Abstract
PTEN is an important tumor suppressor gene that antagonizes the oncogenic PI3K/AKT signaling pathway and has functions in the nucleus for maintaining genome integrity. Although PTEN inactivation by mutation is infrequent in breast cancer, transcript and protein levels are deficient in >25 % of cases. The E3 ubiquitin ligase NEDD4 (also known as NEDD4-1) has been reported to negatively regulate PTEN protein levels through poly-ubiquitination and proteolysis in carcinomas of the prostate, lung, and bladder, but its effect on PTEN in the breast has not been studied extensively. To investigate whether NEDD4 contributes to low PTEN levels in human breast cancer, we analyzed the expression of these proteins by immunohistochemistry across a large Swedish cohort of breast tumor specimens, and their transcript expression levels by microarrays. For both NEDD4 and PTEN, their transcript expression was significantly correlated to their protein expression. However, comparing NEDD4 expression to PTEN expression, either no association or a positive correlation was observed at the protein and transcript levels. This unexpected observation was further corroborated in two independent breast cancer cohorts from The Netherlands Cancer Institute and The Cancer Genome Atlas. Our results suggest that NEDD4 is not responsible for the frequent down-regulation of the PTEN protein in human breast carcinoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
PTEN is a phosphatase that plays an important role in tumor suppression by negatively regulating the oncogenic phosphatidylinositol 3-kinase (PI3K) pathway, as well as through functions in the nucleus that contribute to maintenance of genomic integrity [1]. Germline mutations of PTEN are found in patients with PTEN hamartoma tumor syndrome and are associated with an increased risk for breast, thyroid, and endometrial cancer [2–4]. Moreover, somatic loss-of-function mutations of PTEN are estimated to be present in 30 % of cancer and are found across the entire spectrum of tumor types [5–7]. The PTEN/PI3K pathway is one of the key pathways deregulated in breast cancer. PIK3CA, which encodes the p110-alpha catalytic subunit of PI3K, has activating mutations in one-third of breast tumors, and although mutation rate of PTEN is less than 5 % [8], PTEN expression is found to be greatly diminished in at least 25 % of breast tumors and in near mutual exclusivity to PIK3CA mutation [9, 10]. The mechanisms by which PTEN is down-regulated is poorly delineated in breast cancer, but mutations, copy number loss, rearrangements, epigenetic silencing, as well as post-translational regulation may contribute [9–13]. Of note, PTEN loss is frequent within the poor-prognosis basal-like molecular subtype of breast cancer [13].
Recently, Wang et al. reported that NEDD4 (neural precursor cell expressed, developmentally down-regulated 4, E3 ubiquitin protein ligase; also known as NEDD4-1) is an E3 ubiquitin ligase of PTEN and catalyzes poly-ubiquitination of PTEN in cells leading to proteolysis of the PTEN protein, thereby negatively regulating PTEN abundance [14]. Furthermore, in their analysis of mouse prostate and human bladder cancer samples, high expression of NEDD4 was inversely correlated to PTEN protein levels but not PTEN mRNA levels, suggesting that NEDD4 plays a proto-oncogenic role in tumorigenesis and cancer development via post-translational suppression of PTEN [14]. Negative regulation of PTEN by NEDD4-mediated poly-ubiquitination has since been reported to be involved in several biological and pathological processes, such as axon branching [15, 16], T-cell activation [17], keloid formation [18], and insulin-mediated glucose metabolism [19]. Inverse relationships between the expression of NEDD4 and PTEN have also been observed in human non-small cell lung carcinomas [20] and colon cancer [21].
However, the regulation of PTEN by NEDD4 may be microenvironment and/or cell-type specific. For example, Trotman et al. found that in addition to catalyzing poly-ubiquitination of PTEN, NEDD4 is also responsible for PTEN mono-ubiquitination that leads to PTEN nuclear import and protection from proteasomal degradation, making the role of NEDD4 in regulation of PTEN stability subtle and complex [22]. Moreover, some studies have called into question the interaction between NEDD4 and PTEN. For example, no discernible effect on Pten stability, subcellular localization, or downstream targets was observed in two separate Nedd4 knock-out mouse models [23]. Furthermore, Maddika et al. failed to reproduce the functional interaction between NEDD4 and PTEN, and instead found that WWP2, another E3 ubiquitin ligase within the NEDD4-like protein family, mediated poly-ubiquitination of PTEN [24]. A third group has also failed to demonstrate that PTEN is a substrate of Nedd4, and rather found that PTEN regulated Nedd4 by modulating mTORC1 activity [19]. Lastly, in gastric carcinoma, no relationship was observed between NEDD4 and PTEN expression [25], and in colorectal cancer cell lines and biopsies, NEDD4 modulation and expression level were not associated to the levels of PTEN [26].
NEDD4 and its potential role in PTEN regulation in breast cancer have not been studied. To reveal the pattern of expression of NEDD4 in human breast cancer, and to investigate whether NEDD4-mediated PTEN degradation is a factor that contributes to the frequent loss of PTEN protein, we analyzed NEDD4 and PTEN expression at the protein and mRNA levels in a large cohort of Swedish breast tumors, and verified our findings in two independent breast cancer cohorts from The Netherlands Cancer Institute (NKI) and The Cancer Genome Atlas (TCGA) (Table 1).
Materials and Methods
Breast Cancer Cohorts
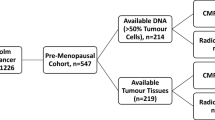
Clinical and demographic information is provided for all cohorts in Table 1. For the Swedish cohort, 132 formalin-fixed paraffin-embedded (FFPE) tissue microarray (TMA) tumor specimens, arrayed in triplicates, were studied for NEDD4 protein expression by IHC, of which 123 had matched PTEN IHC scores previously evaluated [9, 27]. These 123 samples were analyzed for correlation between PTEN and NEDD4 protein levels. Correlation between the PTEN protein and NEDD4 mRNA levels, and correlation between PTEN mRNA and NEDD4 mRNA levels were analyzed in a subset of 105 samples with both PTEN IHC status and microarray gene expression data [27] (NCBI Gene Expression Omnibus accession GSE5325). Correlation between NEDD4 protein and NEDD4 mRNA levels was performed in a subset of 42 samples with NEDD4 IHC and microarray data. For the NKI cohort, gene expression microarray data from 295 tumor samples was analyzed for correlation between gene expression levels of PTEN and NEDD4 [28, 29]. Tissue microarrays containing these 295 NKI cases were stained for PTEN protein, of which 267 samples could be evaluated, and thereafter were analyzed for correlations between PTEN IHC scores and PTEN or NEDD4 mRNA expression levels. For TCGA cohort, level 3 IlluminaHiSeq_RNASeqV2 gene expression data for 970 primary breast tumor samples was used, as well as PTEN protein expression status for 407 cases derived from a reverse phase protein arrays platform. All TCGA data were downloaded from the TCGA data portal (https://tcga-data.nci.nih.gov/tcga/, downloaded on January 20, 2014). The study was approved by the Lund University Hospital ethics committee (LU240-01 and 2009/658), waiving the requirement for informed consent for the study, and all experimental protocols were performed in accordance with approved guidelines.
Immunohistochemistry
The rabbit polyclonal anti-NEDD4 WW2 domain antibody #07–049 (EMD Millipore, Darmstadt, Germany), previously validated to be specific for NEDD4 [14], was used for IHC. The staining was done using an Autostainer Plus instrument and EnVision Plus system (Dako Denmark A/S, Glostrup, Denmark) following manufacturer’s recommended protocol. Antigen retrieval was performed using Dako Targeted Retrieval Buffer pH 6.0 at 98 °C for 20 min, and the primary antibody was used at 1:500 dilution with 30 min incubation time at room temperature. The stained specimens were scanned using a MIRAX MIDI slide scanner (Carl Zeiss AG, Oberkochen, Germany) and viewed with Pannoramic Viewer v1.15.3 (3DHISTECH, Budapest, Hungary). Semi-quantitative scoring was done according to the Dako system 0–3 scoring scale, where scores of 0 were given to tissues with no NEDD4 staining, 1+ to weak NEDD4 staining, 2+ to intermediate NEDD4 staining, and 3+ to strong NEDD4 staining (Fig. 1). The IHC scores of 0 and 1+ were then combined and categorized as NEDD4-negative, and scores of 2+ and 3+ were categorized as NEDD4-positive. PTEN IHC results for the Swedish cohort were reported previously [9, 27]. PTEN IHC was performed on the NKI TMAs using methods previously described [13].
NEDD4 immunohistochemistry. 132 breast tumor tissue microarray specimens were immunohistochemically stained with anti-NEDD4 antibody. Shown are representative examples of tumors with NEDD4 IHC scores of a 0, b 1+, c 2+, and d 3+. Scores 0/1+ were categorized NEDD4-negative, and 2+/3+ as NEDD4-positive. NEDD4 protein was expressed predominantly in the cytoplasm regardless of the staining intensity
Statistical Analysis
The chi-squared test was used to test the significance level of correlations between the NEDD4 protein and different breast cancer biomarkers. The Wilcoxon rank-sum test was used for correlation between PTEN and NEDD4 protein levels. The Student’s t-test was used for correlations between the PTEN/NEDD4 protein and PTEN/NEDD4 mRNA levels. The Pearson’s correlation test was used for correlations between PTEN and NEDD4 mRNA levels from gene expression data and RNA-seq data. All tests were two-tailed, and P < 0.05 was considered significant. All statistical analyses were performed with R version 3.1.0 (http://www.r-project.org).
Results and Discussion
Immunohistochemical (IHC) staining was performed for 132 formalin-fixed paraffin-embedded (FFPE) breast tumor specimens (Swedish cohort) using an antibody previously reported to be specific to NEDD4 [14] (see Methods; Fig. 1). Consistent with previous studies in other tissues [14], NEDD4 protein was predominantly cytoplasmic in breast cancer cells (Fig. 1). Among the 132 stained samples, 60 (45 %) had zero or weak NEDD4 protein staining (classified as NEDD4-negative), whereas 72 (55 %) had intermediate to strong expression (NEDD4-positive). NEDD4 protein expression was positively correlated to estrogen receptor status (ER; P = 0.0017), but not associated to the other clinical variables progesterone receptor (PR; P = 0.12), human epidermal growth factor receptor 2 (HER2; P = 0.12), Nottingham Histologic Grade (P = 0.57), and Ki-67 (P = 0.40) (Table 2). Microarray gene expression data were available for 42 of the 132 cases from a previous study [27]. Using this data, we found NEDD4 protein levels to be significantly correlated to NEDD4 mRNA expression level (P = 0.04) (Fig. 2a), supporting the specificity of the antibody and also indicating that NEDD4 mRNA may be an appropriate surrogate for NEDD4 protein levels in breast cancer.
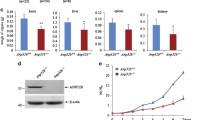
PTEN and NEDD4 protein and mRNA levels in the Swedish cohort. a NEDD4 protein levels were significantly correlated to NEDD4 mRNA levels (N = 42, P = 0.04). PTEN protein levels were not significantly correlated to b NEDD4 protein levels in breast cancer tissues (N = 123, P = 0.12) or c NEDD4 mRNA levels (N = 105, P = 0.73). PTEN mRNA levels were significantly correlated to d PTEN protein levels (N = 105, P < 0.001), e NEDD4 protein levels (N = 42, P = 0.02), and f NEDD4 mRNA levels (N = 105, R = 0.22, P = 0.03)
PTEN protein expression was previously determined by IHC for 123 of the 132 cases [9]. We tested whether NEDD4 protein levels were negatively associated to PTEN protein levels, however no correlation was seen in this Swedish breast cancer material (P = 0.12; Fig. 2b). This was inconsistent with the inverse correlation between the two proteins observed in a mouse prostate cancer model [14] and in lung cancers [20]. In fact, in our Swedish cohort the correlation trended positively, with 77 % of cases being PTEN-positive within the NEDD4-positive group compared to 64 % being PTEN-positive in the NEDD4-negative group (Fig. 2b).
In human bladder carcinoma, Wang et al. reported NEDD4 mRNA expression and PTEN mRNA expression to be uncorrelated, but that NEDD4 mRNA levels were inversely correlated to PTEN protein levels [14]. To investigate if it is also the case in breast tumors, we next considered the transcript levels of these genes using the GSE5325 microarray dataset of 105 breast tumors previously utilized to develop a gene expression signature for PTEN-loss [27]. In contrast to bladder cancer, we found no correlation between NEDD4 mRNA and PTEN protein expression (P = 0.73; Fig. 2c). PTEN mRNA, however, was highly correlated to PTEN protein (P < 0.001; Fig. 2d), which has been previously reported [27]. Unexpectedly, we found PTEN mRNA levels to be significantly positively correlated to NEDD4 protein expression (N = 42, P = 0.02; Fig. 2e) as well as to NEDD4 mRNA levels (N = 105, P = 0.03; Fig. 2f).
To validate these findings, two independent large-scale breast cancer cohorts from the NKI and TCGA were studied. The NKI cohort contained 295 breast tumor samples with microarray gene expression data [28, 29]. Tissue microarray sections were obtained and immunostained for PTEN protein, of which 267 cases were evaluable. Similar to the Swedish cohort, we found no correlation between NEDD4 mRNA and PTEN protein (P = 0.39; Fig. 3a). The strong positive correlation between PTEN mRNA and PTEN protein (P < 0.001; Fig. 3b), as well as the association of our previously published PTEN-loss signature [27] with loss of PTEN protein (P = 0.003; data not shown), were confirmed in this independent dataset. Moreover, the positive association between NEDD4 mRNA and PTEN mRNA found in our Swedish cohort was also validated in the NKI patient material (N = 295, P < 0.001; Fig. 3c).
PTEN mRNA/protein levels and NEDD4 mRNA levels in the NKI cohort. PTEN IHC scores were not associated to a NEDD4 mRNA levels (N = 267, P = 0.39), but were significantly correlated to b PTEN mRNA levels (N = 267, P < 0.001). c PTEN mRNA and NEDD4 mRNA levels were also significantly correlated (N = 295, R = 0.19, P < 0.001)
These associations were further corroborated in the TCGA breast carcinoma cohort containing RNA-sequencing (RNA-seq) gene expression profiles of primary breast tumors from 970 patients, of which 407 also had available PTEN protein expression data derived from reverse phase protein arrays [10]. In this large cohort the correlation between NEDD4 mRNA and PTEN protein was also significantly positive (P < 0.001; Fig. 4a). Additionally, PTEN mRNA and PTEN protein levels were positively correlated (P < 0.001; Fig. 4b), as observed in the Swedish and NKI cohorts. Lastly, the positive correlation between NEDD4 mRNA and PTEN mRNA levels was also confirmed in the TCGA dataset (P < 0.001; Fig. 4c).
PTEN and NEDD4 mRNA levels in the TCGA cohort. PTEN protein levels were significantly correlated to a NEDD4 mRNA levels (N = 407, R = 0.17, P < 0.001), and b PTEN mRNA levels (N = 407, R = 0.46, P < 0.001). c PTEN mRNA levels were significantly correlated to NEDD4 mRNA levels in the 970 primary breast tumors (R = 0.37, P < 0.001)
In conclusion, our study investigated whether PTEN was associated to NEDD4 in three large independent breast cancer sample cohorts. Contrary to reports in some other cancer forms, no inverse relationship was seen between NEDD4 transcript and PTEN protein levels. Rather, there was no correlation between NEDD4 protein and PTEN protein, and the correlation between NEDD4 mRNA/protein and PTEN mRNA was significantly positive. NEDD4-mediated poly-ubiquitination of PTEN may be an important mechanism that contributes to PTEN protein loss in bladder cancer [14] and non-small cell lung carcinoma [20]; whereas the results in gastric and colorectal cancers have been discrepant [25, 26]. Interestingly, in ovarian cancer HeLa cells, PTEN has also been reported to negatively regulate NEDD4 expression via the PI3K/AKT pathway, forming a potential negative feedback loop [30]. Our present study does not support NEDD4 as a major negative regulator of PTEN levels in human breast cancer. Additional studies are necessary to better delineate the underlying mechanisms of PTEN loss in this poor-prognosis subgroup.
References
Maehama T, Dixon JE (1998) The tumor suppressor, PTEN/MMAC1, dephosphorylates the lipid second messenger, phosphatidylinositol 3,4,5-trisphosphate. J Biolumin Chemilumin 273:13375–13378
Arch EM, Goodman BK, Van Wesep RA, Liaw D, Clarke K, Parsons R, et al (1997) Deletion of PTEN in a patient with bannayan-riley-ruvalcaba syndrome suggests allelism with cowden disease. Am J Med Genet 71:489–493
Liaw D, Marsh DJ, Li J, Dahia PL, Wang SI, Zheng Z, et al (1997) Germline mutations of the PTEN gene in cowden disease, an inherited breast and thyroid cancer syndrome. Nat Genet 16:64–67
Hobert JA, Eng C (2009) PTEN hamartoma tumor syndrome: an overview. Genitourin Med 11:687–694
Li J, Yen C, Liaw D, Podsypanina K, Bose S, Wang SI, et al (1997) PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275:1943–1947
Steck PA, Pershouse MA, Jasser SA, Yung WK, Lin H, Ligon AH, et al (1997) Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers. Nat Genet 15:356–362
Shaw RJ, Cantley LC (2006) Ras, PI(3)K and mTOR signalling controls tumour cell growth. Nature 441:424–430
Forbes SA, Beare D, Gunasekaran P, Leung K, Bindal N, Boutselakis H, et al (2015) COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res 43:D805–811
Saal LH, Holm K, Maurer M, Memeo L, Su T, Wang X, et al (2005) PIK3CA mutations correlate with hormone receptors, node metastasis, and ERBB2, and are mutually exclusive with PTEN loss in human breast carcinoma. Cancer Res 65:2554–2559
The Cancer Genome Atlas Network (2012) Comprehensive molecular portraits of human breast tumours. Nature 490:61–70
Simpson L, Parsons R (2001) PTEN: life as a tumor suppressor. Exp Cell Res 264:29–41
Wang X, Jiang X (2008) Post-translational regulation of PTEN. Oncogene 27:5454–5463
Saal LH, Gruvberger-Saal SK, Persson C, Lovgren K, Jumppanen M, Staaf J, et al (2008) Recurrent gross mutations of the PTEN tumor suppressor gene in breast cancers with deficient DSB repair. Nat Genet 40:102–107
Wang X, Trotman LC, Koppie T, Alimonti A, Chen Z, Gao Z, et al (2007) NEDD4-1 is a proto-oncogenic ubiquitin ligase for PTEN. Cell 128:129–139
Drinjakovic J, Jung H, Campbell DS, Strochlic L, Dwivedy A, Holt CE (2010) E3 ligase Nedd4 promotes axon branching by downregulating PTEN. Neuron 65:341–357
Goh CP, Low LH, Putz U, Gunnersen J, Hammond V, Howitt J, et al (2013) Ndfip1 expression in developing neurons indicates a role for protein ubiquitination by Nedd4 E3 ligases during cortical development. Neurosci Lett 555:225–230
Guo H, Qiao G, Ying H, Li Z, Zhao Y, Liang Y, et al (2012) E3 ubiquitin ligase Cbl-b regulates pten via Nedd4 in T cells independently of its ubiquitin ligase activity. Cell Rep 1:472–482
Chung S, Nakashima M, Zembutsu H, Nakamura Y (2011) Possible involvement of NEDD4 in keloid formation; its critical role in fibroblast proliferation and collagen production. Proc Jpn Acad Ser B Phys Biol Sci 87:563–573
Shi Y, Wang J, Chandarlapaty S, Cross J, Thompson C, Rosen N, et al (2014) PTEN is a protein tyrosine phosphatase for IRS1. Nat Struct Mol Biol 21:522–527
Amodio N, Scrima M, Palaia L, Salman AN, Quintiero A, Franco R, et al (2010) Oncogenic role of the E3 ubiquitin ligase NEDD4-1, a PTEN negative regulator, in non-small-cell lung carcinomas. Am J Pathol 177:2622–2634
Hong SW, Moon JH, Kim JS, Shin JS, Jung KA, Lee WK, et al (2014) p34 is a novel regulator of the oncogenic behavior of NEDD4-1 and PTEN. Cell Death Differ 21:146–160
Trotman LC, Wang X, Alimonti A, Chen Z, Teruya-Feldstein J, Yang H, et al (2007) Ubiquitination regulates PTEN nuclear import and tumor suppression. Cell 128:141–156
Fouladkou F, Landry T, Kawabe H, Neeb A, Lu C, Brose N, et al (2008) The ubiquitin ligase Nedd4-1 is dispensable for the regulation of PTEN stability and localization. Proc Natl Acad Sci U S A 105:8585–8590
Maddika S, Kavela S, Rani N, Palicharla VR, Pokorny JL, Sarkaria JN, et al (2011) WWP2 is an E3 ubiquitin ligase for PTEN. Nat Cell Biol 13:728–733
Yang Z, Yuan XG, Chen J, Lu NH (2012) Is NEDD4-1 a negative regulator of phosphatase and tensin homolog in gastric carcinogenesis? World J Gastroenterol 18:6345–6348
Eide PW, Cekaite L, Danielsen SA, Eilertsen IA, Kjenseth A, Fykerud TA, et al (2013) NEDD4 is overexpressed in colorectal cancer and promotes colonic cell growth independently of the PI3K/PTEN/AKT pathway. Cell Signal 25:12–18
Saal LH, Johansson P, Holm K, Gruvberger-Saal SK, She QB, Maurer M, et al (2007) Poor prognosis in carcinoma is associated with a gene expression signature of aberrant PTEN tumor suppressor pathway activity. Proc Natl Acad Sci U S A 104:7564–7569
van de Vijver MJ, He YD, van't Veer LJ, Dai H, Hart AA, Voskuil DW, et al (2002) A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 347:1999–2009
van 't Veer, LJ, Dai, H, van de Vijver, MJ, He, YD, Hart, AA, Mao, M et al (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415:530–536
Ahn Y, Hwang CY, Lee SR, Kwon KS, Lee C (2008) The tumour suppressor PTEN mediates a negative regulation of the E3 ubiquitin-protein ligase Nedd4. Biochem J 412:331–338
Acknowledgments
We thank Kristina Lövgren for laboratory assistance, Björn Frostner and Susanne André for administrative support, and members of the Translational Oncogenomics Unit, Division of Oncology and Pathology, for valuable discussion. This study was funded in part by the Swedish Research Council, Swedish Cancer Society, Governmental Funding of Clinical Research within National Health Service, Crafoord Foundation, Mrs. Berta Kamprad Foundation, Lund University Medical Faculty, Gunnar Nilsson Cancer Foundation, Skåne University Hospital Foundation, BioCARE Research Program, King Gustav Vth Jubilee Foundation, and the Krapperup Foundation.
Author contributions
Y.C. and L.H.S. conceived the study and performed the experiments. M.J.vdV. and R.P. provided reagents. Y.C., H.H., and L.H.S. analyzed the data. Y.C. and L.H.S wrote and revised the manuscript. L.H.S. supervised the project.
Competing interests
The authors declare no competing financial interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Chen, Y., van de Vijver, M.J., Hibshoosh, H. et al. PTEN and NEDD4 in Human Breast Carcinoma. Pathol. Oncol. Res. 22, 41–47 (2016). https://doi.org/10.1007/s12253-015-9971-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-015-9971-2