Abstract
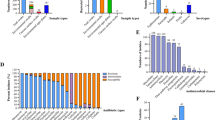
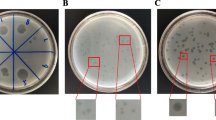
The consumption of fresh produce has led to increase in antibiotic-resistant (AR) Salmonella outbreaks. In this study, indigenous Salmonella was isolated from a total of two hundred-two samples including fresh produce and agricultural environmental samples in Korea. After biochemical confirmation using the Indole, Methyl Red, Voges-Proskauer, Citrate tests, presumable Salmonella isolates were identified by 16S rRNA sequencing. Identified Salmonella isolates were evaluated for antibiotic susceptibility against twenty-two antibiotics. The specificity and the efficiency of plating (EOP) of vB_SalS_KFSSM were evaluated against fifty-three bacterial strains. Twenty-five suspected Salmonella were isolated and confirmed by the positive result for methyl red and citrate, of which ten were identified as Salmonella spp. through 16S rRNA gene sequencing. Eight Salmonella isolates (4.0%, n = 8/202) were resistant to at least one antibiotic, among which five were multi-drug resistant. As a lytic phage against Salmonella spp. CMGS-1, vB_SalS_KFSSM was isolated from cow manure. The phage was observed as a tailed phage belonging to the class Caudoviricetes. It exhibited an intra-broad specificity against four indigenous AR Salmonella isolates, two indigenous Salmonella isolates, and five other Salmonella serotypes with great efficiencies (EOP ≥ 0.75). Thus, this study suggested the potential of vB_SalS_KFSSM to combat indigenous AR Salmonella.


Similar content being viewed by others
Data availability
All data generated during this study are included in this published article.
References
Abakpa, G. O., Umoh, V. J., Ameh, J. B., Yakubu, S. E., Kwaga, J. K. P., & Kamaruzaman, S. (2015). Diversity and antimicrobial resistance of Salmonella enterica isolated from fresh produce and environmental samples. Environmental Nanotechnology, Monitoring & Management, 3, 38–46.
Abatcha, M. G., Effarizah, M. E., & Rusul, G. (2018). Prevalence, antimicrobial resistance, resistance genes and class 1 integrons of Salmonella serovars in leafy vegetables, chicken carcasses and related processing environments in Malaysian fresh food markets. Food Control, 91, 170–180.
Ackermann, H. W., DuBow, M. S., Jarvis, A. W., Jones, L. A., Krylov, V. N., Maniloff, J., Rocourt, J., Safferman, R. S., Schneider, J., Seldin, L., et al. (1992). The species concept and its application to tailed phages. Archives of Virology, 124, 69–82.
Alenazy, R. (2022). Antibiotic resistance in Salmonella: Targeting multidrug resistance by understanding efflux pumps, regulators and the inhibitors. Journal of King Saud University-Science., 34, 102275.
Behera, B. K., Paria, P., Das, A., & Das, B. K. (2022). Molecular identification and pathogenicity study of virulent Citrobacter freundii associated with mortality of farmed Labeo rohita (hamilton 1822), in India. Aquaculture, 547, 737437.
CDC, Centers for Disease Control and Prevention. (2019). Antibiotic Resistance Threats in the United States, 2019. U.S. Department of Health and Human Services, Centers for Disease Control and Prevention. Atlanta, Georgia, USA.
Choi, I. Y., Lee, C., Song, W. K., Jang, S. J., & Park, M. K. (2019). Lytic KFS-SE2 phage as a novel bio-receptor for Salmonella Enteritidis detection. Journal of Microbiology, 57, 170–179.
Choi, I. Y., Lee, J. H., Kim, H. J., & Park, M. K. (2017). Isolation and characterization of a novel broad-host-range bacteriophage infecting Salmonella enterica subsp. enterica for biocontrol and rapid detection. Journal of Microbiology and Biotechnology, 27, 2151–2155.
Choi, I. Y., Park, D. H., Chin, B. A., Lee, C., Lee, J., & Park, M. K. (2020). Exploring the feasibility of Salmonella Typhimurium-specific phage as a novel bio-receptor. Journal of Animal Science and Technology, 62, 668–681.
Clarridge, J. E., 3rd. (2004). Impact of 16S rRNA gene sequence analysis for identification of bacteria on clinical microbiology and infectious diseases. Clinical Microbiology Reviews, 17, 840–862.
CLSI, Clinical and Laboratory Standards Institute. (2012). Performance standards for antimicrobial susceptibility testing. 33rd ed., CLSI supplement M100. Clinical and Laboratory Standards Institute.
Fong, K., Wong, C. W. Y., Wang, S., & Delaquis, P. (2021). How broad is enough: The host range of bacteriophages and its impact on the agri-food sector. PHAGE, 2, 83–91.
Gal-Mor, O., Boyle, E. C., & Grassl, G. A. (2014). Same species, different diseases: How and why typhoidal and non-typhoidal Salmonella enterica serovars differ. Frontiers in Microbiology, 5, 391.
Hailu, W., Helmy, Y. A., Carney-Knisely, G., Kauffman, M., Fraga, D., & Rajashekara, G. (2021). Prevalence and antimicrobial resistance profiles of foodborne pathogens isolated from dairy cattle and poultry manure amended farms in northeastern Ohio, the United States. Antibiotics, 10, 1450.
Hall, T. A. (1999). BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41, 95–98.
Hooton, S. P., Timms, A. R., Rowsell, J., Wilson, R., & Connerton, I. F. (2011). Salmonella Typhimurium-specific bacteriophage ΦSH19 and the origins of species specificity in the Vi01-like phage family. Virology Journal, 8, 498.
Huang, C., Shi, J., Ma, W., Li, Z., Wang, J., Li, J., & Wang, X. (2018a). Isolation, characterization, and application of a novel specific Salmonella bacteriophage in different food matrices. Food Research International, 111, 631–641.
Huang, C., Virk, S. M., Shi, J., Zhou, Y., Willias, S. P., Morsy, M. K., Abdelnabby, H. E., Liu, J., Wang, X., & Li, J. (2018b). Isolation, characterization, and application of bacteriophage LPSE1 against Salmonella enterica in ready to eat (RTE) foods. Frontiers in Microbiology, 9, 1046.
Khan, M. A. S., & Rahman, S. R. (2022). Use of phages to treat antimicrobial-resistant Salmonella infections in poultry. Veterinary Sciences, 9, 438.
Khan Mirzaei, M., & Nilsson, A. S. (2015). Isolation of phages for phage therapy: A comparison of spot tests and efficiency of plating analyses for determination of host range and efficacy. PLoS ONE, 10, e0118557.
Kim, S. H., Adeyemi, D. E., & Park, M. K. (2021). Characterization of a new and efficient polyvalent phage infecting E. coli O157:H7, Salmonella spp., and Shigella sonnei. Microorganisms, 9, 2105.
Kim, S., Kim, S. H., Rahman, M., & Kim, J. (2018). Characterization of a Salmonella Enteritidis bacteriophage showing broad lytic activity against Gram-negative enteric bacteria. Journal of Microbiology, 56, 917–925.
Kowalska, B. (2023). Fresh vegetables and fruit as a source of Salmonella bacteria. Annals of Agricultural and Environmental Medicine, 30, 9–14.
Krumperman, P. H. (1983). Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of fecal contamination of foods. Applied and Environmental Microbiology, 46, 165–170.
Kuang, X., Hao, H., Dai, M., Wang, Y., Ahmad, I., Liu, Z., & Zonghui, Y. (2015). Serotypes and antimicrobial susceptibility of Salmonella spp. isolated from farm animals in China. Frontiers in Microbiology., 6, 602.
Kumar, S., Stecher, G., Li, M., Knyaz, C., & Tamura, K. (2018). MEGA X: Molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution, 35, 1547.
Lane, D. J. (1991). 16S/23S Rrna Sequencing. In E. Stackebrandt & M. Goodfellow (Eds.), Nucleic Acid Techniques in Bacterial Systematic (pp. 115–175). John Wiley and Sons.
Lee, C., Choi, I. Y., Park, D. H., & Park, M. K. (2020). Isolation and characterization of a novel Escherichia coli O157:H7-specific phage as a biocontrol agent. Journal of Environmental Health Science and Engineering, 18, 189–199.
Lee, H. E., Jeon, Y. B., Chin, B. A., Lee, S. H., Lee, H. J., & Park, M. K. (2023). Performance of wild, tailed, humidity-robust phage on a surface-scanning magnetoelastic biosensor for Salmonella Typhimurium detection. Food Chemistry, 409, 135239.
Li, J., Li, Y., Ding, Y., Huang, C., Zhang, Y., Wang, J., & Wang, X. (2021a). Characterization of a novel Siphoviridae Salmonella bacteriophage T156 and its microencapsulation application in food matrix. Food Research International, 140, 110004.
Li, Z., Li, W., Ma, W., Ding, Y., Zhang, Y., Yang, Q., Wang, J., & Wang, X. (2021b). Characterization and application of a lytic phage D10 against multidrug-resistant Salmonella. Viruses, 13, 1626.
Liu, A., Liu, Y., Peng, L., Cai, X., Shen, L., Duan, M., Ning, Y., Liu, S., Li, C., Liu, Y., et al. (2020). Characterization of the narrow-spectrum bacteriophage LSE7621 towards Salmonella Enteritidis and its biocontrol potential on lettuce and tofu. LWT, 118, 108791.
Liu, Y., Demina, T. A., Roux, S., Aiewsakun, P., Kazlauskas, D., Simmonds, P., Prangishvili, D., Oksanen, H. M., & Krupovic, M. (2021). Diversity, taxonomy, and evolution of archaeal viruses of the class Caudoviricetes. PLoS Biology, 19, e3001442.
Lu, M., Liu, H., Lu, H., Liu, R., & Liu, X. (2020). Characterization and genome analysis of a novel Salmonella phage vB_SenS_SE1. Current Microbiology, 77, 1308–1315.
Machado-Moreira, B., Richards, K., Brennan, F., Abram, F., & Burgess, C. M. (2019). Microbial contamination of fresh produce: What, where, and how? Comprehensive Reviews in Food Science and Food Safety, 18, 1727–1750.
Mahmoud, M., Askora, A., Barakat, A. B., Rabie, O. E. F., & Hassan, S. E. (2018). Isolation and characterization of polyvalent bacteriophages infecting multi drug resistant Salmonella serovars isolated from broilers in Egypt. International Journal of Food Microbiology, 266, 8–13.
Makwana, P. P., Nayak, J. B., Brahmbhatt, M. N., & Chaudhary, J. H. (2015). Detection of Salmonella spp. from chevon, mutton and its environment in retail meat shops in anand city (Gujarat), India. Veterinary World, 8, 388–392.
MFDS, Ministry of Food and Drug Safety. (2019). Salmonella spp. Korean Food Standards Codex. https://various.foodsafetykorea.go.kr/fsd/#/ext/Document/FC?searchNm=Salmonella&itemCode=FC0A065002003A223
Oh, J. H., & Park, M. K. (2017). Recent trends in Salmonella outbreaks and emerging technology for biocontrol of Salmonella using phages in foods: A review. Journal of Microbiology and Biotechnology, 27, 2075–2088.
Park, D. W., & Park, J. H. (2021). Characterization of a novel phage depolymerase specific to Escherichia coli O157:H7 and biofilm control on abiotic surfaces. Journal of Microbiology, 59, 1002–1009.
Pelyuntha, W., Ngasaman, R., Yingkajorn, M., Chukiatsiri, K., Benjakul, S., & Vongkamjan, K. (2021). Isolation and characterization of potential Salmonella phages targeting multidrug-resistant and major serovars of Salmonella derived from broiler production chain in Thailand. Frontiers in Microbiology., 12, 662461.
Pereira, C., Moreirinha, C., Rocha, R. J., Calado, R., Romalde, J. L., Nunes, M. L., & Almeida, A. (2016). Application of bacteriophages during depuration reduces the load of Salmonella Typhimurium in cockles. Food Research International, 90, 73–84.
Pérez-Rodríguez, F., & Mercanoglu Taban, B. (2019). A state-of-art review on multi-drug resistant pathogens in foods of animal origin: Risk factors and mitigation strategies. Frontiers in Microbiology, 10, 2091.
Pettengill, J., Tate, H., Gensheimer, K., Hsu, C., Ihrie, J., Markon, A., McDermott, P., Zhao, S., Strain, E., & Bazaco, M. (2020). Distribution of antimicrobial resistance genes across Salmonella enterica isolates from animal and nonanimal foods. Journal of Food Protection, 83, 295–304.
Pires, D. P., Costa, A. R., Pinto, G., Meneses, L., & Azeredo, J. (2020). Current challenges and future opportunities of phage therapy. FEMS Microbiology Reviews, 44, 684–700.
Pławińska-Czarnak, J., Wódz, K., Kizerwetter-Świda, M., Nowak, T., Bogdan, J., Kwieciński, P., Kwieciński, A., & Anusz, K. (2021). Citrobacter braakii yield false-positive identification as Salmonella, a note of caution. Foods, 10, 2177.
Rahman, M., Alam, M. U., Luies, S. K., Kamal, A., Ferdous, S., Lin, A., Sharior, F., Khan, R., Rahman, Z., Parvez, S. M., et al. (2021). Contamination of fresh produce with antibiotic-resistant bacteria and associated risks to human health: A scoping review. International Journal of Environmental Research and Public Health, 19, 360.
Reddy, S. P., Wang, H., Adams, J. K., & Feng, P. C. (2016). Prevalence and characteristics of Salmonella serotypes isolated from fresh produce marketed in the United States. Journal of Food Protection, 79, 6–16.
Rodríguez-Hernández, R., Bernal, J. F., Cifuentes, J. F., Fandiño, L. C., Herrera-Sánchez, M. P., Rondón-Barragán, I., & Verjan Garcia, N. (2021). Prevalence and molecular characterization of Salmonella isolated from broiler farms at the Tolima region—Colombia. Animals, 11, 970.
Sedeik, M. E., El-Shall, N. A., Awad, A. M., Elfeky, S. M., Abd El-Hack, M. E., Hussein, E. O., Alowaimer, A. N., & Swelum, A. A. (2019). Isolation, conventional and molecular characterization of Salmonella spp. from newly hatched broiler chicks. AMB Express, 9, 136.
Terreni, M., Taccani, M., & Pregnolato, M. (2021). New antibiotics for multidrug-resistant bacterial strains: Latest research developments and future perspectives. Molecules, 26, 2671.
Toyting, J., Miura, N., Utrarachkij, F., Tanomsridachchai, W., Belotindos, L. P., Suwanthada, P., Kapalamula, T. F., Kongsoi, S., Koide, K., Kim, H., et al. (2023). Exploration of the novel fluoroquinolones with high inhibitory effect against quinolone-resistant DNA gyrase of Salmonella Typhimurium. Microbiology Spectrum. https://doi.org/10.1128/spectrum.01330-23
Wang, C., Chen, Q., Zhang, C., Yang, J., Lu, Z., Lu, F., & Bie, X. (2017). Characterization of a broad host-spectrum virulent Salmonella bacteriophage fmb-p1 and its application on duck meat. Virus Research, 236, 14–23.
Yang, X., Wu, Q., Huang, J., Wu, S., Zhang, J., Chen, L., Wei, X., Ye, Y., Li, Y., Wang, J., et al. (2020). Prevalence and characterization of Salmonella isolated from raw vegetables in China. Food Control, 109, 106915.
Zafar, U., Taj, M. K., Nawaz, I., Zafar, A., & Taj, I. (2019). Characterization of Proteus mirabilis isolated from patient wounds at bolan medical complex hospital, quetta. Jundishapur Journal of Microbiology, 12, e87963.
Zhao, F., Sun, H., Zhou, X., Liu, G., Li, M., Wang, C., Liu, S., Zhuang, Y., Tong, Y., & Ren, H. (2019). Characterization and genome analysis of a novel bacteriophage vB_SpuP_Spp16 that infects Salmonella enterica serovar pullorum. Virus Genes, 55, 532–540.
Acknowledgements
This work was supported by Korea Institute of Planning and Evaluation for Technology in Food, Agriculture and Forestry (IPET) through High Value-added Food Technology Development Program, funded by Ministry of Agriculture, Food and Rural Affairs (MAFRA)(322014052HD030)
Funding
Korea Institute of Planning and Evaluation for Technology in Food, Agriculture and Forestry (IPET), 322014052HD030, Mi-Kyung Park.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflicts of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Choe, J., Kim, SH., Han, J.M. et al. Prevalence of Indigenous Antibiotic-Resistant Salmonella Isolates and Their Application to Explore a Lytic Phage vB_SalS_KFSSM with an Intra-Broad Specificity. J Microbiol. 61, 1063–1073 (2023). https://doi.org/10.1007/s12275-023-00098-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-023-00098-6