Abstract
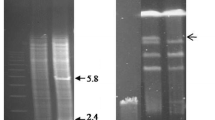
Streptomyces aureofaciens B96 produces several intra- and extracellular enzymes with deoxyribonuclease activity. According to the sequence of the previously published gene exoSc from S. coelicolor A3(2), the DNA sequence from S. aureofaciens B96 was amplified, cloned and expressed in E. coli. The protein product of exoSa gene, recExoSa, was also an exonuclease with DNAase and 5′-phosphomonoesterase activities at optimum temperature 37 °C and pH 8.0. It degraded only linear DNA (chromosomal, double-stranded and single-stranded) and linear plasmid DNA from both ends, with a preference to blunt ends in comparison with overhang ends. The purified enzyme exhibited no RNAase activity. Both exoSc and exoSa genes were interrupted by the apramycin resistance gene; constructed fragments were transformed into particular streptomyces protoplasts. Mutation caused by exoSa disruption in S. aureofaciens chromosome and mutation by interrupted exoSc in S. coelicolor were lethal.
Similar content being viewed by others
Abbreviations
- ccc:
-
covalently closed circular
- MM:
-
minimal medium
- ss:
-
single-stranded
- ds:
-
double-stranded
- mMM:
-
modified MM
References
Blondelet-Rouault M.H., Weiser J., Lebrihi A., Branny P., Pernodet J.L.: Antibiotic resistance gene cassettes derived from the Ω interposon for use in E. coli and Streptomyces. Gene 190, 315–317 (1997).
Brnáková Z., Godány A., Timko J.: An extracellular endodeoxyribonuclease from Streptomyces aureofaciens. Biochim.Biophys.Acta 1721, 116–123 (2005).
Brnáková Z., Godány A., Timko J.: An exodeoxyribonuclease from Streptomyces coelicolor: expression, purification and biochemical characterization. Biochim.Biophys.Acta 1770, 630–637 (2007).
Cal S., Aparicio J.F., De los Reyes-Gavilan C.G., Nicieza R.G., Sanchez J.: A novel exocytoplasmic endonuclease from Streptomyces antibioticus. Biochem.J. 306, 93–100 (1995).
Chase J.W., Richardson C.C.: Exonuclease VII of Escherichia coli. Mechanism of action. J.Biol.Chem. 249, 4553–4561 (1974).
Corpet F.: Multiple sequence alignment with hierarchical clustering. Nucl.Acids Res. 16, 10881–10890 (1988).
Fernández M., Sanchez J.: Nuclease activities and cell death processes associated with the development of surface cultures of Streptomyces antibioticus ETH 7451. Microbiology 148, 405–412 (2002).
Karoonuthaisiri N., Weaver D., Huang J., Cohen S.N., Kao C.M.: Regional organization of gene expression in Streptomyces coelicolor. Gene 353, 53–66 (2005).
Kieser T., Bibb M.J., Buttner M.J., Chater K.F., Hopwood D.A.: Practical Streptomyces Genetics, 2nd ed. John Innes Foundation, Norwich (UK) 2000.
Kirby R., Gan T.K., Hunter I., Herron P., Tilley E.: The genome of Streptomyces rimosus subsp. rimosus shows a novel structure compared to other Streptomyces using DNA/DNA microarray analysis. Antonie van Leeuwenhoek 94, 173–186 (2008).
Kitani S., Hoshika M., Nihira T.: Disruption of sscR encoding a γ-butyrolactone autoregulator receptor in Streptomyces scabie NBRC 12914 affects production of secondary metabolites. Folia Microbiol. 53, 115–124 (2008).
Kuwayama H., Obara S., Morio T., Katoh M., Urushihara H., Tanaka Y.: PCR-mediated generation of a gene disruption construct without the use of DNA ligase and plasmid vectors. Nucl.Acids Res. 30, e2 (2000).
Laemmli U.K.: Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685 (1970).
MacNeil D.J., Gewain K.M., Ruby C.L., Dezeny G., Gibbons P.H., MacNeil T.: Analysis of Streptomyces avermitilis genes required for avermectin biosynthesis utilising a novel integration vector. Gene 111, 61–68 (1992).
Manteca A., Fernández M., Sanchez J.: Cytological and biochemical evidence for an early cell dismantling event in surface cultures of Streptomyces antibioticus. Res.Microbiol. 157, 143–152 (2006).
Miertzschke M., Greiner-Stöffele T.: The xthA gene product of Archaeoglobus fulgidus is an unspecific DNase. Eur.J.Biochem. 270, 1838–1849 (2003).
Miguélez E.M., Hardisson C., Manzanal M.B.: Hyphal death during colony development in Streptomyces antibioticus: morphological evidence for the existence of a process of cell deletion in a multicellular prokaryote. J.Cell Biol. 145, 515–525 (1999).
Miguélez E.M., Hardisson C., Manzanal M.B.: Streptomycetes: a new model to study cell death. Internat.Microbiol. 3, 153–158 (2000).
Nazarov V., Žatkovič P., Godány A.: New shuttle promoter-probe vectors for E. coli and Streptomycetes. Biotechnol.Lett. 12, 639–644 (1990).
Nicieza R.G., Huergo J., Connolly B.A., Sanchez J.: Purification, characterization, and role of nucleases and serine proteases in Streptomyces differentiation. J.Biol.Chem. 274, 20366–20375 (1999).
Patil N.S., Deshmukh S.S., Shankar V.: Extracellular nuclease from a thermophile, Streptomyces thermonitrificans: production, purification and partial characterization of — double strand preferential — deoxyribonuclease activity. Proc.Biochem. 40, 1271–1278 (2005).
Salas-Pacheco J., Setlow B., Setlow P., Pedraza-Reyes M.: Role of the Nfo (YqfS) and ExoA apurinic/apyrimidimic endonucleases in protecting Bacillus subtilis spores from DNA damage. J.Bacteriol. 187, 7374–7381 (2005).
Sambrook J., Fritsch E.F., Maniatis T.: Molecular Cloning: a Laboratory Manual, 2nd ed. Cold Spring Harbor Laboratory, New York 1989.
Studier F.W., Moffatt B.A.: Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J.Mol.Biol. 189, 113–130 (1986).
Wildermuth H.: Development and organization of the aerial mycelium in Streptomyces coelicolor. J.Gen.Microbiol. 60, 43–50 (1970).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brnáková, Z., Godány, A. & Timko, J. Characterization and disruption of exonuclease genes from Streptomyces aureofaciens B96 and S. coelicolor A3(2). Folia Microbiol 54, 97–104 (2009). https://doi.org/10.1007/s12223-009-0014-9
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-009-0014-9