Abstract
Chronic ventricular pacing can lead to pacing-induced cardiomyopathy (PICM). Clinical data alone is insufficient to predict who will develop PICM. Our study aimed to evaluate the circulating miR profile associated with chronic right ventricular pacing in children with congenital complete AV block (CCAVB) and to identify candidate miRs for longitudinal monitoring. Clinical data and blood were collected from chronically paced children (N = 9) and compared with non-paced controls (N = 13). miR microarrays from the buffy coat revealed 488 differentially regulated miRs between groups. Pathway analysis predicted both adaptive and maladaptive miR signaling associated with chronic pacing despite preserved ventricular function. Greater profibrotic signaling (miRs-92a, 130, 27, 29) and sodium and calcium channel dysregulation (let-7) were seen in those paced > 10 years with the most dyregulation seen in a patient with sudden death vs. those paced < 10 years. These miRs may help to identify early adverse remodeling in this population.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Congenital complete atrioventricular block (CCAVB) is estimated to occur in ~ 1 in 15,000 live-born infants [1], and often requires chronic pacing starting at a very young age. Chronic right ventricular pacing produces electromechanical and inter- and intraventricular dyssynchrony which can lead to left ventricular remodeling, dysfunction, and heart failure resulting in pacing-induced cardiomyopathy (PICM) [2,3,4]. These findings likely occur late in the pathogenesis of PICM when significant adverse myocardial remodeling has already occurred, impacting the effectiveness of strategies such as cardiac resynchronization to improve ventricular function. Indeed, the success of cardiac resynchronization therapy in chronic RV paced children who develop LV dysfunction has shown mixed results [5]. However, there is currently no mechanism to monitor chronically paced patients other than by echocardiography, nor is there a mechanism to monitor for the development of adverse remodeling while ventricular function is still preserved since these patients do not undergo routine myocardial biopsy.
Circulating microRNAs (miRs) are non-coding RNA which regulate gene expression by RNA degradation or translational inhibition [6], and are emerging as key biomarkers of heart failure in cardiomyopathies as well as ischemic and congenital heart disease [7,8,9,10,11]. Marfella et al. discovered a miR profile in chronically paced adult heart failure patients which correlated with clinical improvement after cardiac resynchronization [12]. miRs-26b-5p, 145-5p, 92a-3p, 30e-5p, and 29a-3p were decreased in heart failure but increased after 1 year of cardiac resynchronization therapy, correlating with improved ejection fraction. The objectives of our study were to (i) evaluate the circulating miR profile associated with chronic right ventricular pacing and preserved ventricular function in children with CCAVB, and (ii) identify miRs which may be candidates for longitudinal monitoring.
Methods
Study Population
This was a single center, cross-sectional, pilot study of pediatric patients presenting for an electrophysiology outpatient evaluation at a tertiary center between December 2015 and June 2018. The study was approved by the Stanford University Institutional Review Board and assent and consent was obtained from all patients. Inclusion criteria for all patients included age 10–20 years, structurally normal hearts, and normal left ventricular function by echocardiography (left ventricular ejection fraction (LVEF) > 55%). Patients were recruited into two groups — (1) paced group (N = 9): patients with a diagnosis of CCAVB and right ventricular pacing for greater than 5 years, and (2) control group (N = 13): patients with well-controlled supraventricular tachycardia (atrioventricular node reentrant tachycardia (N = 9) or Wolff-Parkinson-White syndrome (N = 4)). Patients with hepatic dysfunction, renal dysfunction, pulmonary disease, or febrile illness were excluded due to the potential for these co-morbidities to affect miR expression, clearance, and excretion. Baseline patient demographics and clinical data including New York Heart Association (NYHA) heart failure class were collected. Follow-up data were collected at 3 years. Electrocardiogram and echocardiogram for each patient were performed during a single outpatient visit, at which time a peripheral blood sample was also collected.
Electrocardiogram and Pacemaker Support
Electrocardiograms (ECGs) performed at the time of a routine outpatient visit were reviewed and the following parameters were recorded — PR interval, QRS duration, and left bundle branch block. Pacemaker information including pacing system, site(s), duration, and percent atrial or ventricular pacing was also obtained. Follow-up data were collected at 3 years.
Echocardiography
Echocardiograms performed at the time of a routine outpatient visit were reviewed. Data on left ventricular function and size were collected from the reports — LVEF, left ventricular end diastolic dimension in systole (LVEDS) and in diastole (LVEDD), and indexed to body surface area (BSA). Follow-up data were collected at 3 years.
MicroRNA Expression Profiling and Analysis
Peripheral blood samples were centrifuged then the buffy coat was collected and stored at − 80 °C. Total RNA was isolated from frozen buffy coats (200 μL) using TRIzol (Invitrogen, Carlsbad, CA, USA) and purified using an RNeasy Mini Kit (QIAGEN, Dusseldorf, Germany) according to the manufacturer’s instructions. The expression of some miRs may vary in the different blood fractions (plasma, serum, buffy coat) [13, 14]. The concentration of miRs was the greatest in the buffy coat in our hands based on our prior animal data which showed that the most highly expressed cardiac miR-21 was highly expressed in the buffy coat vs. plasma. Similarly, Glinge et al. have shown that some of the most highly expressed cardiac miRs are highly expressed in the buffy coat vs. plasma [15]. We therefore assessed miR expression in the buffy coat vs. the plasma. RNA quality and quantity were measured using a QIAxpert kit (QIAGEN, Cat. #9,002,340, Germany). Gel electrophoresis was performed on the extracted RNA to ensure sample integrity following which small RNA were quantified using Agilent Bioanalyzer 2100 NANO analysis (Agilent, Santa Clara, USA). Total RNA (100 ng) was labeled with cyanine-3 to generate fluorescent miR, then purified and hybridized onto the SurePrint custom G3 miRNA Microarray (8 × 60 k, p/n G4871A). The slides were scanned and data extracted using Agilent feature extraction (FE) software for miR expression. miR expression analysis was performed using GeneSpring GX 14.9.1 software. Normalized data between the 20th and 100th percentiles with detected probes were used for further analysis. Quality control was performed, following which unpaired t-test with Benjamini–Hochberg multiple testing correction was applied to the data. Significantly altered miRs with a corrected p value of < 0.05 and with a fold change ≥ 2 up- or downregulated in paced vs. control groups were considered for further analysis. Putative target genes of significantly dysregulated miRs were identified using mirPATH v.7 DIANA tools. Gene Ontology (GO) and pathway analyses were performed to identify important biologic processes, nodal points, and pathways unique to the paced and control groups using Ingenuity Pathway Analysis and Cytoscape software.
Reverse Transcriptase Polymerase Chain Reaction
The expressions of a subset of dysregulated miRs identified by microarray were validated by Taqman two-step qRT-PCR: upregulated miR-205-3p, miR-210-5p, miR-214-3p, and miR-92a-3p, and downregulated miR-15b-5p, miR-126-5p, miR-130b-5p, miR-148-5p, miR-190a-5p, miR-29a-3p, and miR-27a-3p. 50 ng of RNA was reverse transcribed to cDNA followed by amplification (Applied Biosystems). Ambion mirVana qRT-PCR Primer Sets were used (Supplement Table 1). C. elegans miR-39 was used as the spike in control. Fold change (FC) in expression was assessed between paced vs. control patients using 2 power ΔΔCt method [16].
Statistical Analysis
Demographic, clinical, ECG, and echocardiogram data are presented as mean ± SEM. An unpaired, 2-tailed Student’s t-test was utilized for two-group comparisons of continuous variables with normal distribution and Mann–Whitney test when distribution was not normal. p ≤ 0.05 was considered significant.
Results
Patient Characteristics
Patient demographics and clinical data for the control and paced groups are shown in Table 1. The mean patient age was 15.03 ± 2 years in the control group and 15.7 ± 2.4 years in the paced group. Forty-six percent were males in the control group and 33% were males in the paced group. All were clinically asymptomatic at the time of sample collection with no hepatic or renal dysfunction and no acute illnesses. Thirty-three percent of the paced group had anti-Rho and anti-La antibodies at the time of diagnosis. Medications included one patient on a beta blocker and another on aspirin in the control group, and one patient on “as needed” albuterol for well-controlled asthma in the paced group. At the time of recruitment, none of the control patients had undergone an ablation or any other surgical procedure. All of the paced patients had undergone a pacemaker implantation and no other surgical procedure. Forty-four percent of the paced patients underwent pacemaker placement in the first week of life, 12% in the first year of life, and the remaining 44% between 5 and 10 years. As expected, the QRS duration was longer in the paced group. Data were collected from transthoracic echocardiograms performed at the time of blood collection (Table 1). LVEF was 63.3 ± 0.01% in controls vs. 60 ± 0.1% in paced patients (p = 0.04). All paced patients had dual chamber pacemakers (epicardial systems in 78%, and transvenous systems in 12%), were paced for greater than 5 years (per our inclusion criteria), and had ventricular pacing > 95% of the time (Table 2).
Unique Circulating miR Expression Profile Is Detected in Chronically Paced Children
miR quality was excellent in all blood samples (Fig. 1a, b). Normalized signal intensity in miR expression among the control (N = 13) and paced (N = 9) groups demonstrated normal distribution across all samples (Fig. 1c, d). A total of 488 miRs were differentially regulated between the control and paced groups (FC > 2/ < − 2, corrected p < 0.05). 296 miRs were upregulated in the paced group and 192 miRs were downregulated (Fig. 1e, f). To better understand the observed global miRNA expression profile changes, we performed pathway analysis on both upregulated and downregulated miRs.
Sample quality control and miR expression in control vs. paced patients. a RNA was isolated from the buffy coat and the presence of small RNA with no degradation was confirmed by 2D gel electrophoresis, b the presence of small RNAs between 25 and 50 nucleotides including miR (arrows) was confirmed by an electropherogram, c violin plot of normalized signal intensity in miR expression among the control (N = 13) and paced (N = 9) samples demonstrates normal distribution across all samples. d Scatter plot demonstrates global miR expression by microarray in controls vs. paced patients. e, f Volcano plot and heatmap showing significantly up- (red) and downregulated miRs (blue) in paced vs. controls groups by > twofold change in expression and corrected FDR p < 0.05. miR, microRNA
miRs Upregulated with Chronic Pacing Predict Poor Protein Quality Control and Cell–Cell Mechanical Uncoupling
Upregulated miRs predicted downregulation in 15 pathways which clustered into 4 groups (Table 3, Fig. 2), (i) structural proteins: O-glycan biosynthesis (protein folding, stability, trafficking to the right cellular location), glycosphingolipid biosynthesis of membranes; (ii) extracellular matrix (ECM): proteoglycans (non-structural components of ECM), adherens junctions which mechanically couple and reinforce cardiomyocytes, Wnt/TGF-β signaling which mediate fibrosis; (iii) metabolism: FoxO signaling pathway which regulates gluconeogenesis and glycogenolysis by insulin signaling and counteracts oxidative stress; and (iv) protein turnover and degradation: lysine degradation which regulates essential amino acid metabolites, ubiquitin proteosomal and lysosomal systems mediating protein quality control, Hippo signaling which is an antiproliferative and cell death pathway.
miR regulation with chronic ventricular pacing. Network map highlights significant Gene Ontology (GO) biological processes predicted by the miRs dysregulated in paced patients vs. controls. miRs upregulated with chronic pacing predict downregulation of proteins involved in cell adhesion, glucose metabolism, and protein quality control. miRs downregulated with chronic pacing predict upregulation of oxidative stress and neurotransmitter signaling, with decreased response to estrogen and nitric oxide signaling. miRs dysregulated with chronic pacing also predicted mixed regulation of components involved in protein folding and extracellular matrix. GO biological processes mapped using Cytoscape version 3.8.2. TNF, tumor necrosis factor. TGF, transforming growth factor beta. NO, nitric oxide
miRs Downregulated with Chronic Pacing Predict Altered Parasympathetic Tone as well as Upregulation of TGF-β and Fatty Acid Metabolism
Downregulated miRs predicted upregulation in 16 pathways which clustered into 4 groups (Table 4, Fig. 2), (i) O-glycan biosynthesis; (ii) extracellular matrix: profibrotic TGF-β signaling, ECM-receptor interaction which includes the crosstalk between ECM structural proteins such as collagens, laminin and fibronectin, and cells via transmembrane proteins such as integrins and proteoglycans; (iii) fatty acid biosynthesis; and (iv) neurotransmitter signaling enhancing both vagal nerve inhibition via GABA, vagal nerve activation via muscarinic receptors, and inhibition of estrogen signaling which impairs nitric oxide metabolism and vasodilation, L-type Ca and K channel regulation, and intracellular signal transduction pathways. In summary, components of structural and extracellular matrix are both up- and downregulated simultaneously in chronically paced patients while there is preserved fatty acid metabolism and impaired vascular signaling.
Unique miR signature seen Based on Duration of Pacing
We next evaluated whether patient demographics and the duration of pacing influenced the circulating miR signature. The circulating miR signature was not influenced by patient age, gender, or race. Interestingly, with longer duration of pacing 14.17 ± 0.7 years (N = 6) vs. 8.49 ± 0.6 years (N = 3) (p = 0.001), 194 miRs were upregulated and 97 miRs were downregulated (Fig. 3). Age at the time of sample collection was similar between the two groups (15.6 ± 1.1 vs. 15.9 ± 1.5 years, p = 0.87). The longer duration of pacing group had pacemakers placed between 0 and 1 year of age (with one exception), while the < 10 year paced group had pacemakers placed at > 5 years of age. Upregulated miRs predicted downregulation of target pathways involved in focal adhesion via cadherins and protocadherins (miR-548, miR-449c-5p), protein phosphorylation, ATP binding, kinase activity (miR-758, miR-183), and protein quality control via protein ubiquitination, conjugation, and ligase activity (miR-513a, 548, 514b) (Table 5). Downregulated miRs predicted upregulation of target pathways involved in extracellular matrix (miR-195-5p, 374a-5p, 942-5p), inflammation (miR-126-5p, miR-199a, miR-182-5p), and zinc finger protein family of transcription factors (miR-181a, miR-26a, miR-148b, miR-15b, miR-374a) (Table 5). Predicted target proteins of select dysregulated miRs are shown in Fig. 4. Among the patients with longer duration of pacing, one patient died following a ventricular fibrillation induced cardiac arrest. Clinical characteristics of this patient were similar to the remainder of the chronically paced group at the time of sample collection and at 3-year follow-up. However, the miR signature differed. Upregulated miRs predicted downregulation of pathways involved in myocardial sodium and calcium channel homeostasis (let-7), as well as pathways important for maintaining vascular endothelial health and preventing fibrosis (miR-92a, miR-130). Downregulated miRs predicted upregulation of profibrotic signaling (miR-29, miR-27) (Fig. 5).
miR expression in paced patients by duration of pacing. a Violin plot of normalized signal intensity in miR expression among all the paced samples demonstrates normal distribution. b, c Volcano plot and heatmap showing significantly up- (red) and downregulated miRs (blue) in patients paced > 10 years vs. patients paced < 10 years by > twofold change in expression and corrected FDR p < 0.05. miR, microRNA
Target proteins of select dysregulated miRs secondary to chronic pacing. Bioinformatically predicted target genes of dysregulated miRs secondary to chronic pacing were mapped using Cytoscape version 3.8.2. Downregulations of miR-195-5p and miR-26a-5p predict upregulation in extracellular matrix proteins; downregulation in miR-183-5p predicts upregulation in protein phosphatases and integrins; downregulation in miR-182-5p predicts upregulation in vascular inflammation via HDAC9 and MITF; and downregulation in miR-126-5p predicts upregulation in cell–cell interaction via ADAM9 and extracellular matrix via MMP7 and THBS1. Upregulation of miR-373-3p predicts downregulation of target genes involved in DNA repair and synthesis of transmembrane receptors. Red, upregulated miR; blue, downregulated miR; green, target proteins
Target proteins of dysregulated miRs in the deceased patient compared to patients paced for < 10 years. Bioinformatically predicted target genes of dysregulated miRs secondary to chronic pacing were mapped using Cytoscape version 3.8.2. Upregulation of let-7 predicts downregulation of proteins involved in myocardial sodium and calcium channel homeostasis. Upregulations of miR-92a and miR-130 predict downregulation of proteins important for maintaining vascular endothelial health and preventing fibrosis. Downregulations of miR-29 and miR-27 predict upregulation of profibrotic signaling. Red, upregulated miR; blue, downregulated miR; green, target proteins
qPCR Validation of miR Microarray Data
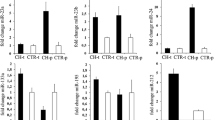
qPCR validation of miR microarray results was performed and confirmed on 3 select upregulated miRs: miR-205-5p, miR-210-5p, and miR-214-3p, and on 5 select downregulated miRs: miR-15b-3p, miR-126-5p, miR-130b-5p, miR-148b-5p, and miR-190a-5p (Fig. 6). These particular miRs were selected for qPCR validation based on (i) their high expression, (ii) being upregulated (miR-205-5p, miR-210-5p, miR-214-3p, Table 3) or downregulated (miR-15b-3p, miR-126-5p, miR-130b-5p, miR-148b-5p, miR-190a-5p, Table 4) with chronic pacing, and (iii) previous studies showing their high expression and roles in regulating key biologic processes in other forms of heart failure [17,18,19,20,21,22,23]. Interestingly, miRs reported as having favorable prognostic value when downregulated in adult heart failure were among the most downregulated in children who were paced for < 10 years compared with children paced for > 10 years, namely miR-15b-5p (FC-7), miR-148b-5p (FC-7), and miR-190a-5p (FC-15). Additional qPCR validation was performed for expression of select miRs which were further dysregulated in the deceased patient when compared to the rest of the chronically paced group. Consistent with the microarray data, there was significant downregulation of miR-29a-3p (FC-2.2) and miR-27a-3p (FC-2.5), as well as upregulation of miR-92a-3p (FC-3.3).
qPCR validation of miRs. Select miRs known to be dysregulated in adult heart failure were validated using qPCR. qPCR expression of the miRs in control vs. paced patients (N = 6/group) correlated with miR expression via microarray. We tested three upregulated miRs: miR-214-3p, miR-210-5p, and miR-205-5p and five downregulated miRs: miR-130b-5p, miR-190a-5p, miR-148b-5p, miR-126-5p, and miR-15b-3p. Data are represented as mean and standard error of mean. Unpaired, Student’s t-test was used where distribution was normal (miR-205, 210, 126, and 148) and Mann–Whitney test was used where distribution was not normal (miR-214, 15, 130, 190). p ≤ 0.05 (✱); p ≤ 0.001 (✱✱). miR, microRNA; FC, fold change
Discussion
Chronic ventricular pacing in children with CCAVB and normal cardiac structure can lead to pacing-induced cardiomyopathy in 5–10% of patients. This pilot study in children with CCAVB evaluated the circulating miR signature to understand the response to chronic ventricular pacing. We identified both adaptive and maladaptive miR signaling at a time when ventricular function is still preserved with a shift toward more maladaptive signaling with longer duration of pacing. Based on the circulating miR dysregulation, fatty acid metabolism and cell survival were upregulated, while nitric oxide signaling and cell–cell and cell–matrix continuity were downregulated and inflammation and oxidative stress were upregulated. We also show a subset of dysregulated miRs which could be implicated in the development of an arrhythmogenic substrate with chronic pacing.
Similar to other etiologies of heart failure, impaired fatty acid oxidation has been reported in animal models of pacing-induced cardiomyopathy [24,25,26,27,28]. In contrast, we observed downregulation of miR-195 which predicts heightened fatty acid biosynthesis, highlighting the ongoing ability to meet the increased demands of the stressed heart. A transition to upregulation of miR-195 may predict the development of heart failure and could be used as a tool to follow disease progression [29]. Adaptive signaling was also predicted by preservation of pathways involving crosstalk between ECM and transmembrane proteins, as well as maintenance of structural integrity, and cell survival and function [30].
Maladaptive miR signaling associated with chronic ventricular pacing was characterized by an upregulation in the miR-548 family of miRs, which are known to inhibit cell–cell and cell–matrix continuity as well as protein quality control, and may therefore induce electromechanical uncoupling. Chronic right ventricular pacing is known to cause electromechanical uncoupling [31,32,33,34], which over time may lead to ventricular dysfunction and arrhythmias [3, 35]. While heightened oxidative stress and inflammation have been demonstrated in animal models of chronic ventricular pacing [36, 37], we show here for the first time that this adverse myocardial remodeling may be reflected in the circulating miR signature, thereby raising the possibility of its utility in following disease progression noninvasively. [36, 38]. The miR signature in our chronically paced patients also implicates impaired proteosomal-ubiquitin systems which maintain protein quality and prevent accumulation of damaged proteins such as quality control of gap junction proteins [39,40,41].
It is unclear whether the duration of pacing is an independent risk factor for the development of PICM. Long-term follow-up studies suggest that chronic pacing can result in significant histopathological changes but the overall incidence of LV dysfunction remains quite low over time [3, 42,43,44]. We show a miR signature predicting ongoing maladaptive signaling and reduced adaptive signaling with longer duration of pacing despite preserved ventricular function. Downregulations of miR15b-3p and miR 190a-5p have been associated with preservation of cardiac function in adult patients with heart disease [17, 18]. Our data is consistent with this. However, patients paced > 10 years showed an upregulation in these miRs compared to those paced < 10 years suggesting a preclinical shift away from preserved ventricular function. Patients paced for > 10 years also showed downregulation of miR-126 supporting vascular endothelial dysfunction and heightened inflammation with longer pacing duration [19]. Interestingly, one patient who was paced for > 10 years had a miR profile predicting dysregulation of myocardial sodium and calcium channels (let-7), endothelial dysfunction (miR-92a), and fibrosis (miR-27, miR-29, miR-130). This arrhymogenic and profibrotic substrate was out of proportion to the remainder of the chronically paced group. This patient experienced an unexplained terminal ventricular arrhythmia 3 years later. These findings are consistent with studies demonstrating that fibrosis and myocardial ion channel dysregulation provide a mechanistic substrate for ventricular arrhythmias across multiple etiologies of heart failure [45, 46]. Taken together, these findings suggest that the let-7 family as well as miRs-92a, 130, 27 and 29, which have shown promise as candidate biomarkers in other forms of heart failure, may also be helpful for predicting morbidity in chronically paced children [47,48,49,50,51].
The present study does not elucidate whether the circulating miR signature we identified in paced patients with CCAVB originates from the stressed myocardium itself, affected target organs, or represents a larger systemic response to myocardial stress. Additionally, we cannot eliminate the possibility that this miR signature could reflect other sources of myocardial stress such as the initial immunologic insult caused by anti-SSA/SSB antibodies—the cause of CCAVB in our study population. However, studies of anti-SSA/SSB induced cardiomyopathy suggest that patients generally present earlier in life and it is additionally characterized by endomyocardial fibroelastosis and decline in cardiac function which was not seen in our patients [52, 53]. In addition, while we show enhanced miR dysregulation with longer pacing duration, we cannot elimitate the potential influences of age at the time of pacemaker placement, gender, or ethnicity on miR regulation. Lastly, it is not yet known whether the adverse remodeling pathways identified in this study are those which ultimately cause PICM.
Conclusions
In summary, we identified a unique, noninvasive, circulating miR signature in chronically paced children with normal function predicting a balance of adaptative and maladaptive signaling which may be important for preserving ventricular function. Longer duration of pacing (> 10 years) was associated with ongoing maladaptive signaling as well as a dampening of adaptive signaling when compared to those with shorter pacing duration (< 10 years). Given that ventricular function was maintained in both subgroups, this difference could signify that ongoing remodeling is occurring but is not yet sufficient to cause measurable changes in ventricular function. miR-15b, miR-126, and miR-130, which were significantly dysregulated with longer pacing duration in the present study and known to be dysregulated in other forms of heart failure, may be good candidates for following this disease progression. Finally, miR-29a, miR-27a, and miR-92a, which predicted enhanced ion channel dysregulation and fibrosis and were further dysregulated in the chronically paced patient who suffered a terminal ventricular arrhythmia, may be important for predicting risk for later morbidity.
Data Availability
The data generated and analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- PICM:
-
Pacing-induced cardiomyopathy
- miR(s):
-
MicroRNA(s)
- CCAVB:
-
Congenital complete atrioventricular block
- LVEF:
-
Left ventricular ejection fraction
- LVEDS:
-
Left ventricular end diastolic dimension in systole
- LVEDD:
-
Left ventricular end diastolic dimension in diastole
- RV FAC:
-
Right ventricular fractional area change
- BSA:
-
Body surface area
- FC:
-
Fold change
- ECM:
-
Extracellular matrix
- PBMCs:
-
Peripheral blood mononuclear cells
References
Michaëlsson M, Engle MA. Congenital complete heart block: an international study of the natural history. Cardiovasc Clin. 1972;4:85–101.
van Geldorp IE, Vanagt WY, Prinzen FW, Delhaas T. Chronic ventricular pacing in children: toward prevention of pacing-induced heart disease. Heart Fail Rev. 2011;16:305–14. https://doi.org/10.1007/s10741-010-9207-1.
Karpawich PP, Rabah R, Haas JE. Altered cardiac histology following apical right ventricular pacing in patients with congenital atrioventricular block. Pacing Clin Electrophysiol. 1999;22:1372–7. https://doi.org/10.1111/j.1540-8159.1999.tb00631.x.
Thambo J-B, Bordachar P, Garrigue S, Lafitte S, Sanders P, Reuter S, et al. Detrimental ventricular remodeling in patients with congenital complete heart block and chronic right ventricular apical pacing. Circulation. 2004;110:3766–72. https://doi.org/10.1161/01.CIR.0000150336.86033.8D.
Horigome H. Dilated cardiomyopathy in children with isolated congenital complete atrioventricular block. Circ J. 2016;80:1110–2. https://doi.org/10.1253/circj.CJ-16-0284.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97. https://doi.org/10.1016/s0092-8674(04)00045-5.
Gholaminejad A, Zare N, Dana N, Shafie D, Mani A, Javanmard SH. A meta-analysis of microRNA expression profiling studies in heart failure. Heart Fail Rev. 2021;26:997–1021. https://doi.org/10.1007/s10741-020-10071-9.
Vogel B, Keller A, Frese KS, Leidinger P, Sedaghat-Hamedani F, Kayvanpour E, et al. Multivariate miRNA signatures as biomarkers for non-ischaemic systolic heart failure. Eur Heart J. 2013;34:2812–23. https://doi.org/10.1093/eurheartj/eht256.
Chen F, Yang J, Li Y, Wang H. Circulating microRNAs as novel biomarkers for heart failure. Hellenic J Cardiol. 2018;59:209–14. https://doi.org/10.1016/j.hjc.2017.10.002.
Abu-Halima M, Meese E, Keller A, Abdul-Khaliq H, Rädle-Hurst T. Analysis of circulating microRNAs in patients with repaired Tetralogy of Fallot with and without heart failure. J Transl Med. 2017;15:156. https://doi.org/10.1186/s12967-017-1255-z.
Weldy CS, Syed SA, Amsallem M, Hu D-Q, Ji X, Punn R, et al. Circulating whole genome miRNA expression corresponds to progressive right ventricle enlargement and systolic dysfunction in adults with tetralogy of Fallot. PLoS One. 2020;15:e0241476. https://doi.org/10.1371/journal.pone.0241476.
Marfella R, Di Filippo C, Potenza N, Sardu C, Rizzo MR, Siniscalchi M, et al. Circulating microRNA changes in heart failure patients treated with cardiac resynchronization therapy: responders vs. non-responders. Eur J Heart Fail. 2013;15:1277–88. https://doi.org/10.1093/eurjhf/hft088.
Mompeón A, Ortega-Paz L, Vidal-Gómez X, Costa TJ, Pérez-Cremades D, Garcia-Blas S, et al. Disparate miRNA expression in serum and plasma of patients with acute myocardial infarction: a systematic and paired comparative analysis. Sci Rep. 2020;10:5373. https://doi.org/10.1038/s41598-020-61507-z.
Wang K, Yuan Y, Cho J-H, McClarty S, Baxter D, Galas DJ. Comparing the microRNA spectrum between serum and plasma Ahuja SK editor. PLoS One. 2012;7:e41561. https://doi.org/10.1371/journal.pone.0041561.
Glinge C, Clauss S, Boddum K, Jabbari R, Jabbari J, Risgaard B, et al. Stability of circulating blood-based microRNAs — pre-analytic methodological considerations Calin G editor. PLoS One. 2017;12:e0167969. https://doi.org/10.1371/journal.pone.0167969.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C(T)) method. Methods. 2001;25:402–8. https://doi.org/10.1006/meth.2001.1262.
Wong LL, Armugam A, Sepramaniam S, Karolina DS, Lim KY, Lim JY, et al. Circulating microRNAs in heart failure with reduced and preserved left ventricular ejection fraction. Eur J Heart Fail. 2015;17:393–404. https://doi.org/10.1002/ejhf.223.
Porrello ER, Mahmoud AI, Simpson E, Johnson BA, Grinsfelder D, Canseco D, et al. Regulation of neonatal and adult mammalian heart regeneration by the miR-15 family. PNAS. 2013;110:187–92. https://doi.org/10.1073/pnas.1208863110.
Harris TA, Yamakuchi M, Ferlito M, Mendell JT, Lowenstein CJ. MicroRNA-126 regulates endothelial expression of vascular cell adhesion molecule 1. Proc Natl Acad Sci U S A. 2008;105:1516–21. https://doi.org/10.1073/pnas.0707493105.
Yang K, Shi J, Hu Z, Hu X. The deficiency of miR-214-3p exacerbates cardiac fibrosis via miR-214-3p/NLRC5 axis. Clin Sci. 2019;133:1845–56. https://doi.org/10.1042/CS20190203.
Guan Y, Song X, Sun W, Wang Y, Liu B. Effect of hypoxia-induced microRNA-210 expression on cardiovascular disease and the underlying mechanism. Oxid Med Cell Longev. 2019;2019:e4727283. https://doi.org/10.1155/2019/4727283.
Lkhagva B, Lin Y-K, Kao Y-H, Chazo T-F, Chung C-C, Chen S-A, et al. Novel histone deacetylase inhibitor modulates cardiac peroxisome proliferator-activated receptors and inflammatory cytokines in heart failure. Pharmacol. 2015;96:184–91. https://doi.org/10.1159/000438864.
Du Y, Ma X, Ma L, Li S, Zheng J, Lv J, et al. Inhibition of microRNA-148b-3p alleviates oxygen-glucose deprivation/reoxygenation-induced apoptosis and oxidative stress in HT22 hippocampal neuron via reinforcing Sestrin2/Nrf2 signalling. Clin Exp Pharmacol Physiol. 2020;47:561–70. https://doi.org/10.1111/1440-1681.13231.
Neubauer S. The failing heart — an engine out of fuel. N Engl J Med. 2007;356:1140–51. https://doi.org/10.1056/NEJMra063052.
Taegtmeyer H. Genetics of energetics: transcriptional responses in cardiac metabolism. Ann Biomed Eng. 2000;28:871–6. https://doi.org/10.1114/1.1312187.
Stanley WC, Recchia FA, Lopaschuk GD. Myocardial substrate metabolism in the normal and failing heart. Physiol Rev. 2005;85:1093–129. https://doi.org/10.1152/physrev.00006.2004.
Osorio JC, Stanley WC, Linke A, Castellari M, Diep QN, Panchal AR, et al. Impaired myocardial fatty acid oxidation and reduced protein expression of retinoid X receptor-α in pacing-induced heart failure. Circulation. 2002;106:606–12. https://doi.org/10.1161/01.CIR.0000023531.22727.C1.
Zhang X, Ji R, Liao X, Castillero E, Kennel PJ, Brunjes DL, et al. MicroRNA-195 regulates metabolism in failing myocardium via alterations in sirtuin 3 expression and mitochondrial protein acetylation. Circulation. 2018;137:2052–67. https://doi.org/10.1161/CIRCULATIONAHA.117.030486.
Kenny HC, Abel ED. Heart failure in type 2 diabetes mellitus. Circ Res. 2019;124:121–41. https://doi.org/10.1161/CIRCRESAHA.118.311371.
Song R, Zhang L. Cardiac ECM: its epigenetic regulation and role in heart development and repair. Int J Mol Sci. 2020;21:E8610. https://doi.org/10.3390/ijms21228610.
van Oosterhout MF, Prinzen FW, Arts T, Schreuder JJ, Vanagt WY, Cleutjens JP, et al. Asynchronous electrical activation induces asymmetrical hypertrophy of the left ventricular wall. Circulation. 1998;98:588–95. https://doi.org/10.1161/01.cir.98.6.588.
Prinzen FW, Hunter WC, Wyman BT, McVeigh ER. Mapping of regional myocardial strain and work during ventricular pacing: experimental study using magnetic resonance imaging tagging. J Am Coll Cardiol. 1999;33:1735–42. https://doi.org/10.1016/s0735-1097(99)00068-6.
Tse HF, Lau CP. Long-term effect of right ventricular pacing on myocardial perfusion and function. J Am Coll Cardiol. 1997;29:744–9. https://doi.org/10.1016/s0735-1097(96)00586-4.
Skalidis EI, Kochiadakis GE, Koukouraki SI, Chrysostomakis SI, Igoumenidis NE, Karkavitsas NS, et al. Myocardial perfusion in patients with permanent ventricular pacing and normal coronary arteries. J Am Coll Cardiol. 2001;37:124–9. https://doi.org/10.1016/s0735-1097(00)01096-2.
Vernooy K, Dijkman B, Cheriex EC, Prinzen FW, Crijns HJGM. Ventricular remodeling during long-term right ventricular pacing following His bundle ablation. Am J Cardiol. 2006;97:1223–7. https://doi.org/10.1016/j.amjcard.2005.11.044.
Cesselli D, Jakoniuk I, Barlucchi L, Beltrami AP, Hintze TH, Nadal-Ginard B, et al. Oxidative stress-mediated cardiac cell death is a major determinant of ventricular dysfunction and failure in dog dilated cardiomyopathy. Circ Res. 2001;89:279–86. https://doi.org/10.1161/hh1501.094115.
Sengupta A, Molkentin JD, Paik J-H, DePinho RA, Yutzey KE. FoxO transcription factors promote cardiomyocyte survival upon induction of oxidative stress. J Biol Chem. 2011;286:7468–78. https://doi.org/10.1074/jbc.M110.179242.
Marín-García J, Goldenthal MJ, Moe GW. Abnormal cardiac and skeletal muscle mitochondrial function in pacing-induced cardiac failure. Cardiovasc Res. 2001;52:103–10. https://doi.org/10.1016/S0008-6363(01)00368-6.
Glickman MH, Ciechanover A. The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol Rev. 2002;82:373–428. https://doi.org/10.1152/physrev.00027.2001.
Ciechanover A. The ubiquitin proteolytic system: from a vague idea, through basic mechanisms, and onto human diseases and drug targeting. Neurol. 2006;66:S7-19. https://doi.org/10.1212/01.wnl.0000192261.02023.b8.
Laing JG, Tadros PN, Green K, Saffitz JE, Beyer EC. Proteolysis of connexin43-containing gap junctions in normal and heat-stressed cardiac myocytes. Cardiovasc Res. 1998;38:711–8. https://doi.org/10.1016/s0008-6363(98)00060-1.
Kim JJ, Friedman RA, Eidem BW, Cannon BC, Arora G, Smith EO, et al. Ventricular function and long-term pacing in children with congenital complete atrioventricular block. J Cardiovasc Electrophysiol. 2007;18:373–7. https://doi.org/10.1111/j.1540-8167.2006.00741.x.
Udink ten Cate FEA, Breur JMPJ, Cohen MI, Boramanand N, Kapusta L, Crosson JE, et al. Dilated cardiomyopathy in isolated congenital complete atrioventricular block: early and long-term risk in children. J Am College Cardiol. 2001;37:1129–34. https://doi.org/10.1016/S0735-1097(00)01209-2.
Pordon CM, Moodie DS. Adults with congenital complete heart block: 25-year follow-up. Cleve Clin J Med. 1992;59:587–90. https://doi.org/10.3949/ccjm.59.6.587.
Rahm A-K, Lugenbiel P, Schweizer PA, Katus HA, Thomas D. Role of ion channels in heart failure and channelopathies. Biophys Rev. 2018;10:1097–106. https://doi.org/10.1007/s12551-018-0442-3.
Luo X, Zhang H, Xiao J, Wang Z. Regulation of human cardiac ion channel genes by microRNAs: theoretical perspective and pathophysiological implications. Cell Physiol Biochem. 2010;25:571–86. https://doi.org/10.1159/000315076.
Bao M-H, Feng X, Zhang Y-W, Lou X-Y, Cheng Y, Zhou H-H. Let-7 in cardiovascular diseases, heart development and cardiovascular differentiation from stem cells. Int J Mol Sci. 2013;14:23086–102. https://doi.org/10.3390/ijms141123086.
Wang W, Li Z, Zheng Y, Yan M, Cui Y, Jiang J. Circulating microRNA-92a level predicts acute coronary syndrome in diabetic patients with coronary heart disease. Lipids Health Dis. 2019;18:22. https://doi.org/10.1186/s12944-019-0964-0.
Chiti E, Paolo MD, Turillazzi E, Rocchi A. MicroRNAs in hypertrophic, arrhythmogenic and dilated cardiomyopathy. Diagnostics (Basel). 2021;11:1720. https://doi.org/10.3390/diagnostics11091720.
Liu M-N, Luo G, Gao W-J, Yang S-J, Zhou H. miR-29 family: a potential therapeutic target for cardiovascular disease. Pharmacol Res. 2021;166:105510. https://doi.org/10.1016/j.phrs.2021.105510.
Teng L, Huang Y, Guo J, Li B, Lin J, Ma L, et al. Cardiac fibroblast miR-27a may function as an endogenous anti-fibrotic by negatively regulating early growth response protein 3 (EGR3). J Cell Mol Med. 2021;25:73–83. https://doi.org/10.1111/jcmm.15814.
Nield LE, Silverman ED, Taylor GP, Smallhorn JF, Mullen JBM, Silverman NH, et al. Maternal anti-Ro and anti-La antibody-associated endocardial fibroelastosis. Circulation. 2002;105:843–8. https://doi.org/10.1161/hc0702.104182.
Moak JP, Barron KS, Hougen TJ, Wiles HB, Balaji S, Sreeram N, et al. Congenital heart block: development of late-onset cardiomyopathy, a previously underappreciated sequela. J Am Coll Cardiol. 2001;37:238–42. https://doi.org/10.1016/S0735-1097(00)01048-2.
Acknowledgements
We thank our human subjects research coordinators Sara Sherman-Levine and Aihua Zhu, supported by NIH/National Center for Advancing Translational Sciences/Clinical and Translational Science Awards grant UL1 TR001085, the Lucile Packard Foundation for Children’s Health, and the Child Health Research Institute. We thank our human subjects research coordinator Sandra Moon from the Lucile Packard Heart Center Translational Research Program.
Funding
This work was supported by the National Institutes of Health [K08 HL127277-01 NIH/NHLBI to SR]; Reddy Foundation grant to SR; American Heart Association Grant-in-Aid to SR; NIH/NHLBI F32HL160067 to CW; and NIH/NHLBI L30HL159413 to CW.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by BMN, KLC, SR, and XJ. The first draft of the manuscript was written by BMN and KLC, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
All procedures involving human participants in this study were performed in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. This study was approved by the Stanford University Institutional Review Board and consent was obtained for all patients and assent was obtained for those > 7 years of age. No animal studies were carried out by the authors for this article.
Conflict of Interest
The authors declare no competing interests.
Additional information
Associate Editor Junjie Xiao oversaw the review of this article
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Navarre, B.M., Clouthier, K.L., Ji, X. et al. miR Profile of Chronic Right Ventricular Pacing: a Pilot Study in Children with Congenital Complete Atrioventricular Block. J. of Cardiovasc. Trans. Res. 16, 287–299 (2023). https://doi.org/10.1007/s12265-022-10318-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12265-022-10318-w