Abstract
Glycine-rich RNA-binding proteins (GRPs) are essential for many physiological and biochemical processes in plants, especially the response to environmental stresses. GRPs exist widely in angiosperms and gymnosperms plant species; however, their roles in Vitis vinifera are still poorly understood. To characterize VviGRP gene family, we performed a genomic survey, bioinformatics and expression analysis of VviGRPs in grape. We identified nineteen VviGRPs gene family members. The result of bioinformatics analysis showed their motif distribution, gene structure characteristics and chromosomal locations. Then we carried out synteny and phylogenetic analysis to study the origin and evolutionary relationship of GRPs. Tissue-specific expression analysis showed that VviGRPs have different expression patterns. Meanwhile, we studied expression profiles of seventeen ovule-expressed genes during seed development of stenospermocarpic seedless and seeded grapes, and the result showed that most of them have much higher relative expression levels in stenospermocarpic seedless grapes than that of seeded one before 25 days after full bloom (DAFB). It is suggested that VviGRPs may involve in the seed development process. Taken together, our research indicated that VviGRPs are related to seed development and will be beneficial for further investigations into the seed abortion mechanism under stenospermocarpic grapes.






Similar content being viewed by others
References
Albà MM, Pagès M (1998) Plant proteins containing the RNA recognition motif. Trends Plant Sci 3(97):15–21
Aneeta, Sanan-Mishra N, Tuteja N, Sopory SK (2002) Salinity-and ABA-induced up-regulation and light-mediated modulation of mRNA encoding glycine-rich RNA-binding protein from Sorghum bicolor. Biochem Biophys Res Commun 296(5):1063–1068
Bergeron D, Beauseigle D, Bellemare G (1993) Sequence and expression of a gene encoding a protein with RNA-binding and glycine-rich domains in Brassica napus. Biochem Biophys Acta 1216:123–125
Burd CG, Dreyfuss G (1994) Conserved structures and diversity of functions of RNAbinding proteins. Science 265(5172):615–621
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55(4):611–622
Cabezas JA, Cervera MT, Ruiz-García L, Carreño J, Martínez-zapater JM (2006) A genetic analysis of seed and berry weight in grapevine. Genome 12:1572–1585
Chen X, Zeng QC, Lu XP, Yu DQ, Li WZ (2010) Characterization and expression analysis of four glycine-rich RNA-binding proteins involved in osmotic response in tobacco (Nicotiana tabacum cv. Xanthi). Agric Sci China 9(11):1577–1587
De Oliveira DE, Seurinck J, Inzé D, Van Montagu M, Botterman J (1990) Differential expression of five Arabidopsis genes encoding glycine-rich proteins. Plant Cell 2(5):427–436
Dudley JW (1993) Molecular markers in plant improvement manipulation of genes affecting quantitiative. Crop Sci 33:660–668
Dunn MA, Brown K, Lightowlers R, Hughes MA (1996) A low-temperature-responsive gene from barley encodes a protein with single-stranded nucleic acid-binding activity which is phosphorylated in vitro. Plant Mol Biol 30:947–959
Fedoroff NV (2002) RNA-binding proteins in plants: the tip of an iceberg? Curr Opin Plant Biol 5:452–459
Fischer BM, Salakhutdinov I, Akkurt M, Eibach R, Edwards KJ, Töpfer R, Zyprian EM (2004) Quantitative trait locus analysis of fungal disease resistance factors on a molecular map of grapevine. Theor Appl Genet 108:501–515
Fusaro A, Sachrtto-Martins G (2007) Bloming time for plant glycine-rich proteins. Plant Signal Behav 2:e1–e2
Fusaro AF, Bocca SN, Ramos RL, Barrôco RM, Magioli C, Jorge VC, Coutinho TC, Rangel-Lima CM, Rycke RD, Inzé D, Engler G, Sachetto-Martins G (2007) A cold-induced nucleo-cytoplasmic RNA-binding protein, has a role in flower and seed development. Planta 225(6):1339–1351
Gomez J, Sanchez-Martinez D, Stiefel V, Rigau J, Puigdomenech P, Pages M (1988) A gene induced by the plant hormone abscisic acid in response to water stress encodes a glycinerich protein. Nature 334:262–264
Grimplet J, Adamblondon AF, Bert PF, Bitz O, Cantu D, Davies C, Delrot S, Pezzotti M, Rombauts S, Cramer GR (2014) The grape gene nomenclature system. BMC Genom 15:1077
Hanano S, Sugita M, Sugiura M (1996) Isolation of a novel RNA-binding protein and its association with a large ribonucleoprotein particle present in the nucleoplasm of tobacco cells. Plant Mol Biol 31:57–68
Hirose T, Sugita M, Sugiura M (1993) cDNA structure, expression and nucleic-acid binding properties of three RNA binding proteins in tobacco: occurrence of tissue alternative splicing. Nucleic Acids Res 21:3981–3987
Hirose T, Sugita M, Sugiura M (1994) Characterization of a cDNA encoding a novel type of RNA-binding protein in tobacco: its expression and nucleic acid-binding properties. Mol Gen Genet 244:360–366
Jaillon O, Aury JM, Noel B, Policriti A, Clepet C, Casagrande A, Choisne N, Aubourg S, Vitulo N, Jubin C, Vezzi A, Legeai F, Hugueney P, Dasilva C, Horner D, Mica E, Jublot D, Poulain J, Bruyere C, Billault A, Segurens B, Gouyvenoux M, Ugarte E, Cattonaro F, Anthouard V, Vico V, Del Fabbro C, Alaux M, Di Gaspero G, Dumas V, Felice N, Paillard S, Juman I, Moroldo M, Scalabrin S, Canaguier A, Le Clainche I, Malacrida G, Durand E, Pesole G, Laucou V, Chatelet P, Merdinoglu D, Delledonne M, Pezzotti M, Lecharny A, Scarpelli C, Artiguenave F, Pe ME, Valle G, Morgante M, Caboche M, Adam-Blondon AF, Weissenbach J, Quetier F, Wincker P (2007) The grape genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449:463–467
Jung HJ, Park SJ, Kang HS (2013) Regulation of RNA metabolism in plant development and stress responses. J Plant Biol Res 56(3):123–129
Kim JS, Park SJ, Kwak KJ, Kim YO, Kim JY, Song J, Jang B, Jung CH, Kang H (2007a) Cold shock domain proteins and glycine-rich RNA-binding proteins from Arabidopsis thaliana can promote the cold adaptation process in Escherichia coli. Nucleic Acids Res 35(2):506–516
Kim YJ, Park SJ, Jang B, Jung CH, Ahn SJ, Goh CH, Cho K, Han O, Kang H (2007b) Functional characterization of a glycine-rich RNA-binding protein 2 in Arabidopsis thaliana under abiotic stress conditions. Plant J 50:439–451
Kim YO, Pan S, Jung CH, Kang H (2007c) A zinc finger-containing glycine-rich RNA-binding protein, at RZ-1a, has a negative impact on seed germination and seedling growth of Arabidopsis thaliana under salt or drought stress conditions. Plant Cell Physiol 48:1170–1181
Kim JY, Kim WY, Kwak KJ, Oh SH, Han YS, Kang H (2010a) Zinc finger-containing glycine-rich RNA-binding protein in Oryza sativa has an RNA chaperone activity under cold stress conditions. Plant Cell Environ 33(5):759–768
Kim JY, Kim WY, Kwak KJ, Oh SH, Han YS, Kang H (2010b) Glycine-rich RNA binding proteins are functionally conserved in Arabidopsis thaliana and Oryza sativa during cold adaptation process. J Exp Bot 61(9):2317–2325
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645
Kwak KJ, Kim YO, Kang H (2005) Characterization of transgenic Arabidopsis plants overexpressing GR-RBP4 under high salinity, dehydration, or cold stress. J Exp Bot 56(421):3007–3016
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Liu SM, Sykes SR, Clingeleffer PR (2008) Effect of culture medium, genotype, and year of cross on embryo development and recovery from in vitro cultured ovules in breeding stenospermocarpic seedless grape varieties. Aust J Agric Res 59:175–182
Lu Y, Sun J, Yang Z, Zhao C, Zhu M, Ma D, Dong T, Zhou Z, Liu M, Yang D, Li Z, Xu T (2019) Genome-wide identification and expression analysis of glycine-rich RNA-binding protein family in sweet potato wild relative Ipomoea trifida. Gene 686:177–186
Ludevid MD, Freire MA, Gomez J, Burd CG, Albericio F, Giralt E, Dreyfuss G, Pages M (1992) RNA binding characteristics of a 16 kDa glycine‐rich protein from maize. Plant J 2:999–1003
Mangeon A, Magioli C, Menezes-Salgueiro AD, Cardeal V, de Oliveira C, Galvão VC, Margis R, Engler G, Sachetto-Martins G (2009) AtGRP5, a vacuole-located glycine-rich protein involved in cell elongation. Planta 230(2):253–265
Mangeon A, Junqueira RM, Sachetto-Martins G (2010) Functional diversity of the plant glycine-rich proteins superfamily. Plant Signal Behav 5(2):99–104
Maruyama K, Sato N, Ohta N (1999) Conservation of structure and cold-regulation of RNA-binding proteins in cyanobacteria: probable convergent evolution with eukaryotic glycine-rich RNA-binding proteins. Nucleic Acids Res 27:2029–2036
Molina A, Mena M, Carbonero P, Garcýa Olmedo F (1997) Differential expression of pathogen-responsive genes encoding two types of glycine-rich proteins in barley. Plant Mol Biol 33:803–810
Moriguchi K, Sugita M, Sugiura M (1997) Structure and subcellular localization of a small RNA-binding protein from tobacco. Plant J 6:825–834
Nishiyama H, Higashitsuji H, Yokoi H, Itoh K, Danno S, Matsuda T, Fujita J (1997) Cloning and characterization of human CIRP (cold-inducible RNA-binding protein) cDNA and chromosomal assignment of the gene. Gene 19:115–120
Nocker S, Vierstra RD (1993) Two cDNAs from Arabidopsis thaliana encode putative RNA binding proteins containing glycine-rich domains. Plant Mol Biol 21:695–699
Nomata T, Kabeya Y, Sato N (2004) Cloning and characterization of glycine-rich RNA-binding protein cDNAs in the moss Physcomitrella patens. Plant Cell Physiol 45:48–56
Notsuka K, Tsuru T, Shiraishi M (2001) Seedless-seedless grape hybridization via in-ovulo embryo culture. J Jpn Soc Hortic Sci 70:7–15
Park AR, Cho SK, Yun UJ, Jin MY, Lee SH, Sachetto-Martins G, Park OK (2001) Interaction of the Arabidopsis receptor protein kinase wak1 with a glycine-rich protein, AtGRP-3. J Biol Chem 276:26688–26693
Potenza C, Thomas SH, Sengupta-Gopalan C (2001) Genes induced during early response to Meloidogyne incongnita in roots of resistant and susceptible alfalfa cultivars. Plant Sci 161:289–299
Roytchev V (1998) Inheritance of grape seedlessness in seeded and seedless hybrid cominations of grape cultivars with complex genealogy. Am J Enol Vitic 49:302–305
Sachetto-Martins G, Franco LO, Oliveira DE (2000) Plant glycine-rich proteins: a family or just proteins with a common motif? Biochim Biophys Acta 1492(1):1
Sahi C, Agarwal M, Singh A, Grover A (2007) Molecular characterization of a novel isoform of rice (Oryza sativa L.) glycine rich-RNA binding protein and evidence for its involvement in high temperature stress response. Plant Sci 173(2):144–155
Schoning JC, Streitner C, Page DR, Hennig S, Uchida K, Wolf E, Furuya M, Staiger D (2007) Autoregulation of the circadian slave oscillator component AtGRP7 and regulation of its targets is impaired by a single RNA recognition motif point mutation. Plant J 52(6):1119–1130
Shinozuka H, Hisano H, Yoneyama S, Shimamoto Y, Jones ES, Forster JW, Yamada T, Kanazawa A (2006) Gene expression and genetic mapping analyses of a perennial ryegrass glycine-rich RNA-binding protein gene suggest a role in cold adaptation. Mol Genet Genom 275(4):399–408
Staiger D, Zecca L, Wieczorek Kirk DA, Apel K, Eckstein L (2003) The circadian clock regulated RNA-binding protein At-GRP7 autoregulates its expression by influencing alternative splicing of its own pre-mRNA. Plant J 33(2):361–371
Steinert PM, Mack JW, Korge BP, Gan SQ, Haynes SR, Steven AC (1991) Glycine loops in proteins: their occurrence in certain intermediate filament chains, loricrins and singlestranded RNA-binding proteins. Int J Biol Macromol 13:130–139
Stephen JR, Dent KC, Finch-Savage WE (2003) A cDNA encoding a cold-induced glycine-rich RNA binding protein from Prunus avium expressed in embryonic axes. Gene 320:177–183
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tang H, Wang X, Bowers JE, Ming R, Alam M, Paterson AH (2008) Unraveling ancient hexaploidy through multiply-aligned angiosperm gene maps. Genome Res 18:1944–1954
Tao TY, Ouellet T, Dadej K, Miller SS, Johnson DA, Singh J (2006) Characterization of a novel glycine-rich protein from the cell wall of maize silk tissues. Plant Cell Report 25:848–858
Ueki S, Citovsky V (2002) The systemic movement of a tobamovirus is inhibited by a cadmium-ion-induced glycine-rich protein. Nat Cell Biol 4:478–486
Ueki S, Citovsky V (2005) Identification of an interactor of cadmium ion-induced glycine-rich protein involved in regulation of callose levels in plant vasculature. Proc Natl Acad Sci USA 34:12089–12094
Varner JE, Cassab GI (1986) Molecular biology: a new protein in petunia. Nature 323:110
Vermel M, Guermann B, Delage L, Grienenberber JM, Marechal-Drouard L, Gualberto JM (2002) A family of RRM-type RNA-binding proteins specific to plant mitochondria. Proc Natl Acad Sci USA 99:5866–5871
Wang L, Xie XQ, Yao WK, Wang J, Ma FL, Wang C, Yang YZ, Tong WH, Zhang JX, Xu Y, Wang XP, Zhang CH, Wang YJ (2017) RING-H2 type E3 gene VpRH2 from Vitis pseudoreticulata improves resistance to powdery mildew by interacting with VpGRP2A. J Exp Bot 68(7):1669–1687
Zhang JH, Zhao YH, Xiao HL, Zheng YL, Yue B (2014) Genome-wide identification, evolution, and expression analysis of RNA-binding glycine-rich protein family in maize. J Integr Plant Biol 56(10):1020–1031
Acknowledgements
This work was supported by the National Key R&D Program of China [2019YFD1001405]; the Modern Agro-industry Technology Research System [No. CARS-30-yz-7]; and the Natural Science Foundation of China [No. 32072554].
Funding
National Key R&D Program of China [2019YFD1001405]; the Modern Agro-industry Technology Research System [No. CARS-30-yz-7]; and the Natural Science Foundation of China [No. 32072554].
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. CZ and YL conceived and designed the whole experiments; YT and CH performed the experiments; YT analysed the data; CZ, YW contributed reagents/materials/analysis tools; YT and CH wrote the manuscript; YL reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12298_2021_1082_MOESM1_ESM.tif
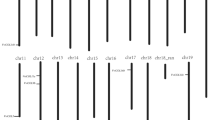
ESM_1 RNA extraction of six tissues or organs and ovules at different development stages from ‘Pinot Noir’ and ‘Thompson Seedless’. (a) RNA extraction of six tissues or organs from ‘Pinot Noir’. Line 1, RNA of leaf; Line 2, RNA of stem; Line 3, RNA of flower; Line 4, RNA of flower bud; Line 5, RNA of ovule; Line 6, RNA of tendril. (b) RNA extract from ‘Pinot Noir’ ovules at different development stages. Line 1, RNA of ‘Pinot Noir’ ovules in 10 day after full-bloom (DAFB); Line 2, RNA of ‘Pinot Noir’ ovules in 15 DAFB; Line 3, RNA of ‘Pinot Noir’ ovules in 20 DAFB; Line 4, RNA of ‘Pinot Noir’ ovules in 25 DAFB; Line 5, RNA of ‘Pinot Noir’ ovules in 30 DAFB; Line 6, RNA of ‘Pinot Noir’ ovules in 35 DAFB. (c) RNA extract from ‘Pinot Noir’ and ‘Thompson Seedless’ovules at different development stages. Line 1, RNA of ‘Pinot Noir’ ovules in 40 DAFB; Line 2, RNA of ‘Pinot Noir’ ovules in 45 DAFB; Line 3, RNA of ‘Thompson Seedless’ ovules in 10 DAFB; Line 4, RNA of ‘Thompson Seedless’ ovules in 15 DAFB; Line 5, RNA of ‘Thompson Seedless’ ovules in 20 DAFB; Line 6, RNA of ‘Thompson Seedless’ ovules in 25 DAFB; Line 7, RNA of ‘Thompson Seedless’ ovules in 30 DAFB; Line 8, RNA of ‘Thompson Seedless’ ovules in 35 DAFB. (d) RNA extract from ‘Thompson Seedless’ovules at different development stages. Line 1, RNA of ‘Thompson Seedless’ ovules in 40 DAFB; Line 2, RNA of ‘Thompson Seedless’ ovules in 45 DAFB (TIF 12504 KB)
12298_2021_1082_MOESM5_ESM.tif
ESM_5. RT-qPCR primers location and dissolution curve of VviGRP5Lh. (a) RT-qPCR primers location of VviGRP5Lh. (a) Dissolution curve of VviGRP5Lh. (TIF 14309 KB)
12298_2021_1082_MOESM6_ESM.tif
ESM_6 Dissolution curve of VviGRP in RT-qPCR. (a) Dissolution curve of VviACTIN. (a) Dissolution curve of VviACTIN. (b) Dissolution curve of VviGADPH. (c) Dissolution curve of VviRZ-1A. (d) Dissolution curve of VviGRP5Le. (e) Dissolution curve of VviGRP5Lg. (f) Dissolution curve of VviGRP5Lh. (g) Dissolution curve of VviGRP4L. (h) Dissolution curve of VviGRP5Ld. (i) Dissolution curve of VviGRP5Lf. (j) Dissolution curve of VviGRP5Lc. (k) Dissolution curve of VviGRP3. (l) Dissolution curve of VviRZ-1B. (m) Dissolution curve of VviGRP7. (n) Dissolution curve of VviGRP2. (o) Dissolution curve of VviRZ-1C. (p) Dissolution curve of VviGRP5Li. (q) Dissolution curve of VviGRP5Lb. (r) Dissolution curve of VviGRP5. (s) Dissolution curve of VviSUA. (TIF 27180 KB)
Rights and permissions
About this article
Cite this article
Tang, Y., Huang, C., Li, Y. et al. Genome-wide identification, phylogenetic analysis, and expression profiling of glycine-rich RNA-binding protein (GRPs) genes in seeded and seedless grapes (Vitis vinifera). Physiol Mol Biol Plants 27, 2231–2243 (2021). https://doi.org/10.1007/s12298-021-01082-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-021-01082-3