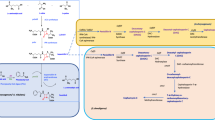
Abstract
Our previous study provided an insight into DoxR as a negative regulator of doxorubicin production in Streptomyces peucetius ATCC 27952. Streptomyces hosts are advantageous in terms of producing a number of pharmaceuticals in low titer. Antisense RNAs (asRNAs) silencing strategy acts as an alternative tool for metabolic engineering of microorganisms for construction of an efficient cell factory. In this study, a paired-termini antisense RNAs (PTasRNAs) silencing strategy was employed for inhibition of DoxR to enhance doxorubicin production. To continue this endeavor, we designed and constructed the piBR702 vector for the expression of PTasRNAs in monocistronic mode. Further, two variants of asRNA, adoxR and bdoxR were designed and cloned into piBR702. All the rDNAs were transformed into S. peucetius to generate engineered strains. The engineered strains, S. peucetius A and S. peucetius B produced enhanced titers of doxorubicin, daunorubicin, and e-rhodomycinone; however, no such change was seen in S. peucetius AB. Moreover, RT-PCR analysis of doxR from S. peucetius A and S. peucetius B, together with the higher production from S. peucetius A, confirmed adoxR as a better asRNA than bdoxR. The reason behind this could be due to the simple secondary structure and low binding free energy of adoxR (−419 kcal/mol) than bdoxR (−358.8 kcal/mol). Our study demonstrated that antibiotic production was enhanced significantly by inhibiting DoxR, a negative regulator in S. peucetius using PTasRNAs. In addition, this study further provides an insight into PTasRNAs as an effective tool for gene silencing in Streptomyces and its use as an effective tool for metabolic engineering.
Similar content being viewed by others
References
Ahn, J. H., K. H. Lee, J. W. Shim, E. Y. Lee, and D. M. Kim (2013) Streamlined cell-free protein synthesis from sequence information. Biotechnol. Bioproc. Eng. 18: 1101–1108.
Borodina, I. and J. Nielsen (2014) Advances in metabolic engineering of yeast Saccharomyces cerevisiae for production of chemicals. Biotechnol. J. 9: 609–620.
Chaudhary, A. K., J. W. Park, Y. J. Yoon, B. G. Kim, and J. K. Sohng (2013a) Re-engineering of genetic circuit for 2-deoxystreptamine (2-DOS) biosynthesis in Escherichia coli BL21 (DE3). Biotechnol. Lett. 35: 285–293.
Li, Y. and B. A. Pfeifer (2014) Heterologous production of plantderived isoprenoid products in microbes and the application of metabolic engineering and synthetic biology. Curr. Opin. Plant Biol. 19: 8–13.
Na, D., S. M. Yoo, H. Chung, H. Park, J. H. Park, and S. Y. Lee (2013) Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs. Nat. Biotechnol. 31: 170–174.
Ro, D. K., E. M. Paradise, M. Ouellet, K. J. Fisher, K. L. Newman, J. M. Ndungu, K. A. Ho, R. A. Eachus, T. S. Ham, J. Kirby, M. C. Chang, S. T. Withers, Y. Shiba, R. Sarpong, and J. D. Keasling (2006) Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature 440: 940–943.
Xu, P., A. Vansiri, N. Bhan, and M. A. Koffas (2012) ePathBrick: A synthetic biology platform for engineering metabolic pathways in Escherichia coli. ACS. Synth. Biol. 1: 256–266.
Medema, M. H., R. Breitling, and E. Takano (2011) Synthetic biology in Streptomyces bacteria. Methods Enzymol. 497: 485–502.
Herai, S., Y. Hashimoto, H. Higashibata, H. Maseda, H. Ikeda, S. Omura, and M. Kobayashi (2004) Hyper-inducible expression system for streptomycetes. Proc. Natl. Acad. Sci. USA. 101: 14031–14035.
Luo, Y., H. Huang, J. Liang, M. Wang, L. Lu, Z. Shao, R. E. Cobb, and H. Zhao (2013) Activation and characterization of a cryptic polycyclic tetramate macrolactam biosynthetic gene cluster. Nat. Commun. 4: 2894.
Sugimoto, Y., L. Ding, K. Ishida, and C. Hertweck (2014) Rational design of modular polyketide synthases: morphing the aureothin pathway into a luteoreticulin assembly line. Angew. Chem. Int. Ed. Engl. 53: 1560–1564.
Uguru, G. C., M. Mondhe, S. Goh, A. Hesketh, M. J. Bibb, L. Good, and J. E. Stach (2013) Synthetic RNA silencing of actinorhodin biosynthesis in Streptomyces coelicolor A3(2). PLoS One 8: 1–9.
Zhuo, Y., T. Zhang, Q. Wang, P. Cruz-Morales, B. Zhang, M. Liu, F. Barona-Gómez, and L. Zhang (2014) Synthetic biology of avermectin for production improvement and structure diversification. Biotechnol. J. 9: 316–325.
Chaudhary, A. K., D. Dhakal, and J. K. Sohng (2013b) An insight into the “-omics” based engineering of streptomycetes for secondary metabolite overproduction. Biomed. Res. Int. 2013: 968518.
Liang, J. C., R. J. Bloom, and C. D. Smolke (2011) Engineering biological systems with synthetic RNA molecules. Mol. Cells 43: 915–926.
Lucks, J. B., L. Qi, V. K. Mutalik, D. Wang, and A. P. Arkin (2011) Versatile RNA-sensing transcriptional regulators for engineering genetic networks. Proc. Natl. Acad. Sci. USA. 108: 8617–8622.
Sharma, V., A. Yamamura, and Y. Yokobayashi (2012) Engineering artificial small RNAs for conditional gene silencing in Escherichia coli. ACS. Synth. Biol. 1: 6–13.
Deshmukh, R. and H. J. Purohit (2014) siRNA mediated gene silencing in Fusarium sp. HKF15 for overproduction of bikaverin. Bioresour. Technol. 157: 368–371.
Liu, Y., Y. Zhu, J. Li, H.D. Shin, R. R. Chen, G. Du, L. Liu, and J. Chen (2014) Modular pathway engineering of Bacillus subtilis for improved N-acetylglucosamine production. Metab. Eng. 23: 42–52.
Jha, A. K., A. R. Anaya, A. K. Chaudhary, S. W. Park, W. J. Cho, and J. K. Sohng (2014) Metabolic engineering of rational screened Saccharopolyspora spinosa for the enhancement of spinosyns A and D Production. Mol. Cells 37: 727–733.
Yang, J., J. Li, Y. Hu, L. Li, L. Long, F. Wang, and S. Zhang (2015) Characterization of a thermophilic hemoglobin-degrading protease from Streptomyces rutgersensis SCSIO 11720 and its application in antibacterial peptides production. Biotechnol. Bioproc. Eng. 20: 79–90.
D'Alia, D., K. Nieselt, S. Steigele, J. Müller, I. Verburg, and E. Takano (2010) Noncoding RNA of glutamine synthetase I modulates antibiotic production in Streptomyces Coelicolor A3(2). J. Bacteriol. 192: 1160–1164.
Pánek, J., J. Bobek, K. Mikulík, M. Basler, and J. Vohradský (2008) Biocomputational prediction of small non-coding RNAs in Streptomyces. BMC. Genomics 9: 1–11
Swiercz, J. P., Hindra, J. Bobek, H. J. Haiser, C. D. Berardo, B. Tjaden, and M. A. Elliot (2008) Small non-coding RNAs in Streptomyces coelicolor. Nucleic Acids Res. 36: 7240–7251.
Tezuka, T., H. Hara, Y. Ohnishi, and S. Horinouchi (2009) Identification and gene disruption of small noncoding RNAs in Streptomyces griseus. J. Bacteriol. 191: 4896–4904.
Vockenhuber, M. P., C. M. Sharma, M. G. Statt, D. Schmidt, Z. Xu, S. Dietrich, H. Liesegang, D. H. Mathews, and B. Suess (2011) Deep sequencing-based identification of small non-coding RNAs in Streptomyces coelicolor. RNA. Biol. 8: 468–477.
Hindra., P. Pak, and M. A. Elliot (2010) Regulation of a novel gene cluster involved in secondary metabolite production in Streptomyces coelicolor. J. Bacteriol. 192: 4973–4982.
Waters, L. S. and G. Storz (2009) Regulatory RNAs in bacteria. Cell 136: 615–628.
Romero, N. M. and R. P. Mellado (1995) Activation of the actinorhodin biosynthetic pathway in Streptomyces lividans. FEMS Microbiol. Lett. 127: 79–84.
Romero, N. M., V. Parro, F. Malpartida, and R. P. Mellado (1992) Heterologous activation of the actinorhodin biosynthetic pathway in Streptomyces lividans. Nucleic Acids Res. 20: 2767–2772.
Chaudhary, A. K., B. Singh, S. Maharjan, A. K. Jha, B. G. Kim, and J. K. Sohng (2014) Switching antibiotics production on and off in Actinomycetes by an IclR family transcriptional regulator from Streptomyces peucetius ATCC27952. J. Microbiol. Biotechnol. 24: 1065–1072.
Kieser, T., M. J. Bibb, M. J. Buttner, K. F. Chater, and D. A. Hopwood (2000) Practical Streptomyces Genetics. John Innes Foundation, Norwich, UK.
Sambrook, J., E. F. Fritsch, and T. Maniatis (2001) Molecular cloning: A laboratory manual. 3rd ed., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, USA.
Vara, J., M. Lewandowska-Skarbek, Y. G. S. Wang, Donadio, and C. R. Hutchinson (1989) Cloning of genes governing the deoxysugar portion of the erythromycin biosynthesis pathway in Saccharopolyspora erythraea (Streptomyces erythreus). J. Bacteriol. 171: 5872–5881.
Sthapit. B., T. J. Oh, R. Lamichhane, K. Liou, H. C. Lee, C. G. Kim, and J. K. Sohng (2004) Neocarzinostatin naphthoate synthase: An unique iterative type I PKS from neocarzinostatin producer Streptomyces carzinostaticus. FEBS. Lett. 566: 201–206.
Bibb, M. J., G. R. Janssen, and J. M. Ward (1985) Cloning and analysis of the promoter region of the erythromycin resistance gene (ermE*) of Streptomyces erythraeus. Gene 38: 215–226.
Pulido, D. and A. Jiménez (1987) Optimization of gene expression in Streptomyces lividans by a transcription terminator. Nucleic Acids Res. 15: 4227–4240.
Nepal, K. K., T. J. Oh, B. Subba, J. C. Yoo, and J. K. Sohng (2008) Characterization of RbmD through neomycin production reconstituted from the engineered Streptomyces fradiae BS1. Mol. Cells 27: 83–88.
Singh, B., C. B. Lee, and J. K. Sohng (2010) Precursor for biosynthesis of sugar moiety of doxorubicin depends on rhamnose biosynthetic pathway in Streptomyces peucetius ATCC27952. Appl. Microbiol. Biotechnol. 85: 1565–1574.
Li, M., J. Wang, Y. Geng, Y. Li, Q. Wang, Q. Liang, and Q. Qi (2012) A strategy of gene overexpression based on tandem repetitive promoters in Escherichia coli. Microb. Cell Fact. 11: 1–10.
Nakashima, N., T. Tamura, and L. Good (2006) Paired-termini stabilize antisense RNAs and enhance conditional gene silencing in Escherichia coli. Nucleic Acids Res. 34: e138.
Nakashima, N. and T. Tamura (2009) Conditional gene silencing of multiple genes with antisense RNAs and generation of a mutator strain of Escherichia coli. Nucleic Acids Res. 37: 1–11.
Hofacker, I. L. (2003) Vienna RNA secondary structure server. Nucleic Acids Res. 31: 3429–3431.
Malla, S., N. P. Niraula, K. Liou, and J. K. Sohng (2009) Selfresistance mechanism in Streptomyces peucetius: Overexpression of drrA, drrB and drrC for doxorubicin enhancement. Microbiol. Res. 165: 259–267.
Aiba, H. (2007) Mechanism of RNA silencing by Hfq-binding small RNAs. Curr. Opin. Microbiol. 10: 134–139.
Nakashima, N., S. Goh, L. Good, and T. Tamura (2012) Multiple-gene silencing using antisense RNAs in Escherichia coli. Methods Mol. Biol. 815: 307–319.
Arcamone, F., G. Cassinelli, G. Fantini, A. Grein, P. Orezzi, C. Pol, and C. Spalla (1969) 14-Hydroxydaunomycin, a new antitumor antibiotic from Streptomyces peucetius var. caesius. Biotechnol. Bioeng. 11: 1101–1110.
Kapranov, P., A. T. Willingham, and T. R. Gingeras (2007) Genomewide transcription and the implications for genomic organization. Nat. Rev. Genet. 8: 413–423.
Tjaden, B., S. Sarah, Goodwin, A. Jason, Opdyke, A. Jason G. Maude, X. F. Daniel, G. Susan, and S. Gisela (2006) Target prediction for small, noncoding RNAs in bacteria. Nucleic Acids Res. 34: 2791–2802.
Georg, J. and W. R. Hess (2011) cis-Antisense RNA, another level of gene regulation in bacteria. Microbiol. Mol. Biol. Rev. 75: 286–300.
Thomason, M. K. and G. Storz (2010). Bacterial antisense RNAs: How many are there, and what are they doing? Annu. Rev. Genet. 44: 167–188.
Desai, R. P. and E. T. Papoutsakis (1999) Antisense RNA strategies for metabolic engineering of Clostridium acetobutylicum. Appl. Environ. Microbiol. 65: 936–945.
Hebert, C. G., J. J. Valdes, and W. E. Bentley (2008) Beyond silencing-engineering applications of RNA interference and antisense technology for altering cellular phenotype. Curr. Opin. Biotechnol. 19: 500–505.
Yoo, S. M., D. Na, and S. Y. Lee (2013) Design and use of synthetic regulatory small RNAs to control gene expression in Escherichia coli. Nat. Protoc. 8: 1694–1707.
Chaudhary, A. K., D. Na, and E. Y. Lee (2015). Rapid and highthroughput construction of microbial cell-factories with regulatory noncoding RNAs. Biotechnol. Adv. doi number: 10.1016/ j.biotechadv.2015.05.009.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chaudhary, A.K., Pokhrel, A.R., Hue, N.T. et al. Paired-termini antisense RNA mediated inhibition of DoxR in Streptomyces peucetius ATCC 27952. Biotechnol Bioproc E 20, 381–388 (2015). https://doi.org/10.1007/s12257-014-0810-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-014-0810-1