bridging the epistemic cut. Bio Systems, 60, 5–21.
Pattee, H. H. (2006). The physics of autonomous biological information. Biological Theory, 1(3), 224–226.
Peirce, Charles S. (1931–1935). Collected Papers of Charles Sanders Peirce, Vol. 1–6. C. Hartshorne & P. Weiss (Eds.). Harvard University Press.
Schrödinger, E. (1944). What Is Life? Cambridge University Press.
Shannon, C. E. (1948). A mathematical theory of communication. Bell System Technical Journal, 27, 623–656.
Spencer-Brown, G. (1972). Laws of Form. USA: The Julian Press.
Watson, J. D., & Crick, F. H. (1953). A structure for deoxyribose nucleic acid. Nature, 171, 737–738.
Funding
Partial support for the preparation of this manuscript was provided by The Human Energy Project and the Stanford University Boundaries of Humanity Project.
Author information
Authors and Affiliations
Contributions
100%.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare that he has no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
In honor of Howard Pattee’s (1969) “How Does a Molecule Become a Message?”.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Deacon, T.W. How Molecules Became Signs. Biosemiotics 14, 537–559 (2021). https://doi.org/10.1007/s12304-021-09453-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12304-021-09453-9