Abstract
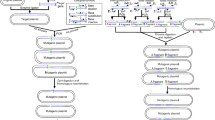
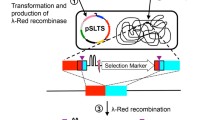
The subject of this review covers modern experimental procedures for chromosomal gene replacement in Escherichia coli and related bacteria, which enable the specific substitution of targeted genome sequences with copies of those carrying defined mutations. Two principal methods for gene replacement were established. The first “in–out” method is based on integration of plasmid into bacterial chromosome and subsequent resolving of the cointegrate. The “linear fragment” method (recombineering) is based on homologous recombination mediated by short homology arms at the ends of linear DNA molecule. Many new protocols and improvements in targeted gene replacement were introduced during the last 10 years. These methods are well suited for high-throughput functional gene studies and for many biotechnological applications.





Similar content being viewed by others
Abbreviations
- λ Red:
-
Genes gam, bet, and exo from bacteriophage λ mediating linear DNA recombination
- FLP:
-
Site-specific flip recombinase of the 2-μ plasmid of Saccharomyces cerevisiae
- FRT:
-
FLP recombinase recognition site
- Cre:
-
Site-specific recombinase from bacteriophage P1
- loxP:
-
Cre recombinase recognition site
- BAC:
-
Vector(s), bacterial artificial chromosome(s)
References
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, Datsenko KA, Tomita M, Wanner BL, Mori H (2006) Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2:2006–2008
Bartra SS, Styer KL, O'Bryant DM, Nilles ML, Hinnebusch BJ, Aballay A, Plano GV (2008) Resistance of Yersinia pestis to complement-dependent killing is mediated by the ail outer membrane protein. Infect Immun 76:612–622
Bloor AE, Cranenburgh RM (2006) An efficient method of selectable marker gene excision by Xer recombination for gene replacement in bacterial chromosomes. Appl Environ Microbiol 72:2520–2525
Burian J, Nguyen T, Kľučár Ľ, Guller L, Lloyd-Jones G, Stuchlík S, Siekel P, Fejdi P, Turňa J (1998) In vivo and in vitro cloning and phenotype characterization of tellurite resistance determinant conferred by plasmid pTE53 of a clinical isolate of Escherichia coli. Folia Microbiol 43:589–599
Cascioferro A, Boldrin F, Serafini A, Provvedi R, Palù G, Manganelli R (2010) Xer site-specific recombination, an efficient tool to introduce unmarked deletions into mycobacteria. Appl Environ Microbiol 76:5312–5316
Court DL, Sawitzke JA, Thomason LC (2002) Genetic engineering using homologous recombination. Annu Rev Genet 36:361–388
Cox MM, Layton SL, Jiang T, Cole K, Hargis BM, Berghman LR, Bottje WG, Kwon YM (2007) Scarless and site-directed mutagenesis in Salmonella enteritidis chromosome. BMC Biotechnol 7:59
Datsenko KA, Wanner BL (2000) Onestep inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97:6640–6645
Datta S, Costantino N, Zhou X, Court DL (2008) Identification and analysis of recombineering functions from Gram-negative and Gram-positive bacteria and their phages. Proc Natl Acad Sci USA 105:1626–1631
DeVito JA (2007) Recombineering with tolC as a selectable/counter-selectable marker: remodeling the rRNA operons of Escherichia coli. Nucleic Acids Res 36:e4
Doublet B, Douard G, Targant H, Meunier D, Madec JY, Cloeckaert A (2008) Antibiotic marker modifications of λ Red and FLP helper plasmids, pKD46 and pCP20, for inactivation of chromosomal genes using PCR products in multidrug resistant strains. J Microbiol Methods 75:359–361
Dutra BE, Sutera VA Jr, Lovett ST (2007) RecA-independent recombination is efficient but limited by exonucleases. Proc Natl Acad Sci USA 104:216–221
Ellis HM, Yu D, DiTizio T, Court DL (2001) High efficiency mutagenesis, repair, and engineering of chromosomal DNA using single-stranded oligonucleotides. Proc Natl Acad Sci USA 98:6742–6746
Filutowicz M, McEachern MJ, Helinski DR (1986) Positive and negative roles of an initiator protein at an origin of replication. Proc Natl Acad Sci USA 83:9645–9649
Fukiya S, Mizoguchi H, Mori H (2004) An improved method for deleting large regions of Escherichia coli K-12 chromosome using a combination of Cre/loxP and λ Red. FEMS Microbiol Lett 234:325–331
Gamper M, Kast P (2005) Strategy for chromosomal gene targeting in RecA-deficient Escherichia coli strains. Biotechniques 38:405–408
Gerlach RG, Jäckel D, Hölzer SU, Hensel M (2009) Rapid oligonucleotide-based recombineering of the chromosome of Salmonella enterica. Appl Environ Microbiol 75:1575–1580
Gutterson NI, Koshland DE Jr (1983) Replacement and amplification of bacterial genes with sequences altered in vitro. Proc Natl Acad Sci USA 80:4894–4898
Hamilton CM, Aldea M, Washburn BK, Babitzke P, Kushner SR (1989) New method for generating deletions and gene replacements in Escherichia coli. J Bacteriol 171:4617–4622
Hashimoto M, Ichimura T, Mizoguchi H, Tanaka K, Fujimitsu K, Keyamura K, Ote T, Yamakawa T, Yamazaki Y, Mori H, Katayama T, Kato JI (2005) Cell size and nucleoid organization of engineered Escherichia coli cells with a reduced genome. Mol Microbiol 55:137–149
Heermann R, Zeppenfeld T, Jung K (2008) Simple generation of site-directed point mutations in the Escherichia coli chromosome using Red®/ET® recombination. Microb Cell Fact 7:14
Herring CD, Blattner FR (2004) Conditional lethal amber mutations in essential Escherichia coli genes. J Bacteriol 186:2673–2681
Herring CD, Glasner JD, Blattner FR (2003) Gene replacement without selection: regulated suppression of amber mutations in Escherichia coli. Gene 311:153–163
Husseiny MI, Hensel M (2005) Rapid method for the construction of Salmonella enterica serovar typhimurium vaccine carrier strains. Infect Immun 73:1598–1605
Kang Y, Durfee T, Glasner JD, Qiu Y, Frisch D, Winterberg KM, Blattner FR (2004) Systematic mutagenesis of the Escherichia coli genome. J Bacteriol 186:4921–4930
Katashkina JI, Hara Y, Golubeva LI, Andreeva IG, Kuvaeva TM, Mashko SV (2009) Use of the λ Red-recombineering method for genetic engineering of Pantoea ananatis. BMC Mol Biol 10:34
Kim K, Kim KP, Choi J, Lim JA, Lee J, Hwang S, Ryu S (2010) Outer membrane proteins A (OmpA) and X (OmpX) are essential for basolateral invasion of Cronobacter sakazakii. Appl Environ Microbiol 76:5188–5198
Kolisnychenko V, Plunkett G III, Herring CD, Fehér T, Pósfai J, Blattner FR, Pósfai G (2002) Engineering a reduced Escherichia coli genome. Genome Res 12:640–647
Kowalczykowski SC, Dixon DA, Eggleston AK, Lauder SD, Rehrauer WM (1994) Biochemistry of homologous recombination in Escherichia coli. Microbiol Rev 58:401–465
Kuhlman TE, Cox EC (2010) Site-specific chromosomal integration of large synthetic constructs. Nucleic Acids Res 38:e92
Lee EC, Yu D, Martinez de Velasco J, Tessarollo L, Swing DA, Court DL, Jenkins NA, Copeland NG (2001) A highly efficient Escherichia coli-based chromosome engineering system adapted for recombinogenic targeting and subcloning of BAC DNA. Genomics 73:56–65
Lee DJ, Bingle LE, Heurlier K, Pallen MJ, Penn CW, Busby SJ, Hobman JL (2009) Gene doctoring: a method for recombineering in laboratory and pathogenic Escherichia coli strains. BMC Microbiol 9:252
Leenhouts K, Buist G, Bolhuis A, Berge TA, Kiel J, Mierau I, Dabrowska M, Venema G, Kok J (1996) A general system for generating unlabelled gene replacements in bacterial chromosomes. Mol Gen Genet 253:217–224
Leslie NR, Sherratt DJ (1995) Site-specific recombination in the replication terminus region of Escherichia coli: functional replacement of dif. EMBO J 14:1561–1570
Liang R, Liu J (2010) Scarless and sequential gene modification in Pseudomonas using PCR product flanked by short homology regions. BMC Microbiol 10:209
Link AJ, Phillips D, Church GM (1997) Methods for generating precise deletions and insertions in the genome of wild-type Escherichia coli: application to open reading frame characterization. J Bacteriol 179:6228–6237
Marinelli LJ, Piuri M, Swigoňová Z, Balachandran A, Oldfield LM, van Kessel JC, Hatfull GFBRED (2008) A simple and powerful tool for constructing mutant and recombinant bacteriophage genomes. PLoS ONE 3:e3957
Merlin C, McAteer S, Masters M (2002) Tools for characterization of Escherichia coli genes of unknown function. J Bacteriol 184:4573–4581
Murphy KC (1998) Use of bacteriophage λ recombination functions to promote gene replacement in Escherichia coli. J Bacteriol 180:2063–2071
Murphy KC, Campellone KG (2003) Lambda Red-mediated recombinogenic engineering of enterohemorrhagic and enteropathogenic E. coli. BMC Mol Biol 4:11
Murphy KC, Campellone KG, Poteete AR (2000) PCR-mediated gene replacement in Escherichia coli. Gene 246:321–330
Muyrers JPP, Zhang Y, Benes V, Testa G, Ansorge W, Stewart AF (2000) Point mutation of bacterial artificial chromosomes by ET recombination. EMBO Rep 1:239–243
Oppenheim AB, Rattray AJ, Bubunenko M, Thomason LC, Court DL (2004) In vivo recombineering of bacteriophage λ by PCR fragments and single-strand oligo nucleotides. Virology 319:185–189
Palmeros B, Wild J, Szybalski W, Le Borgne S, Hernández-Chávez G, Gosset G, Valle F, Bolivar F (2000) A family of removable cassettes designed to obtain antibiotic-resistance-free genomic modifications of Escherichia coli and other bacteria. Gene 247:255–264
Pósfai G, Koob M, Hradecná Z, Hasan N, Filutowicz M, Szybalski W (1994) In vivo excision and amplification of large segments of the Escherichia coli genome. Nucleic Acids Res 22:2392–2398
Pósfai G, Kolisnychenko V, Bereczki Z, Blattner FR (1999) Markerless gene replacement in Escherichia coli stimulated by a double-strand break in the chromosome. Nucleic Acids Res 27:4409–4415
Pósfai G, Plunkett G III, Fehér T, Frisch D, Keil GM, Umenhoffer K, Kolisnychenko V, Stahl B, Sharma SS, De Arruda M, Burland V, Harcum SW, Blattner FR (2006) Emergent properties of reduced-genome Escherichia coli. Science 312:1044–1046
Ranallo RT, Barnoy S, Thakkar S, Urick T, Venkatesan MM (2006) Developing live Shigella vaccines using λ Red recombineering. FEMS Immunol Med Microbiol 47:462–469
Rivero-Müller A, Lajic S, Huhtaniemi I (2007) Assisted large fragment insertion by Red/ET-recombination (ALFIRE)—an alternative and enhanced method for large fragment recombineering. Nucleic Acids Res 35:1–8
Rothstein R (1991) Targeting, disruption, replacement, and allele rescue: integrative DNA transformation in yeast. Methods Enzymol 194:281–301
Russell CB, Thaler DS, Dahlquist FW (1989) Chromosomal transformation of Escherichia coli recD strains with linearized plasmids. J Bacteriol 171:2609–2613
Serra-Moreno R, Acosta S, Hernalsteens JP, Jofre J, Muniesa M (2006) Use of the lambda Red recombinase system to produce recombinant prophages carrying antibiotic resistance genes. BMC Mol Biol 7:31
Sharan SK, Thomason LC, Kuznetsov SG, Court DL (2009) Recombineering: a homologous recombination-based method of genetic engineering. Nat Protoc 4:206–223
Swaminathan S, Ellis HM, Waters LS, Yu D, Lee EC, Court DL, Sharan SK (2001) Rapid engineering of bacterial artificial chromosomes using oligonucleotides. Genesis 29:14–21
Swingle B, Markel E, Costantino N, Bubunenko MG, Cartinhour S, Court DL (2010a) Oligonucleotide recombination in Gram-negative bacteria. Mol Microbiol 75:138–148
Swingle B, Bao Z, Markel E, Chambers A, Cartinhour S (2010b) Recombineering using recTE from Pseudomonas syringae. Appl Environ Microbiol 76:4960–4968
Tischer BK, Von Einem J, Kaufer B, Osterrieder N (2006) Two-step Red-mediated recombination for versatile high-efficiency markerless DNA manipulation in Escherichia coli. Biotechniques 40:191–197
Uzzau S, Figueroa-Bossi N, Rubino S, Bossi L (2001) Epitope tagging of chromosomal genes in Salmonella. Proc Natl Acad Sci USA 98:15264–15269
van Kessel JC, Marinelli LJ, Hatfull GF (2008) Recombineering mycobacteria and their phages. Nat Rev Microbiol 6:851–857
Vizváryová M, Stuchlík S, Grones J, Mačor M, Turňa J (1999) Escherichia coli strain with a deletion of the chromosomal ampC gene marked with TcR, suitable for production of penicillin G acylase. Folia Microbiol 44:625–628
Wang S, Zhao Y, Leiby M, Zhu J (2009) A new positive/negative selection scheme for precise BAC recombineering. Mol Biotechnol 42:110–116
Warming S, Costantino N, Court DL, Jenkins NA, Copeland NG (2005) Simple and highly efficient BAC recombineering using galK selection. Nucleic Acids Res 33:1–12
White AP, Allen-Vercoe E, Jones BW, DeVinney R, Kay WW, Surette MG (2007) An efficient system for markerless gene replacement applicable in a wide variety of enterobacterial species. Can J Microbiol 53:56–62
Yu D, Ellis HM, Lee EC, Jenkins NA, Copeland NG, Court DL (2000) An efficient recombination system for chromosome engineering in Escherichia coli. Proc Natl Acad Sci USA 97:5978–5983
Yu BJ, Kang KH, Lee JH, Sung BH, Kim MS, Kim SC (2008) Rapid and efficient construction of markerless deletions in the Escherichia coli genome. Nucleic Acids Res 36:e84
Zhang Y, Buchholz F, Muyrers JP, Stewart AF (1998) A new logic for DNA engineering using recombination in Escherichia coli. Nat Genet 20:123–128
Zhou QM, Fan DJ, Xie JB, Liu CP, Zhou JM (2010) A method for generating precise gene deletions and insertions in Escherichia coli. World J Microbiol Biotechnol 26:1323–1329
Acknowledgments
This publication is the result of project implementation: development of the center of excellence for utilization of information on bio-macromolecules in disease prevention and in improvement of quality of life (ITMS 26240120027) supported by the Research & Development Operational Program funded by the ERDF. The work was also supported by the grant of Slovak grant agency VEGA 1/0344/10.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Madyagol, M., Al-Alami, H., Levarski, Z. et al. Gene replacement techniques for Escherichia coli genome modification. Folia Microbiol 56, 253–263 (2011). https://doi.org/10.1007/s12223-011-0035-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-011-0035-z