Abstract
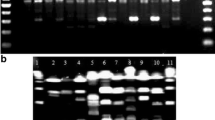
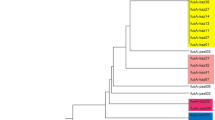
Eleven type strains and 24 Lactobacillus isolates, preliminarily classified to the species due to phenotypic features, were investigated. Standard methods of identification with species-specific PCRs and typing with PFGE (with ApaI, NotI and SmaI restriction enzymes) allowed us to distinguish 16 unique strains belonging to 5 species (L. acidophilus, L. delbrueckii ssp. bulgaricus, L. plantarum, L. rhamnosus, L. salivarius). Alternative approach with 16S–23S rDNA ARDRA identification (with merely two restrictases, BsuRI and TaqI) and PCR-based typing (RAPD with two random- and rep-PCR with (GTG)5 primers) showed to be more discriminative, i.e. 21 unique strains were classified in the same species as above. As a result, 7 out of 24 phenotypically species-assigned isolates were reclassified. The alternative procedure of rapid identification and typing of Lactobacillus isolates appeared to be equally effective and shortened from 1 week to 2–3 d (in comparison to the standard methods).
Similar content being viewed by others
Abbreviations
- ARDRA:
-
amplified ribosomal DNA restriction analysis
- CFU:
-
colony forming units
- DSM:
-
Deutsche Sammlung von Mikroorganismen
- PCR(s):
-
polymerase chain reaction(s)
- PFGE:
-
pulsed-field gel electrophoresis
- RAPD:
-
randomly amplified polymorphic DNA
- rep-PCR:
-
PCR of repetitive bacterial DNA
- UPGMA:
-
unweighted pair group method with arithmetic mean
References
Bouton Y., Guyot P., Beuvier E., Tailliez P., Grappin R.: Use of PCR-based methods and PFGE for typing and monitoring homofermentative lactobacilli during Comte’ cheese ripening. Internat.J.Food Microbiol.76, 27–38 (2002).
Collado M.C., Hernandez M.: Identification and differentiation of Lactobacillus, Streptococcus and Bifidobacterium species in fermented milk products with bifidobacteria. Microbiol.Res.162, 86–92 (2007).
Coudeyras S., Marchandin H., Fajon C., Forestier C.: Taxonomic and strain-specific identification of the probiotic strain Lactobacillus rhamnosus 35 within the Lactobacillus casei group. Appl.Environ.Microbiol.74, 2679–2689 (2008).
De Man J.C., Rogosa M., Sharpe M.E.: A medium for the cultivation of lactobacilli. J.Appl.Bacteriol.23, 130–135 (1960).
Gevers D., Huys G., Swings J.: Applicability of rep-PCR fingerprinting for identification of Lactobacillus species. FEMS Microbiol. Lett.205, 31–36 (2001).
Giraffa G., Andrighetto C., Antonello C., Gatti M., Lazzi C., Marcazzan G., Lombardi A., Neviani E.: Genotypic and phenotypic diversity of Lactobacillus delbrueckii subsp. lactis strains of dairy origin. Internat.J.Food Microbiol.91, 129–139 (2004).
Guan L.L., Hagen K.E., Tannock G.W., Korver D.R., Fasenko G.M., Allison G.E.: Detection and identification of Lactobacillus species in crops of broilers of different ages by using PCR-denaturing gradient gel electrophoresis and amplified ribosomal DNA restriction analysis. Appl.Environ.Microbiol.69, 6750–6757 (2003).
Hamad S.H., Dieng M.C., Ehrmann M.A., Vogel R.F.: Characterization of the bacterial flora of Sudanese sorghum flour and sorghum sourdough. J.Appl.Microbiol.83, 764–770 (1997).
Jacobsen C.N., Rosenfeldt Nielsen V., Hayford A.E., Müller P.L., Michalsen K.F., Párregaard A., Sandström B., Tvede M., Jakobsen M.: Screening of probiotic activities of forty-seven strains of Lactobacillus spp. by in vitro techniques and evaluation of the colonization ability of five selected strains in humans. Appl.Environ.Microbiol.65, 4949–4956 (1999).
Křížová J., Španová A., Rittich B.: RAPD and rep-PCR fingerprinting for characterization of Bifidobacterium species. Folia Microbiol.53, 99–104 (2008).
Leser T.D., Lindecrona R.H., Jensen T.K., Jensen B.B., Müller K.: Changes in bacterial community structure in the colon of pigs fed different experimental diets and after infection with Brachyspira hyodysenteriae. Appl.Environ.Microbiol.66, 3290–3296 (2000).
Markiewicz L., Biedrzycka E.: Identification of Lactobacillus and Bifidobacterium species by PCR applied to quality control of fermented dairy beverages. Polish J.Food Nutr.Sci.55, 359–365 (2005).
Markiewicz L., Wasilewska E., Bielecka M.: Identification of Lactobacillus strains present in fermented dairy products and their differentiation using molecular methods. Polish J.Food Nutr.Sci.58, 251–256 (2008).
Markiewicz L., Biedrzycka E., Wasilewska E., Bielecka M.: Differentiation of strains identified as Bifidobacterium animalis subsp. lactis. Acta Alim.38, 293–301 (2009).
Mooreira J.L.S., Mota R.M., Horta M.F., Teixeira S.M.R., Neumann E., Nicoli J.R., Nunes Á.C.: Identification to the species level of Lactobacillus isolated in probiotic prospecting studies of human, animal or food origin by 16S–23S rRNA restriction profiling. BMC Microbiol. DOI:10.1186/1471-2180-5-15 (2005).
Müller M.R.A., Ehrmann M.A., Vogel R.F.: Multiplex PCR for the detection of Lactobacillus pontis and two related species in a sourdough fermentation. Appl.Environ.Microbiol.66, 2113–2116 (2000).
Roy D., Sirois S., Vincent D.: Molecular discrimination of lactobacilli used as starter and probiotic cultures by amplified ribosomal DNA restriction analysis. Curr.Microbiol.42, 282–289 (2001).
Song Y.-L., Kato N., Liu C.-X., Matsumiya Y., Kato H., Watanabe K.: Rapid identification of 11 human intestinal Lactobacillus species by multiplex PCR assays using group- and species-specific primers derived from the 16S-23S rRNA intergenic spacer region and its flanking 23S rRNA. FEMS Microbiol.Lett.187, 167–173 (2000).
Torriani S., Zapparoli G., Dellaglio F.: Use of PCR-based methods for rapid differentiation of Lactobacillus delbrueckii subsp. bulgaricus and Lactobacillus delbrueckii subsp. lactis. Appl.Environ.Microbiol.65, 4351–4356 (1999).
Tynkkynen S., Satokari R., Saarela M., Mattila-Sandholm T., Saxelin M.: Comparison of ribotyping, randomly amplified polymorphic DNA analysis, and pulsed-field gel electrophoresis in typing of Lactobacillus rhamnosus and L. casei strains. Appl.Environ.Microbiol.65, 3908–3914 (1999).
Ventura M., Callegari M.L., Reniero R., Zink R.: Molecular microbial analysis of Bifidobacterium isolates from different environments by the species-specific amplified ribosomal DNA restriction analysis (ARDRA). FEMS Microbiol.Ecol.36, 113–121 (2001).
Walter J., Tannock G.W., Tilsala-Timisjarvi A., Rodtong S., Loach D.M., Munro K., Alatossava T.: Detection and identification of gastrointestinal Lactobacillus species by using denaturing gradient gel electrophoresis and species-specific PCR primers. Appl.Environ.Microbiol.66, 297–303 (2000).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Markiewicz, L.H., Biedrzycka, E., Wasilewska, E. et al. Rapid molecular identification and characteristics of Lactobacillus strains. Folia Microbiol 55, 481–488 (2010). https://doi.org/10.1007/s12223-010-0080-z
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-010-0080-z