Abstract
Draba (Brassicaeae), a model group for diversification and evolution in Arctic and mountain habitats, is taxonomically challenging and many of its species are insufficiently investigated. One such species is D. pacheri, an endemic of the eastern European Alps and the western Carpathians (here presumably extinct). Several hypotheses exist with respect to the phylogenetic position and the taxonomy of this species, but none of these has ever been tested using molecular data. In this article we examine (i) DNA sequence data to assess the phylogenetic position of D. pacheri within the genus and (ii) AFLP fingerprint data as well as morphometric data to address whether this species can be divided taxonomically into species or subspecies. DNA sequence data firmly place D. pacheri within the Core Draba Group III, whose internal relationships are, however, insufficiently resolved to precisely identify the closest relative of D. pacheri. AFLP data identify several genetically divergent lineages corresponding to geographically distinct regions. Although these lineages are congruent with hypotheses distinguishing either two species (D. pacheri s. str., D. norica) or one species with several subspecies, the lack of clear morphological separation, both with respect to the entire set of traits and single presumably diagnostic characters such as trichome morphology, renders recognition of a single species D. pacheri, as suggested previously, the best taxonomic solution. The deep and geographically strongly structured splits of D. pacheri likely are the result of isolation in several Pleistocene refugia and warrant that conservation efforts should involve populations from each of the main geographic subgroups.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Draba is the largest within the family of Brassicaceae, comprising more than 370 species worldwide (Warwick et al. 2006). Draba is mainly distributed in Arctic, Subarctic, alpine and montane areas (Jordon-Thaden et al. 2010). Despite considerable progress with respect to the delimitation and infrageneric structure of Draba (Jordon-Thaden et al. 2010), many species are yet to be studied using molecular phylogenetic means and numerous taxonomic questions at the species level remain, affecting our understanding of this model group as to its diversification and evolution in Arctic and (high-)mountain habitats.
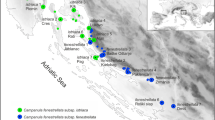
One of these insufficiently investigated species is D. pacheri Stur. After discovering it in the Austrian Alps, in the eastern Hohe Tauern on Stern mountain (= Sternspitze; pop. 3, Fig. 1), R. Gussenbauer gave the plant to D. Pacher, who sent it further to D. Stur (Melzer and Prugger 1986). Stur eventually described it as a new species in 1855 (Stur 1855) and named it, after its presumed discoverer, D. pacheri (Melzer and Prugger 1986). In 1931, F. Widder reported populations of a morphologically similar species from the mountain range of Koralpe in the Lavanttaler Alpen (pop. 7, Fig. 1), more than 100 km east of Stern mountain. Allegedly differing from D. pacheri in plant height, number of cauline leaves, silique length, fruit pedicel length and especially trichome morphology (Widder 1931, 1934; Table 1), Widder considered the population from Koralpe to be sufficiently distinct to describe it as a separate locally endemic species, D. norica Widder (Widder 1931). Later, apart from further populations in the eastern Hohe Tauern (Melzer and Prugger 1986; Stöhr et al. 2007), geographically intervening populations of D. pacheri and D. norica in the Gurktaler Alpen (pop. 4) and the Seetaler Alpen (pops 5 and 6; Fig. 1) were found (Melzer 1962; Melzer and Prugger 1986). Buttler (1967) reported a specimen from the Belianské Tatry in Slovakia (Fig. 1), where the species is presumably extinct (Peniašteková et al. 2000). As populations from the Seetaler Alpen and from the Belianské Tatry could not be unambiguously assigned to either D. pacheri or D. norica (Table 1), Buttler (1967; the population from the Gurktaler Alpen was not known to him) regarded all thitherto known populations as belonging to a single, though morphologically quite variable, species D. pacheri, a treatment followed in modern floras and checklists (e.g. Euro+Med 2006–, Fischer et al. 2008). Nonetheless, emphasizing some morphological differences and the pronounced geographic separation among populations, Melzer and Prugger (1986) suggested that the taxonomic recognition of geographically separated subspecies may be advisable. As D. pacheri is a species of conservation concern, listed in both national and regional Red data lists (Kniely et al. 1995; Niklfeld and Schratt-Ehrendorfer 1999), clarification of taxonomic uncertainties is necessary.
Distribution of Draba pacheri in Europe (insert) and in the Austrian Alps (population numbers as in Table 2; the unsampled westernmost population is indicated by a diamond)
Several hypotheses have been put forward with respect to the taxonomic position of D. pacheri. Pacher (1885) suggested a close relationship to the Arctic D. glabella (as D. hirta L. siliculis glabris), an idea seconded by Ekman (1917), who even put D. pacheri into the synonymy of D. glabella (using for the latter the name D. magellanica subsp. borea). Weingerl (1923) suggested that D. pacheri is a neo-endemic derivative of D. tomentosa, whereas Schulz (1927) suggested a closer relationship to D. dubia. Widder (1931, 1934) considered D. pacheri close to an ancestral form of the group including, among others, D. dubia, D. tomentosa and D. incana. Finally, based on morphological and cytological evidence, Buttler (1967) considered D. pacheri part of the Arctic-Asian D. glabella-complex, yet, in contrast to Pacher (1885) and Ekman (1917), with a closer relationship to Central Asian than to Arctic species. None of these hypotheses has, however, been tested using molecular phylogenetic data.
Here we address the phylogenetic position of D. pacheri and assess whether the disjoint populations in the Alps are genetically and morphologically distinct and so may merit taxonomic recognition. To this end, we sequenced two markers (nuclear ITS and plastid trnLF) allowing the species to be placed in the molecular phylogenetic framework established recently (Jordon-Thaden et al. 2010), and generated molecular (AFLP fingerprint data) and morphometric data (focusing on suggested diagnostic characters such as leaf trichomes or fruit dimensions: Table 1) to detect entities worthy of taxonomic recognition.
Material and Methods
Plant Material
The plant material of D. pacheri was collected in four regions (Hohe Tauern, Gurktaler Alpen, Seetaler Alpen and Koralpe) between June and August 2013. In 2014, a search for the population in Slovakia, in the Belianské Tatry, was undertaken, but was not successful. All fieldwork was carried out according to the permissions granted by the responsible public authorities. Thus, no plant vouchers were collected, and only single leaves were taken. GPS coordinates were acquired using a Garmin eTrex 10 GPS device. Possible sampling sites were taken from the literature (Widder 1931; Melzer 1962; Melzer and Prugger 1986); sampling sites within a 2-km distance were grouped into populations, well reflecting the regional aggregation of sampling sites. The westernmost population in the Hohe Tauern (Stöhr et al. 2007), located about 18 km west of population 1, was not visited for logistical reasons. The sampling locations and populations are listed in Table 2 and the populations are shown in Fig. 1. From each sampling site, one to five plants were chosen for the collection of leaf samples for further analysis, resulting in 36 individuals of D. pacheri and six individuals of the outgroup species D. siliquosa (five from the Gurktaler Alpen, pop. 4, and one from the cirque ‘Seekar’ in the Koralpe, pop. 7). Whenever possible, usually three rosette leaves or, in their absence, cauline leaves were collected and dried in silica gel to be used for both DNA extraction and trichome analysis.
Molecular Data
DNA was extracted using the peqGOLD Plant DNA Mini Kit (peqlab, Erlangen, Germany) according to the manufacturer’s instructions. Because of the critical limitation of plant material, only 2–5 mg of each plants’ leaf tissue could be used for extraction to keep some of the leaf material for a second extraction if necessary. The extracted DNA was eluted in 100 μl HPLC water and stored at −20°C.
For DNA sequencing, five samples of D. pacheri (two from the Hohe Tauern, one from the Gurktaler Alpen, one from the Seetaler Alpen and one from the Koralpe) and one of D. siliquosa (from the Gurktaler Alpen) were used. The nuclear ITS region was amplified using primers ITS 4 and ITS 5 (White et al. 1990); the plastid trnLF region (trnL-intron and trnL-trnF intergenic spacer) was amplified using primers c and f (Taberlet et al. 1991). The PCRs were carried out on an Applied Biosystems GeneAmp PCR System 9700 thermocycler (ABI, Foster City, CA, USA). The PCR mix contained 0.9 μl of DNA extract of unknown concentration, 9 μl 1.1X ReaddyMix PCR (Thermo Fisher, Braunschweig, Germany), 9 μl HPLC-water and 0.7 μl of each primer. For ITS, the PCR conditions were 4 min at 94°C; 35 cycles each with 1 min at 95°C, 1 min at 51°C, 1 min at 68°C, and a final elongation step of 10 min at 72°C. For trnLF, the conditions were 1.5 min at 94°C, 35 cycles each with 15 s at 94°C, 15 s at 51°C, and 1.5 min at 72°C; and extension for 10 min at 72°C. PCR products were cleaned with 1 µl FastAP and 0.5 µl Exo I (Thermo Fisher) following the manufacturer’s instructions. Cycle sequencing was carried out in 10-μl volumes, containing 5.3 μl of the PCR product, 2 μl trehalose, 1.6 μl sequencing buffer, 0.6 μl Big Dye and 0.5 μl primers. After cleaning the products with Sephadex G-50 Fine (Sigma-Aldrich, St. Louis, USA), they were separated on a cycle sequencer (ABI 3730 DNA Analyzer capillary sequencer; Applied Biosystems, Foster City, CA, USA).
Sequence reads were assembled in SeqMan 5.05 (DNAStar, Madison, WI, USA). Using Bioedit 7.2.5 (Hill 1999), the ITS and trnL sequences were aligned to the datasets of Jordon-Thaden et al. (2010) containing ITS and trnL sequences of 169 Draba species. The three datasets (nuclear sequences, plastid sequences and combined sequences) were analysed using maximum likelihood with the fast bootstrap option with 1,000 replicates (Stamatakis et al. 2008) using RAxML 8.0.14 (Stamatakis 2014).
For AFLPs, 38 samples (36 of D. pacheri and two of D. siliquosa) were analysed. The AFLP protocol followed Vos et al. (1995) with the modifications described in Schönswetter et al. (2009) and Rešetnik et al. (2014). Three primer combinations were employed for the selective PCR (fluorescent dye in brackets): EcoRI (6-FAM)-ACA/MseI CAT, EcoRI (NED)-ACC/MseI-CAT and EcoRI (VIC)-AAG/MseI-CTG. Purification and visualization of PCR products were done as described in Rebernig et al. (2010). All samples were processed in a single PCR round. Eleven samples (26%) were used as replicates to test reproducibility. Raw AFLP profiles were aligned with the internal size standard using ABI Prism GeneScan 3.7.1 (Applied Biosystems) and scored manually with Genographer 1.6.0 (version no longer available). Following Bonin et al. (2004), the error rate was calculated as the ratio of mismatches over matches in AFLP profiles of replicated individuals.
For the whole dataset (i.e. including the outgroup), a neighbour joining tree based on uncorrected p-distances with 1,000 bootstrap replicates was constructed using SplitsTree 4.13.1 (Huson and Bryant 2006). Subsequent analyses were done for D. pacheri only, removing bands that were either monomorphic or present/absent in a single individual only. AFLPdat (Ehrich 2006) was used to calculate ‘rarity1’ (henceforth referred to as the rarity index), corresponding to the frequency down-weighed marker value of Schönswetter and Tribsch (2005), and Nei’s gene diversity (Nei 1973) for each of the four regions (Hohe Tauern, Gurktaler Alpen, Seetaler Alpen, Koralpe) as well as for each population (except population 6, where only a single individual was found). Using SplitsTree 4.13.1, a NeighbourNet with 1,000 bootstrap replicates was created. A principal coordinate analysis based on Jaccard’s similarity coefficient was done using FAMD 1.31 (Schlüter and Harris 2006). Population structure was inferred using the Bayesian clustering approach in Structure 2.3.4 (Pritchard et al. 2000; Falush et al. 2007) run in parallel using the R-package ParallelStructure 1.0 (Besnier and Glover 2013). Using an admixture model with correlated allele frequencies (as suggested for closely related populations: Falush et al. 2003) and recessive alleles, ten replicate runs for each K (number of groups) ranging from 1 to 10 were calculated using a burn-in of 5 × 105 iterations followed by 2 × 106 additional MCMC iterations. The optimal number of groups was identified using the deltaK approach (Evanno et al. 2005) as implemented by the StructureHarvester web 0.6.94 (Earl and vonHoldt 2012).
Morphometric Data
Herbarium material from various Austrian herbaria (GJO, GZU, KL and LI; abbreviations follow the Index Herbarium: http://sweetgum.nybg.org/science/ih, accessed on 16 October 2020), amounting to 161 vouchers excluding material grown in botanical gardens, as well as fresh material from 32 plants was examined for ten characters (Table S1 in the Electronic supplementary material): (1) aboveground plant height; (2) number of cauline leaves; (3) length of a central rosette leaf; (4) length of the stem between rosette leaves and lowest pedicel; (5) angle between fruit pedicel and stem; (6) length of fruit pedicel; (7) fruit length; (8) fruit width; (9) ratio of fruit length and fruit width; (10) ratio of fruit pedicel length and fruit length. Characters (7) to (9) were obtained by averaging over measurements from the three lowest, sufficiently developed siliques whereas characters (5) and (6) were averaged over three measurements from the three lowest fruits, irrespective of their developmental status (properly developed or putatively aborted). For plants selected in the field, these data were collected in vivo in order not to damage the plants. The final sample sizes for each region were n = 124 for the Hohe Tauern, n = 9 for the Gurktaler Alpen, n = 24 for the Seetaler Alpen, and n = 36 for Koralpe.
Because many of the herbarium vouchers as well as some of the fresh material were in bad condition, trichome analysis was carried out on fully developed leaves collected in the field (thirteen samples from the Hohe Tauern, five samples from the Gurktaler Alpen, five samples from the Seetaler Alpen, one sample from the Koralpe, totalling 24 samples) and from selected herbarium vouchers (three from the Seetaler Alpen and six from the Koralpe). The proportion of different trichome types (see below) on the lower surface of the proximal half of the rosette leaves was determined using a stereomicroscope, and images were taken with a digital camera. To this end, the investigated trichome types were marked on the images with specific symbols to prevent counting errors; in case of uncertainties concerning the classification of trichomes on the digital images, a stereomicroscope was used to examine the leaf surface directly. Only stellate trichomes composed of four primary branches were counted after being classified into two types differing in the size and number of secondary branches, as this character was considered of particular relevance by Widder (1931) to differentiate D. norica from D. pacheri. Both types have none to several tooth-like secondary branches (i.e. with a length : width ratio < 2 and being maximally 0.15 times as long as the primary branch) but differ in the number of long secondary branches (i.e. with a length : width ratio of at least 2 and being at least 0.15 times as long as the primary branch): type A has maximally one long secondary branch, while type B has at least two long secondary branches on the same or on different primary branches (Fig. 2).
Morphometric data were analysed using PAST 4.03 (Hammer et al. 2001). Differences in single morphometric characters between the four geographically distinct regions were tested via Kruskal-Wallis tests (only for character 7, fruit length, a normal distribution for residuals was not rejected: Shapiro-Wilk W = 0.9943, p = 0.6783) and subsequent Dunn’s post hoc tests applying sequential Bonferroni correction. Principal component analyses (PCA) were done on both the entire morphometric dataset (i.e. morphometric characters 1–10 plus proportion of hair type A) as well as a reduced dataset excluding the single discrete character (character 2: number of cauline leaves) as well as hair type (due to a high proportion of missing data: 83%) using a normalized variance-covariance matrix (correlation) and using iterative imputation of missing values.
Results
In Austria, all visited populations of D. pacheri were confirmed in all four regions (Hohe Tauern, Gurktaler Alpen, Seetaler Alpen, Koralpe). In populations with more than one sampling site, however, D. pacheri could not be confirmed for some sampling sites, most prominently the type locality of D. norica on the eastern slopes of the Koralpe (Seekar; Table 2).
Molecular Data
Newly obtained DNA sequence data (from five accessions of D. pacheri and one of D. siliquosa) are available from the GenBank database under accession numbers OK336415–OK336420 (ITS) and OK346544–OK346549 (trnLF). From DNA sequence data, D. pacheri, without any sequence variation, was inferred across all three datasets (log-likelihood scores of −6,418.9745 for nuclear ITS, −6,333.9862 for trnLF, and -14,489.7883 for the combined data) to fall into the clade of Core Draba Group III as defined by Jordon-Thaden et al. (2010; Fig. 3; phylogenetic trees from all three datasets available as a zip file in the Electronic supplementary material). Because of the overall low and/or poor resolution within this clade, precise relationships of D. pacheri could not be ascertained.
For AFLPs, in the larger dataset (D. pacheri and the outgroup D. siliquosa) 194 polymorphic fragments were scored with an error rate of 0.0007. In the neighbour joining tree (Fig. 4), rooted with D. siliquosa, strong and well supported (bootstrap support [BS] 98–100) differentiation was not only found among the four geographic regions, but also within the Hohe Tauern (pops 1–3). Here, pop. 2 (BS 98) was as distinct from the remaining populations (pops 1 and 3: BS 99) as it was from the single population from the Gurktaler Alpen (pop. 4: BS 100). However, relationships among these three clades (pops 1 and 3; pop. 2; pop. 4) were not resolved. Subsequent sister groups to pops 1–4 were the population (pop. 7; BS 100) from the Koralpe (BS 93) and the ones (pops 5–6; BS 100) from the Seetaler Alpen (BS 100).
The dataset including only accessions of D. pacheri comprised, after exclusion of 88 monomorphic bands and ten bands that were present or absent in only a single individual, 59 polymorphic fragments. Calculated for the four regions (Hohe Tauern, Gurktaler Alpen, Seetaler Alpen, Koralpe), gene diversity varied approximately thirteen-fold, ranging from 0.0068 (Gurktaler Alpen) to 0.0866 (Hohe Tauern), and the rarity values varied approximately two-fold from 1.2575 (Hohe Tauern) to 2.5122 (Koralpe; Table 3). At the population level, gene diversity varied approximately 2.7-fold, ranging from 0.0068 (pop. 4 from Gurktaler Alpen and pop. 2 from Hohe Tauern) to 0.0181 (pop. 3 from Hohe Tauern), and the rarity values varied approximately two-fold from 1.2119 (pop. 1 from the Hohe Tauern) to 2.5122 (pop. 7 from Koralpe; Table 3).
The NeighbourNet analysis (Fig. 5a) revealed three major groups corresponding to the Koralpe (pop. 7; BS 100), Seetaler Alpen (pops 5, 6; BS 100) and Gurktaler Alpen plus Hohe Tauern (pops 1–4; BS 97). Within the latter group, the same three subgroups already identified by the neighbour joining analysis were found: Gurktaler Alpen (pop. 4; BS 100), Hohe Tauern pop. 2 (BS 98) and Hohe Tauern pops 1 and 3 (BS 96); relationships among those three subgroups were unclear because of reticulations (Fig. 5a).
Genetic structure in Draba pacheri inferred from AFLP data: (a) NeighbourNet (numbers at splits are bootstrap values, those in parentheses at the population number indicate the number of individuals of this genotype), (b) principal coordinate analysis, (c) genetic clusters identified via the Bayesian clustering approach in Structure with two and five clusters, K, respectively
In the principal coordinate analysis (Fig. 5b), populations were separated according to geography. Specifically, the Koralpe population (pop. 7) was separated from the others along the first axis (explaining 56.28% of the variation), the Seetaler Alpen ones (pops 5–6) along the second axis (explaining 24.35% of the variation), and the Gurktaler Alpen (pop. 4) and Hohe Tauern populations (pops 1–3) from each other along the third axis (explaining 12.78% of the variation; Fig. 5b).
At K = 2, identified by deltaK as the optimal number of groups (Fig. S1 in the Electronic supplementary material), D. pacheri was divided into a western group (pops 1–4) and an eastern group (pops 5–7), essentially without any admixture (Fig. 5c). At K = 5, suggested by a distinct peak in the deltaK distribution and by a stable likelihood maximum (Fig. S1), D. pacheri was divided according to the four geographic regions, additionally assigning the populations from the Hohe Tauern to two separate groups (pop 2 vs pops 1 and 3), again essentially without any admixture among groups (Fig. 5c).
Morphometric Data
The full data matrix is available as Table S1 in the Electronic supplementary material. Of the ten morphometric characters, only two differed significantly between one geographic region and the remaining ones. Specifically, plants from the Koralpe had a significantly smaller ratio of fruit length to fruit width whereas plants from the Hohe Tauern had a significantly smaller ratio of fruit pedicel length to fruit length (Fig. 6). For the remaining characters, significant differences between two or three geographic regions were found commonly, but any such differences were blurred by one or more geographic regions showing intermediate values (Fig. 6). For example, plants from the Hohe Tauern had a significantly lower number of cauline leaves than plants from both the Seetaler Alpen and the Koralpe, but plants from the Gurktaler Alpen did not differ significantly from any of the other three regions.
The on average lowest proportion of trichomes with two or more long secondary branches (type B) was found in plants from the Gurktaler Alpen (2%) and from the Koralpe (11%), the on average highest proportion of these trichomes was found in plants from the Seetaler Alpen (23%) and from the Hohe Tauern (26%). However, variation was large in all regions (Hohe Tauern 3–51%, Seetaler Alpen 6–34%, Koralpe 0–56%) except for the Gurktaler Alpen (0–3%).
In the PCA, no separation among the Hohe Tauern, Gurktaler Alpen, Seetaler Alpen, and Koralpe was found irrespective of the dataset used (Fig. 7). Separation along the main axis was mainly due to plant height (char. 1), leaf length (char. 3), scape length (char. 4) and fruit pedicel length (char. 6) whereas separation on the second axis was mainly due to hair type (full dataset only) and fruit breadth (char. 8; Table S2).
Principal component analysis of morphometric data of Draba pacheri based on the (a) complete dataset and (b) a dataset including only continuous data available in at least 99% of the samples. Symbols indicate geographic origin: dots, Hohe Tauern; squares, Gurktaler Alpen; plus sign, Seetaler Alpen; crosses, Koralpe
Discussion
DNA sequence data firmly place D. pacheri in the Core Draba Group III as circumscribed by Jordon-Thaden et al. (2010). However, because of the low resolution at the backbone of the phylogenetic tree and the generally low levels of support, identifying the closest relative of D. pacheri is not possible. A closer relationship with the Nordic D. glabella, as suggested by Buttler (1967) based on morphology and ploidy level, is not refuted by sequence data, but the complexity of D. glabella, whose accessions do not form a cohesive group and intermix with other species (Fig. 3), renders even the delimitation of a D. glabella complex (as defined by Buttler 1967) impossible. Evidently, further studies using better-resolving phylogenetic markers and a broader sampling (e.g. including more samples of the western North American D. praealta, which is inferred, though without support, as the closest relative of D. pacheri) will be necessary to address the precise phylogenetic position of D. pacheri.
Several hypotheses have been suggested with respect to the taxonomic treatment of D. pacheri. Widder (1931, 1934) suggested, based on morphological differences especially concerning trichome morphology, separating D. norica from the Koralpe from D. pacheri from the Hohe Tauern (the geographically intermediate populations from the Seetaler Alpen and Gurktaler Alpen were not known yet). Buttler (1967) found that populations from the Seetaler Alpen and from the Tatras could not be readily assigned to D. norica or D. pacheri and blurred the distinction between these two species; consequently, he merged them in a single species. Finally, Melzer and Prugger (1986), additionally including the population from the Gurktaler Alpen, suggested that the four groups identified by Buttler (1967; Hohe Tauern, Seetaler Alpen, Koralpe, Tatry) could be treated as subspecies. Indeed, genetic data were compatible with the distinction of two or three (infraspecific) taxa in the Alps. Specifically, our structure analysis (Fig. 5c) suggests two lineages (Hohe Tauern and Gurktaler Alpen; Seetaler Alpen and Koralpe) corresponding to the delimitation of D. pacheri s. str. and to D. norica used by Widder (1931, 1934), Melzer (1962), and Melzer and Prugger (1986) whereas our NeighbourNet and PCoA analyses (Fig. 5a,b) suggest three lineages (Hohe Tauern and Gurktaler Alpen; Seetaler Alpen; Koralpe) corresponding to subspecies as envisaged by Melzer and Prugger (1986). Neither of these groups can, however, be diagnosed morphologically, as morphological characters either only define single groups (e.g. individuals in Seetaler Alpen are taller than in all other regions, see Fig. 7, but this character is expected to be little reliable as plant growth will be affected by random fertilization from sheep or chamois droppings: Melzer and Prugger 1986) or are incongruent with genetic grouping (the high proportion of strongly branched trichomes considered characteristic for D. pacheri s. str. joins the Seetaler Alpen with the Hohe Tauern to the exclusion of the Gurktaler Alpen). In the absence of any clear and consistent morphological differentiation, keeping a single species without any infraspecific taxa, as done by Buttler (1967), seems to be the most prudent option.
Each of the four geographic groups within D. pacheri from the Alps is associated with well-known Pleistocene refugia (Tribsch and Schönswetter 2003). An association with local refugia may also explain the unexpectedly pronounced differentiation within the Hohe Tauern (pop. 2 versus pops 1 and 3). The deep genetic split separating the Koralpe and the Seetaler Alpen populations from each other and from the remaining populations (Fig. 5a,b) is likely due to a combination of persistent isolation in Pleistocene refugia and small populations sizes.
Draba pacheri is a species of protection concern. In Slovakia it is considered ‘regionally extinct’ (Turis et al. 2014). During this study, a few subpopulations reported from the Austrian Alps could not be confirmed either. The most prominent of those is on the eastern slope of the Koralpe (cirque ‘Seekar’), the type locality of D. norica, where several searching campaigns were not successful; hence, we cannot exclude that D. pacheri may be locally extinct there (the species is still occurring on the western slope of the Koralpe, less than 1 km from the type locality). Because the majority of the subpopulations in Austria could be confirmed, there appears to be no immediate threat to this species, but low genetic diversity within some regions (Table 3) and global warming are expected to negatively affect the survival of small populations at rather low elevations, especially those in the Gurktaler and Seetaler Alpen. The pronounced genetic divergence among different geographic regions renders it necessary to protect each of the populations individually.
References
Besnier F, Glover KA (2013) ParallelStructure: a R package to distribute parallel runs of the population genetics program STRUCTURE on multi-core computers. PLOS ONE 8:e70651
Bonin A, Bellemain E, Bronken Eidesen P, Pompanon F, Brochmann C, Taberlet P (2004) How to track and assess genotyping errors in population genetics studies. Molec Ecol 13:3261–3273
Buttler KP (1967) Zytotaxonomische Untersuchungen an mittel- und südeuropäischen Draba-Arten. Mitt Bot Staatssamml München 6:275–362
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genet Resources 4:359–361
Ehrich D (2006) aflpdat: a collection of r functions for convenient handling of AFLP data. Molec Ecol Notes 6:603–604
Ekman E (1917) Zur Kenntnis der nordischen Hochgebirgs-Drabae. Kungl Svenska Vetensk Acad Handl 57:1–68
Euro+Med (2006–) Euro+Med PlantBase – the information resource for Euro-Mediterranean plant diversity. Available at: https://ww2.bgbm.org/EuroPlusMed (Accessed 8 November 2019)
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molec Ecol 14:2611–2620
Falush D, Stephens M., Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Molec Ecol Notes 7:574–578
Fischer MA, Oswald K, Adler W (2008) Exkusionsflora für Österreich, Liechtenstein und Südtirol, 3. Auflage. Biologiezentrum der Oberösterreichischen Landesmuseen, Linz
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: Paleontological statistics software package for education and data analysis. Palaeontol Electronica 4:art9
Hill TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Molec Biol Evol 23:254–267
Jordon-Thaden I, Hase I, Al-Shehbaz I, Koch MA (2010) Molecular phylogeny and systematics of the genus Draba (Brassicaceae) and identification of its most closely related genera. Molec Phylogen Evol 55:524–540
Kniely G, Niklfeld H, Schratt-Ehrendorfer L (1995) Rote Liste der gefährdeten Farn- und Blütenpflanzen Kärntens. Carinthia II 185:353–392
Melzer H (1962) Neues zur Flora von Steiermark (V). Mitt Naturwiss Vereines Steiermark 92:77–100
Melzer H, Prugger O (1986) Zur Kenntnis von Draba pacheri Stur, Pachers Felsenblümchen. Carinthia II 176:321–331
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Niklfeld H, Schratt-Ehrendorfer L (1999) Farn- und Blütenpflanzen. In Niklfeld H (ed.) Rote Listen gefährdeter Pflanzen Österreichs. austria medien service, Graz, pp 33–152
Pacher D (1885) IV. Systematische Aufzählung der in Kärnten wildwachsenden Gefässpflanzen. II. Abtheilung. Dicotyledones. Familie: Umbelliferae bis Violaceae. Jahrb Naturhist Landesmus Kärnten 17:49–216
Peniašteková M, Petrík A, Kliment J (2000) Vysokohorské druhy rodu Draba L. (Brassicaceae) zo sekcie Draba L. na Slovensku. Acta Environm Univ Comen 10:111–116
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rebernig CA, Weiss-Schneeweiss H, Schneeweiss GM, Schönswetter P, Obermayer R, Villaseñor JL, Stuessy TF (2010) Quaternary range dynamics and polyploid evolution in an arid brushland plant species (Melampodium cinereum, Asteraceae). Molec Phylogen Evol 54: 594–606
Rešetnik I, Frajman B, Bogdanović S, Ehrendorfer F, Schönswetter P (2014) Disentangling relationships among the diploid members of the intricate genus Knautia (Caprifoliaceae, Dipsacoideae). Molec Phylogen Evol 74:97–110
Schlüter PM, Harris SA (2006) Analysis of multilocus fingerprinting data sets containing missing data. Molec Ecol Notes 6:569–572
Schönswetter P, Tribsch A (2005) Vicariance and dispersal in the alpine perennial Bupleurum stellatum L. (Apiaceae). Taxon 54:725–732
Schönswetter P, Solstad H, Escobar Garcia P, Elven R (2009) A combined molecular and morphological approach to the taxonomically intricate European mountain plant Papaver alpinum s.l. (Papaveraceae) – Taxa or informal phylogeographical groups? Taxon 58:1326–1343
Schulz OE (1927) Cruciferae-Draba et Erophila. In Engler A (ed.) Das Pflanzenreich. Regni vegetabilis conspectus IV/105 Heft 89. Wilhelm Engelmann, Leipzig, pp 1–396
Stamatakis A, Hoover P, Rougemont J (2008) A rapid bootstrap algorithm for the RAxML web servers. Syst Biol 57:758–771
Stamatakis A (2014) RAxML Version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Stöhr O, Pilsl P, Essl F, Hohla M, Schröck C (2007) Beiträge zur Flora von Österreich, II. Linzer Biol Beitr 39:155–292
Stur D (1855) Draba Pacheri Stur. Österr Bot Wochenbl 5:49–50
Taberlet T, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Pl Molec Biol 17:1105–1109
Tribsch A, Schönswetter P (2003) Patterns of endemism and comparative phylogeography confirm palaeo-environmental evidence in the Eastern European Alps. Taxon 52:477–497
Turis P, Kliment J, Feráková V, Dítě D, Eliáš P, Hrivnák R, Košťál J, Šuvada R, Mráz P, Bernátová D (2014) Red List of vascular plants of the Carpathian part of Slovakia. Thaiszia 24:35–87
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucl Acids Res 23: 4407–4414
Warwick SI, Francis A, Al-Shehbaz IA (2006) Brassicaceae: species checklist and database on CD-Rom. Pl Syst Evol 259:249–258
Weingerl H (1923) Beiträge zu einer Monographie der europäisch-asiatischen Arten aus der Gattung Draba, sect. Leucodraba. Bot Arch (Königsberg) 4:9–109
Widder FJ (1931) Draba norica, eine neue Ostalpenpflanze. Sitzungsber Akad Wiss Wien Math-Naturwiss Kl Abt 1 140:619–632
Widder FJ (1934) Beobachtungen an Draba pacheri Stur. Österr Bot Z 83:255–265
White TJ, Bruns T, Lee S, Taylor S (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: A guide to methods and applications. Academic Press, New York, pp 315–322
Acknowledgements
Data collection was done in accordance with the requirements of the respective permits (permit SP3-NS-2128/2013 for populations in the Hohe Tauern; permit FE3-NS-1679/2012 for the population in the Gurktaler Alpen; permit WO3-NS-2007/2013 for the western population in the Koralpe; permit ABT13-53S-7/2012-113 for the – not found – eastern population in the Koralpe and the populations in the Seetaler Alpen). We are indebted to M. Magauer and D. Pirkebner for conducting the AFLP lab work.
Funding
Open access funding provided by University of Vienna.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kucs, E., Schönswetter, P. & Schneeweiss, G.M. Deep phylogeographic splits but no taxonomic structure in the disjointly distributed Draba pacheri (Brassicaceae), a subendemic of the Eastern Alps. Folia Geobot 56, 179–192 (2021). https://doi.org/10.1007/s12224-021-09400-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12224-021-09400-z