Abstract
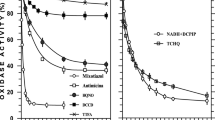
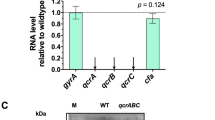
Membrane fragments of two mutant strains of Paracoccus denitrificans genetically modified in the bc 1 complex have been studied for comparison of enzymic activities of succinate-cytochrome-c reductase and its components, viz. succinate dehydrogenase (Complex II) and ubiquinol-cytochrome-c reductase (Complex III) and their response to changes in concentration of succinate, cytochrome c, ionic strength, pH, temperature and sensitivity to antimycin A. The mutants synthesized and assembled the b and c hemes in the ratio characteristic for the wild type strain. The mutant strain M 71 expressing the truncated copy of cytochrome c 1 (devoid of a stretch of 150 mainly acidic amino acids) was less sensitive to increasing concentration of cytochrome c and changes in ionic strength of the medium, but maintained the original affinity to succinate and sensitivity to antimycin A. The mutant strain M 36 with an overexpressed bc 1 content showed the highest response to changes in ionic strength and physical parameters, exhibited the lowest turnover number values with succinate-cytochrome-c reductase, but positively affected the succinate dehydrogenase. In view of the interaction of the redox components in native membranes the functional analyses of separated Complexes II and III should be regarded with caution.
Similar content being viewed by others
Abbreviations
- cyt:
-
cytochrome
- DCIP:
-
2,6-dichloroindophenol
- ISP(s):
-
iron-sulfur cluster(s) (Fe-S protein)
- k (1/s):
-
first-order rate constant
- k cat :
-
turnover number value(s)
- M 36:
-
mutant strain G440/pEG436
- M 71:
-
mutant strain G440/pEG471
- CYO:
-
cytochrome-c oxidase (EC 1.9.3.1)
- QCR:
-
ubiquinol-cytochrome-c reductase (EC 1.10.2.2)
- PMS:
-
phenazine methosulfate
- Q:
-
ubiquinone
- QH2 :
-
ubiquinol
- ROS:
-
reactive oxygen species
- RSD:
-
relative standard deviation
- Suc:
-
succinate
- WT:
-
wild-type (strain)
- SCR:
-
succinate-cytochrome-c reductase (see the text)
- SDH:
-
succinate dehydrogenase (EC 1.3.99.1)
References
Ackrell B.A.: Cytopathies involving mitochondrial complex II. Mol.Aspects Med.23, 369–384 (2002).
Baysal B.E., Ferrell R.E., Willett-Brozik J.E., Lawrence E.C., Myssiorek D., Bosch A., Van Der Mey A., Taschner P.E., Rubinstein W.S., Myers E.N., Richard C.W., Cornelisse C.J., Devilee P., Devlin B.: Mutations in SDHD, a mitochondrial complex II gene, in hereditary paraganglioma. Science287, 848–851 (2000).
Cecchini G.: Function and structure of complex II of the respiratory chain. Ann.Rev.Biochem.72, 77–109 (2003).
Chen Q., Vazgues E.J., Moghaddas S., Hoppel C.L., Lesnefsky E.L.: Production of reactive oxygen species by mitochondria: central role of complex III. J.Biol.Chem.278, 36027–36031 (2003).
Dadák V., Malá G., Kučera I.: Kinetic characterization of succinate dehydrogenase (succinate:ubiquinone oxidoreductase) from Paracoccus denitrificans. Biológia (Bratislava)43, 1077–1085 (1988).
Dadák V., Holík M.: Electrostatic attraction between cytochrome bc1 and cytochrome c affects kinetics of cytochrome c1 reduction. Biochemistry (Moscow)73, 870–880 (2008).
Dröse S., Brandt U.: The mechanism of mitochondrial superoxide production by the cytochrome bc1 complex. J.Biol.Chem.283, 21649–21654 (2008).
Gerhus E., Steinrucke P., Ludwig B.: Paracoccus denitrificans gene replacement mutants. J.Bacteriol.172, 2392–2400 (1990).
Hederstedt L.: Succinate:quinone oxidoreductase in the bacteria Paracoccus denitrificans and Bacillus subtilis. Biochim.Biophys. Acta1553, 74–83 (2002).
Janzon J., Eichhorn A.C., Ludwig B., Malatesta F.: Electron transfer kinetics between soluble modules of Paracoccus denitrificans cytochrome c1 and its physiological redox partners. Biochim.Biophys.Acta1777, 250–259 (2008).
John P., Whatley F.R.: Paracoccus denitrificans DAVIS (Micrococcus denitrificans BEIJERINCK) as a mitochondrion. Adv.Bot.Res.4,51–115 (1977).
Kurowski B., Ludwig B.: The genes of the Paracoccus denitrificans bc1 complex. J.Biol.Chem.262, 13805–13811 (1987).
Ludwig B.: Cytochrome-c oxidase from Paracoccus denitrificans. Methods Enzymol.126, 153–159 (1986).
Oyedotun K.S., Sit C.S., Lemire B.D.: The Saccharomyces cerevisiae succinate dehydrogenase does not require heme for ubiquinone reduction. Biochim.Biophys.Acta1767, 1436–1445 (2007).
Pennoyer J.D., Ohnishi T., Trumpower B.L.: Purification and properties of succinate-ubiquinone oxidoreductase complex from Paracoccus denitrificans. Biochim.Biophys.Acta935, 195–207 (1988).
Schägger H.: Respiratory chain supercomplexes of mitochondria and bacteria. Biochim.Biophys.Acta1555, 154–159 (2002).
Scholes P.B., Smith L.: Composition and properties of the membrane-bound respiratory chain system of Micrococcus denitrificans. Biochim.Biophys.Acta153, 363–375 (1968).
Yang X., Trumpower B.L.: Purification of a three-subunit ubiquinol-cytochrome-c oxidoreductase complex from Paracoccus denitrificans. J.Biol.Chem.261, 12282–12289 (1986).
Yankovskaya V., Horsefield R., Tornroth S., Luna-chavez C., Miyoshi H., Leger C., Byrne B., Cecchini G., Iwata S.: Architecture of succinate dehydrogenase and reactive oxygen species generation. Science299, 700–704 (2003).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dadák, V., Dudák, J. & Zbořil, P. Electron transfer in Paracoccus denitrificans with the modified fbc operon. Folia Microbiol 54, 475–482 (2009). https://doi.org/10.1007/s12223-009-0067-9
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-009-0067-9