Abstract
In-cell NMR, i.e., NMR spectroscopy applied to studying specific macromolecules within living cells, is becoming the technique of choice for the structural and mechanistic description of proteins and nucleic acids within increasingly complex cellular environments, as well as of the temporal evolution of biological systems over a broad range of timescales. Furthermore, in-cell NMR has already shown its potentialities in the early steps of drug development. In this Perspective, we report some of the most recent methodological advancements and successful applications of in-cell NMR spectroscopy, focusing particularly on soluble proteins. We show how the combination of the atomic-level characterization of NMR with its application to a cellular context can provide crucial insights on cellular processes and drug efficacy with unprecedented level of detail. Finally, we discuss the main challenges to overcome and share our vision of the future developments of in-cell NMR and the applications that will be made possible.
Similar content being viewed by others
1 Introduction
Biological systems are highly complex machineries. Understanding their inner workings is critical when developing therapeutic approaches to treat pathologies. To this aim, the structure and mechanism of their molecular components need to be investigated at atomic resolution. In the past 70 years, Structural Biology has been instrumental for understanding the structure and dynamics of biological macromolecules, determining how they work, and providing the necessary knowledge to develop more effective and specific drugs. However, structural and functional characterization is usually done in vitro on isolated macromolecules, therefore far from their physiological context. A holy grail of modern Structural Biology is indeed the ability to investigate at atomic resolution macromolecules in their native environment, where their functional interactions are preserved. Among the existing structural biology techniques, X-ray crystallography, Nuclear Magnetic Resonance (NMR) spectroscopy and cryo-electron microscopy, NMR stands out as the only one able to obtain information on macromolecular structure, kinetics and dynamics at the atomic level in solution. Even more importantly, it can do so in a nondestructive manner, thanks to the low energy of the electromagnetic radiations employed: radio waves (~ 10–6 eV) have energies much lower than the thermal energy at room temperature (~ 10–2 eV), unlike X-rays (~ 104 eV) and accelerated electrons (~ 105 eV). For the above reasons, NMR spectroscopy has since its birth been constantly applied to living cells to study their chemical composition. However, NMR is intrinsically insensitive compared to other spectroscopic techniques; hence, its application has been historically limited to abundant molecules, such as some metabolites present at high concentration in cells. Continuous progresses in the NMR hardware and in the design of efficient pulse sequences have progressively increased the sensitivity of the technique, thus making possible its application to study specific macromolecules of interest within the cellular environment. The latter approach, called in-cell NMR, was first demonstrated on proteins recombinantly expressed in Escherichia coli (Serber et al. 2001), and in the following two decades has evolved, slowly but steadily, into a small branch of biomolecular NMR that actively develops novel approaches to provide structural and mechanistic insights on proteins and nucleic acids within increasingly complex cellular environments. In addition to structural analysis, in-cell NMR can investigate the temporal evolution of biological systems over a broad range of timescales. Thanks to this unique feature, the approach has been watched with interest by the Structural Biology community even while, in the meantime, huge advancements in resolution and sensitivity of cryo-electron microscopy and tomography sparked the “resolution revolution” (Kühlbrandt 2014). The developments and applications of in-cell NMR spectroscopy have been the subject of many reviews and opinion articles, including our own (Kang 2019; Luchinat et al. 2022; Luchinat and Banci 2022; Siegal and Selenko 2019; Theillet 2022; Theillet and Luchinat 2022). Such an extensive coverage has even raised some criticism, that it might create too high expectation in the scientific community, but clearly testifies to the high interest in the methodology. In this perspective, we summarize some recent key advancements and promising applications of in-cell NMR spectroscopy, with particular emphasis on the studies of soluble proteins in human cells, and we share our vision on the challenges and opportunities that the approach will meet in the near future.
2 Adding the fourth dimension to Cellular Structural Biology
As mentioned above, a key advantage of NMR applied to living cells/organisms is the possibility to study time-dependent phenomena over a broad range of timescales: from molecular motions occurring in the nanosecond scale to cellular processes which take hours or days. NMR spectroscopy can conveniently study phenomena in the nanoseconds–seconds range by encoding the time information in the amplitude or frequency of the detected nuclear spins. Quantitative information on the dynamics of molecules such as tumbling, internal motions, chemical exchange with the solvent and transient interactions with other molecules is then obtained from subsequent analysis of the signal amplitudes/shifts (Kovermann et al. 2016). Such experiments are in general applicable to macromolecules in living cells, and they have been instrumental to understand how protein folding and dynamics are affected by weak interactions with the cellular environments (Li and Liu 2013; Monteith and Pielak 2014; Theillet et al. 2016). In fact, understanding how such interactions, termed quinary structure, affect protein function has only recently been made possible by in-cell NMR (Majumder et al. 2015; Monteith et al. 2015; Mu et al. 2017).
Phenomena occurring in the seconds–days range are best studied as they happen by time-resolved NMR. In this approach, NMR spectra are continuously recorded providing a ‘movie’ from which the kinetics of functional processes can be derived from the change of signal amplitude/shift as a function of time. From the spectroscopy side, time-resolved acquisition of 1D NMR spectra is straightforward, and several methods have been developed to increase the time-resolution of multidimensional NMR experiments without sacrificing sensitivity (Gołowicz et al. 2020). When applied to cells, the challenge arises from the need to keep them alive and metabolically stable for hours or even days in a non-ideal condition, i.e., densely packed in a narrow glass tube placed inside the NMR spectrometer. In these conditions, cells rapidly undergo starvation and hypoxia, causing changes of intracellular pH, chemical composition and redox homeostasis, which prevent the correct interpretation of the results. Moreover, dead cells easily rupture and release the molecule of interest, thus invalidating the experiment (Barnes and Pielak 2011).
To overcome this limitation, NMR bioreactors have been introduced. An NMR bioreactor usually consists of a flow cell shaped like a normal NMR tube in which the cells are confined, e.g., by hydrogel encapsulation, and perfused with a continuous flow of medium that provides fresh nutrients and oxygen, and removes the by-products of cell metabolism. Several bioreactor designs have been proposed for in-cell NMR applications, which fit modern 5-mm NMR cryogenic probes and can be applied to both bacteria and insect/mammalian cells (Barbieri and Luchinat 2021; Burz et al. 2019; Cerofolini et al. 2019; Kubo et al. 2013; Sharaf et al. 2010). Such devices have enabled real-time monitoring of cell metabolism (Carvalho et al. 2019; Hertig et al. 2021), intracellular protein–ligand interactions (Breindel et al. 2020; Luchinat et al. 2020, 2021b) and protein redox state regulation (Mochizuki et al. 2018), while keeping the cells viable for up to three days (Fig. 1). Despite these advancements, current NMR bioreactors are still experimental, custom-made devices. Future NMR bioreactors should adhere to community-defined minimum specifications, such as the ability to control medium composition, pH and dissolved O2/CO2 (Hertig et al. 2021), and to ensure uniform cell perfusion, and should be further engineered, e.g., to allow fast injection of reagents independent of the medium flow. Furthermore, to study physio/pathologically relevant tissue culture models, biomimetic hydrogels (Prince and Kumacheva 2019) and/or 3D scaffolds (Chung et al. 2020) similar to those used in regenerative medicine should be implemented, as they are better suited for cell adhesion and tissue growth with respect to agarose, alginate and other polymers commonly used. Finally, in long term, miniaturization and parallelization of flow systems might allow simultaneous detection of multiple cell samples, and could combine the benefits of a perfusion system with the higher throughput of interleaved metabolic analysis of ‘static’ cell samples, which has been recently demonstrated (Alshamleh et al. 2020).
NMR Bioreactor. A From left to right: cells embedded in hydrogel, shown before (top) and after (bottom) 72 h of NMR acquisition under flow conditions, are confined in the active volume of a flow unit, where they are perfused by a constant flow of nutrients, allowing continuous acquisition of in-cell NMR data. B Real-time in-cell.1H NMR spectra (top) show signal intensity changes over time and can be analyzed to monitor intracellular events, such as protein-drug interactions, in real time (bottom)
3 The strive for physiologically relevant cell models
The main rationale for studying a biomolecule of interest in cells is that the cellular environment is preserved. When studying non-specific effects, such as macromolecular crowding or electrostatic interactions, the exact composition of the cellular milieu might not be critical. Such studies have therefore been mostly carried out in E. coli, in which a protein can be recombinantly expressed at high levels. However, for functional studies, the inside of a bacterial cell is clearly very different from that of yeast, and both of them differ from a human cell (Barbieri et al. 2015). Hence, when investigating a protein involved in functional interactions, one may want to mimic as much as possible the environment of the source organism.
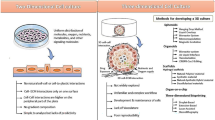
Much effort has been put into enabling NMR studies of macromolecules, both proteins and nucleic acids, in eukaryotic and eventually human-derived cells. Because NMR is an intrinsically insensitive technique, an additional challenge arises: the physiological abundance of most macromolecules is not sufficient, and must be increased to concentrations above ~ 5 µM, regardless of the cell type. Furthermore, to avoid interference from the other molecules present in the cell, isotope labeling of the target molecule becomes compulsory. In bacteria and yeast, these requirements are met when a protein is recombinantly expressed in isotope-labeled medium. In other eukaryotes, however, protein expression is less straightforward, therefore methods have been developed to deliver a protein or nucleic acid into the cells. Exogenous molecules can be microinjected in Xenopus laevis oocytes, owing to their large size (Sakai et al. 2006; Selenko et al. 2006), whereas in human cells, isotope-labeled macromolecules can be delivered to NMR-compatible levels using cell-penetrating peptides (Inomata et al. 2009), pore-forming toxins (Ogino et al. 2009), or by electroporation (Dzatko et al. 2018; Theillet et al. 2016). Alternatively, it has been shown that proteins can be directly overexpressed in insect and human cultured cells (Fig. 2). In insect, this was made possible by employing the highly efficient baculovirus system to infect the cells with coding DNA (Hamatsu et al. 2013), while in human cells an expression approach relying on an efficient and cost-effective DNA transfection was pioneered by our research group (Banci et al. 2013). Delivery and expression strategies have different advantages and shortcomings, which make them complementary. Delivery is currently the only viable approach for in-cell NMR of nucleic acids; it provides excellent isotope-labeling selectivity and, electroporation in particular, has been applied to different human cell lines. On the other hand, protein delivery is quite labor intensive: many cells and large amounts of highly concentrated pure protein are required for each sample preparation. Furthermore, each new protein requires extensive optimization to ensure efficient delivery, and many proteins fail altogether due to aggregation at high concentration, interactions with the plasma membrane preventing delivery or denaturation during electroporation. Conversely, protein expression is much less protein dependent: many different proteins can be overexpressed at high levels, and once the expression is established, large sets of mutants can be easily investigated. Furthermore, it is much faster and less labor intensive than protein delivery, as protein purification is not required, and a small-scale culture is sufficient for each sample. On the other hand, the approach is not suited to nucleic acids, high-level protein overexpression is limited to specific insect (Sf9) and human (HEK293T) cell lines, cell metabolism causes partial isotope labeling of other cellular components resulting in background signals in the NMR spectra, and no artificial protein chemical modifications (e.g., spin labels) are possible.
Protein expression in human cells. From left to right: human cells are first seeded in a culture flask and grown in unlabeled medium for 24 or 48 h; subsequently, the cells are transiently transfected with the genes of interest; protein expression is carried out for 48 h in isotope-labeled medium; cells are collected and analyzed by in-cell NMR
Set aside the specific advantages of each strategy, all methods described above have been optimized towards reaching high enough levels of macromolecule to allow NMR detection. However, the ultimate objective of any in-cell NMR approach should be to reproduce as much as possible the real-life cellular environment. In this respect, the intracellular levels of the investigated molecule should not be increased to the point that its function (or its interaction with the environment) is perturbed, as in that case the very reason for performing in-cell NMR would be lost. Therefore, in general, an important challenge for future applications of in-cell NMR approaches will be to make possible the detection of lower, close-to-physiological levels of the molecule of interest. The NMR bioreactor (see the previous section) already provides a means to compensate for lower levels, by increasing the overall acquisition time. In the long term, efforts should also focus towards increasing the raw sensitivity of NMR applied to biological samples (see the next section). With higher sensitivity, more approaches for both delivery and expression that would fail today will become viable. In this respect, we believe that direct protein expression holds greater potential, as it ensures that the protein undergoes native-like steps of biosynthesis, folding, cofactor binding and further maturation/translocation to specific cellular compartments. Once the threshold for detectability is lowered, advanced genome editing methods already widely adopted by the cell biology community, that allow introducing and/or replacing DNA elements at specific positions in the host genome of cultured human cells, will be applicable to NMR studies (Li et al. 2020). We predict that the combination of stable transfection and inducible protein expression will make possible to obtain protein expression at homogeneous, NMR-accessible levels in physio/pathologically relevant cell culture models, which could be further manipulated, e.g., by inducing the formation of 3D cultures in the form of spheroids and eventually organoids, that better reproduce the behavior of normal and cancerous tissues (Rossi et al. 2018; Zanoni et al. 2020).
4 The ultimate challenge: making the invisible… visible
As discussed above, one main limitation of NMR spectroscopy is its intrinsic poor sensitivity. To further complicate things, large molecules in solution tumble much slower than small molecules. Their slower motion increases the rate of nuclear spin transverse relaxation, causing extensive line broadening, which gets worse with the molecular size and does not improve at higher fields. In cells, if the observed protein interacts with other cellular components—which is often the case—the relaxation broadening increases due to the fact that the average tumbling rate of the protein is decreased, irrespective of its molecular size. In case of interactions with abundant and/or large partners, the effect is exacerbated to the point that NMR detection is no longer possible.
NMR experiments tailored for the detection of large macromolecules in vitro, which exploit transverse cross-relaxation phenomena (TROSY- and CRINEPT-type NMR experiments (Riek et al. 2000)), can improve in-cell spectral resolution and sensitivity at high fields (Luchinat et al. 2021a; Majumder et al. 2015), but would still fail in the case of large intracellular complexes. In vitro, extensive deuteration of large molecules substantially decreases the transverse relaxation of the remaining 1H nuclei, and could offer a solution for interacting proteins in cells. Indeed, it has been shown that deuterated proteins, either expressed and analyzed in bacteria or delivered to human cells, can be detected by NMR and allow studying the interactions with the cellular environment (Majumder et al. 2015). In insect and human cells, direct expression of perdeuterated proteins is not possible due to the toxicity of 2H2O and the fact that all medium components need to be deuterated as well. However, because these cells are unable to synthesize most of the amino acids, amino acid type-selective labeling schemes are possible. In vitro, it has been shown that selective 1H,13C labeling of methyl groups of hydrophobic amino acids (Ala, Val, Leu, Ile, Met) with a deuterated side chain are extremely powerful probes when studying large molecules (Kerfah et al. 2015; Schütz and Sprangers 2020). Methyl-labeled amino acids can be supplemented in H2O-based media; therefore, we envision that similar isotope-labeling schemes will improve the detection of interacting proteins expressed in human cells (Fig. 3).
Existing and future labeling strategies applied to target-based drug screening in cells. For in vitro and in-cell NMR detection, proteins are typically labeled with 13C (light blue) and 15N (pink) isotopes (top left). However, large proteins or proteins interacting with large cellular components cannot be detected with such labeling schemes due to severe line broadening. Methyl-13C labeling (light blue) of specific amino acids coupled with side-chain deuteration (light orange, top right) can overcome the issue and allow detection of otherwise invisible proteins. The incorporation of amino acids containing a fluorine atom (lime green, bottom) also allows the detection of interacting proteins by 1D.19F NMR. An exemplary application of protein-detected in-cell NMR is in-cell drug screening, where compounds active in vitro (left) are screened to select the most active ones in a cellular setting for subsequent preclinical tests (right) (color figure online)
Recently, 19F NMR has emerged as an ideal probe for observing specific molecules in living cells. In terms of sensitivity, the 19F nucleus is second only to 1H, and importantly fluorine is not present in living systems, and therefore analysis of fluorinated molecules in cells is free from any cellular background interference. Such characteristics makes it a powerful alternative to 1H for investigating large complexes in living cells. Indeed, it has been shown that proteins expressed in bacteria with fluorinated amino acids are clearly detected, both directly in bacteria and upon delivery into human cells, also in the presence of interactions (Ye et al. 2013; Zhu et al. 2022). We have further extended the application of 19F NMR to proteins expressed in human cells: we showed that fluorinated aromatic amino acids (3-F-Tyr, 4-F-Phe, 5- and 6-F-Trp) are easily incorporated in the expressed proteins after switching the unlabeled expression medium with one where a selected amino acid is replaced with its fluorinated homolog (Fig. 3). Several proteins, including those that could not be detected in cells by 1H NMR due to extensive interactions, were successfully observed by recording simple 1D 19F NMR spectra, allowing the observation of intracellular protein–protein and protein–ligand interactions (Pham et al. 2023). Importantly, the above works may contribute to unlock the full potential of the 19F nucleus for detecting ‘invisible’ macromolecules in human cells: by combining 19F incorporation with 13C enrichment in specific aromatic side chain positions, 13C-19F TROSY effect could be exploited, which will further enhance the resolution and sensitivity of 13C-, 19F- or even aromatic 1H-detected NMR spectra (Boeszoermenyi et al. 2019), allowing detection of increasingly large intracellular complexes.
Concerning the application of 19F NMR to drug screening in cells, 19F-detection has been shown to be a promising alternative to 1H-detection to observe target proteins (see above) and nucleic acids (Krafčík et al. 2021). In vitro, however, 19F is more often employed in ligand-observed drug screening approaches: fluorine atoms are commonly found in libraries of chemical compounds or fragments, as well as in many approved drugs, and make possible to efficiently screen for target binding, as the shape and frequency of the 19F signals are highly sensitive to the interaction with the target (Buchholz and Pomerantz 2021; Dalvit and Vulpetti 2019). Therefore, we anticipate that 19F will enable ligand-detected in-cell NMR approaches, which will complement target-detected experiments in the study of ligand–target interactions, and will make possible to directly and selectively observe an uncharacterized compound, or a reference ligand, as it penetrates the cells and binds the intracellular target. Small molecules harboring fluorinated functional groups with favorable relaxation properties, such as –CF3, should allow detecting target binding with high sensitivity, even when the target itself is invisible. These approaches will be very useful for in-cell drug screening applications against NMR-invisible targets.
5 Long-term visions of (in-cell) NMR
The advancements described above will expand the capabilities of NMR to investigate macromolecules involved in increasingly large complexes within the cells. However, they do not overcome the main limitation of NMR spectroscopy: sensitivity. Over the years, the capabilities of high-field NMR instruments have improved greatly, both in terms of hardware (higher magnetic fields provide higher sensitivity) and NMR pulse sequences, making possible to record in minutes/hours data that previously required days/weeks. Despite this, at concentrations lower than few tens of µM, which is that of abundant cellular proteins, sensitivity remains a bottleneck. Even in the best-case scenario for transverse relaxation, which is that of (non-interacting) intrinsically disordered proteins, concentrations of the order of µM are necessary. This limits the application of NMR to proteins which are artificially overexpressed or delivered at higher levels than their natural abundance. However, overcoming this limitation is no easy task. Approaches relying on hyperpolarization of the nuclear spins might provide the required boost in sensitivity (Eills et al. 2023). Currently, nuclear hyperpolarization is best achieved in the solid phase. Enhancement of solid-state NMR signals by Dynamic Nuclear Polarization (DNP) is nowadays an established technique, and provides important atomic-resolution insights on the structure of proteins in cryopreserved cells or in native membranes (Kaplan et al. 2016; Narasimhan et al. 2019). In solution, a reagent hyperpolarized by DNP can be rapidly dissolved and injected in the sample where it undergoes chemical reactions, the products of which can be detected with high sensitivity. However, the short lifetime of the hyperpolarization limits dissolution-DNP applications to very fast real-time measurements of metabolic pathways (Jannin et al. 2019). Other approaches could reveal more suitable for applications to cells, such as photo-chemically induced DNP (photo-CIDNP), where the hyperpolarization is provided continuously by a photochemical reaction occurring within the sample. Photo-CIDNP allows NMR detection of molecules in solution at nanomolar concentrations (Yang et al. 2022), and in principle it could be applied to living cells, provided that suitable conditions are found to sustain the required photochemical reaction without compromising cell viability (Fig. 4). Overall, these hyperpolarization methods in solution are still in the initial stage of development, and their applicability to NMR of macromolecules in living cells is yet to be demonstrated.
The (in-cell) NMR spectroscopy of the future. Our vision of the future advancements of the in-cell NMR methodology. A Increased sensitivity will come from hyperpolarization approaches: photo-CIDNP relies on the optical excitation of a sensitizer molecule in the sample (such as fluorescein, left) to hyperpolarize the nuclei of selected moieties in biological samples (such as the indole ring of tryptophan, right). B Microcoils will allow a great reduction in required sample volume, allowing the analysis of smaller cell populations or even single cells. C Probes designed for simultaneous detection of multiple samples using parallel microcoils could be coupled with a flow apparatus for the injection and perfusion of multiple cell samples, while an optical fiber would allow light excitation of the samples. Physiologically and pharmacologically relevant samples that could be analyzed with such a device include organoids D and spheroids E
Finally, cellular applications of NMR spectroscopy could benefit from the development of new types of probes. In this respect, we envision that probe designs substantially different from the current state of the art will have to be developed, moving towards miniaturization/parallelization by exploiting different coil designs, such as microcoils (Bastawrous et al. 2022). Microcoils would allow the analysis of smaller cell populations, such as cells that cannot be easily grown in large numbers, or single spheroids/organoids, while parallelization could enable simultaneous measurement of multiple samples (Kupče et al. 2021). Furthermore, redesigned NMR bioreactors could exploit microfluidics to perfuse simultaneously multiple cell samples (Fig. 4). Clearly, such futuristic designs will only be of use if coupled with methods to boost the NMR signal, such as photo-CIDNP described above, to be able to detect signals from small sample volumes.
6 Conclusions
In this perspective, we started with an overview of what in-cell NMR can do today, in the context of the current (Cellular) Structural Biology field, we analyzed the technical and methodological challenges that will have to be overcome to allow broader application of in-cell NMR and, finally, we envisioned some of the possible future developments of the methodology. We intentionally did not focus on specific applications, as these have been extensively covered in other reviews. In general, however, we strongly believe that future cellular NMR applications will be instrumental to obtain biologically relevant insights on macromolecular dynamics and kinetics in native cellular compartments, cells and tissues, will greatly advance the understanding of molecular mechanisms involved in human diseases and in antimicrobial resistance, and will aid in the development of more efficient therapeutics.
References
Alshamleh I, Krause N, Richter C, Kurrle N, Serve H, Günther UL, Schwalbe H (2020) Real-time NMR spectroscopy for studying metabolism. Angew Chem (Int Ed) 59(6):2304–2308. https://doi.org/10.1002/anie.201912919
Banci L, Barbieri L, Bertini I, Luchinat E, Secci E, Zhao Y, Aricescu AR (2013) Atomic-resolution monitoring of protein maturation in live human cells by NMR. Nat Chem Biol 9(5):297–299. https://doi.org/10.1038/nchembio.1202
Barbieri L, Luchinat E (2021) Monitoring protein-ligand interactions in human cells by real-time quantitative in-cell NMR using a high cell density bioreactor. JoVE (J Vis Exp) 169:e62323. https://doi.org/10.3791/62323
Barbieri L, Luchinat E, Banci L (2015) Protein interaction patterns in different cellular environments are revealed by in-cell NMR. Sci Rep 5:14456. https://doi.org/10.1038/srep14456
Barnes CO, Pielak GJ (2011) In-cell protein NMR and protein leakage. Proteins 79(2):347–351. https://doi.org/10.1002/prot.22906
Bastawrous M, Gruschke O, Soong R, Jenne A, Gross D, Busse F, Nashman B, Lacerda A, Simpson AJ (2022) Comparing the potential of Helmholtz and planar NMR microcoils for analysis of intact biological samples. Anal Chem 94(23):8523–8532. https://doi.org/10.1021/acs.analchem.2c01560
Boeszoermenyi A, Chhabra S, Dubey A, Radeva DL, Burdzhiev NT, Chanev CD, Petrov OI, Gelev VM, Zhang M, Anklin C, Kovacs H, Wagner G, Kuprov I, Takeuchi K, Arthanari H (2019) Aromatic 19F–13C TROSY: a background-free approach to probe biomolecular structure, function, and dynamics. Nat Methods 16(4):333–340. https://doi.org/10.1038/s41592-019-0334-x
Breindel L, Burz DS, Shekhtman A (2020) Active metabolism unmasks functional protein–protein interactions in real time in-cell NMR. Commun Biol. https://doi.org/10.1038/s42003-020-0976-3
Buchholz CR, Pomerantz WCK (2021) 19F NMR viewed through two different lenses: ligand-observed and protein-observed 19F NMR applications for fragment-based drug discovery. RSC Chem Biol 2(5):1312–1330. https://doi.org/10.1039/D1CB00085C
Burz DS, Breindel L, Shekhtman A (2019) Improved sensitivity and resolution of in-cell NMR spectra. Methods Enzymol 621:305–328. https://doi.org/10.1016/bs.mie.2019.02.029
Carvalho J, Alves S, Castro MMCA, Geraldes CFGC, Queiroz JA, Fonseca CP, Cruz C (2019) Development of a bioreactor system for cytotoxic evaluation of pharmacological compounds in living cells using NMR spectroscopy. J Pharmacol Toxicol Methods 95:70–78. https://doi.org/10.1016/j.vascn.2018.11.004
Cerofolini L, Giuntini S, Barbieri L, Pennestri M, Codina A, Fragai M, Banci L, Luchinat E, Ravera E (2019) Real-time insights into biological events: in-cell processes and protein-ligand interactions. Biophys J 116(2):239–247. https://doi.org/10.1016/j.bpj.2018.11.3132
Chung JJ, Im H, Kim SH, Park JW, Jung Y (2020) Toward biomimetic scaffolds for tissue engineering: 3D printing techniques in regenerative medicine. Front Bioeng Biotechnol. https://doi.org/10.3389/fbioe.2020.586406
Dalvit C, Vulpetti A (2019) Ligand-based fluorine NMR screening: principles and applications in drug discovery projects. J Med Chem 62(5):2218–2244. https://doi.org/10.1021/acs.jmedchem.8b01210
Dzatko S, Krafcikova M, Hänsel-Hertsch R, Fessl T, Fiala R, Loja T, Krafcik D, Mergny J-L, Foldynova-Trantirkova S, Trantirek L (2018) Evaluation of the stability of DNA i-Motifs in the nuclei of living mammalian cells. Angew Chem (Int Ed) 57(8):2165–2169. https://doi.org/10.1002/anie.201712284
Eills J, Budker D, Cavagnero S, Chekmenev EY, Elliott SJ, Jannin S, Lesage A, Matysik J, Meersmann T, Prisner T, Reimer JA, Yang H, Koptyug IV (2023) Spin hyperpolarization in modern magnetic resonance. Chem Rev 123(4):1417–1551. https://doi.org/10.1021/acs.chemrev.2c00534
Gołowicz D, Kasprzak P, Orekhov V, Kazimierczuk K (2020) Fast time-resolved NMR with non-uniform sampling. Prog Nucl Magn Reson Spectrosc 116:40–55. https://doi.org/10.1016/j.pnmrs.2019.09.003
Hamatsu J, O’Donovan D, Tanaka T, Shirai T, Hourai Y, Mikawa T, Ikeya T, Mishima M, Boucher W, Smith BO, Laue ED, Shirakawa M, Ito Y (2013) High-resolution heteronuclear multidimensional NMR of proteins in living insect cells using a baculovirus protein expression system. J Am Chem Soc 135(5):1688–1691. https://doi.org/10.1021/ja310928u
Hertig D, Maddah S, Memedovski R, Kurth S, Moreno A, Pennestri M, Felser A, Nuoffer J-M, Vermathen P (2021) Live monitoring of cellular metabolism and mitochondrial respiration in 3D cell culture system using NMR spectroscopy. Analyst 146(13):4326–4339. https://doi.org/10.1039/D1AN00041A
Inomata K, Ohno A, Tochio H, Isogai S, Tenno T, Nakase I, Takeuchi T, Futaki S, Ito Y, Hiroaki H, Shirakawa M (2009) High-resolution multi-dimensional NMR spectroscopy of proteins in human cells. Nature 458(7234):106–109. https://doi.org/10.1038/nature07839
Jannin S, Dumez J-N, Giraudeau P, Kurzbach D (2019) Application and methodology of dissolution dynamic nuclear polarization in physical, chemical and biological contexts. J Magn Reson 305:41–50. https://doi.org/10.1016/j.jmr.2019.06.001
Kang C (2019) Applications of in-cell NMR in structural biology and drug discovery. Int J Mol Sci 20(1):139. https://doi.org/10.3390/ijms20010139
Kaplan M, Narasimhan S, de Heus C, Mance D, van Doorn S, Houben K, Popov-Čeleketić D, Damman R, Katrukha EA, Jain P, Geerts WJC, Heck AJR, Folkers GE, Kapitein LC, Lemeer S, van Bergen EnHenegouwen PMP, Baldus M (2016) EGFR dynamics change during activation in native membranes as revealed by NMR. Cell 167(5):1241–1251.e11. https://doi.org/10.1016/j.cell.2016.10.038
Kerfah R, Plevin MJ, Sounier R, Gans P, Boisbouvier J (2015) Methyl-specific isotopic labeling: a molecular tool box for solution NMR studies of large proteins. Curr Opin Struct Biol 32:113–122. https://doi.org/10.1016/j.sbi.2015.03.009
Kovermann M, Rogne P, Wolf-Watz M (2016) Protein dynamics and function from solution state NMR spectroscopy. Q Rev Biophys 49:e6. https://doi.org/10.1017/S0033583516000019
Krafčík D, Ištvánková E, Džatko Š, Víšková P, Foldynová-Trantírková S, Trantírek L (2021) Towards profiling of the G-quadruplex targeting drugs in the living human cells using NMR spectroscopy. Int J Mol Sci. https://doi.org/10.3390/ijms22116042
Kubo S, Nishida N, Udagawa Y, Takarada O, Ogino S, Shimada I (2013) A gel-encapsulated bioreactor system for NMR studies of protein-protein interactions in living mammalian cells. Angew Chem (Int Ed) 52(4):1208–1211. https://doi.org/10.1002/anie.201207243
Kühlbrandt W (2014) The resolution revolution. Science 343(6178):1443–1444. https://doi.org/10.1126/science.1251652
Kupče Ē, Frydman L, Webb AG, Yong JRJ, Claridge TDW (2021) Parallel nuclear magnetic resonance spectroscopy. Nat Rev Methods Prim. https://doi.org/10.1038/s43586-021-00024-3
Li C, Liu M (2013) Protein dynamics in living cells studied by in-cell NMR spectroscopy. FEBS Lett 587(8):1008–1011. https://doi.org/10.1016/j.febslet.2012.12.023
Li H, Yang Y, Hong W, Huang M, Wu M, Zhao X (2020) Applications of genome editing technology in the targeted therapy of human diseases: mechanisms, advances and prospects. Signal Transduct Target Ther. https://doi.org/10.1038/s41392-019-0089-y
Luchinat E, Banci L (2022) In-cell NMR: From target structure and dynamics to drug screening. Curr Opin Struct Biol 74:102374. https://doi.org/10.1016/j.sbi.2022.102374
Luchinat E, Barbieri L, Campbell TF, Banci L (2020) Real-time quantitative in-cell NMR: ligand binding and protein oxidation monitored in human cells using multivariate curve resolution. Anal Chem 92(14):9997–10006. https://doi.org/10.1021/acs.analchem.0c01677
Luchinat E, Barbieri L, Cremonini M, Banci L (2021a) Protein in-cell NMR spectroscopy at 1.2 GHz. J Biomol NMR 75(2):97–107. https://doi.org/10.1007/s10858-021-00358-w
Luchinat E, Barbieri L, Cremonini M, Pennestri M, Nocentini A, Supuran CT, Banci L (2021b) Determination of intracellular protein-ligand binding affinity by competition binding in-cell NMR. Acta Cryst 77(Pt 10):1270–1281. https://doi.org/10.1107/S2059798321009037
Luchinat E, Cremonini M, Banci L (2022) Radio signals from live cells: the coming of age of in-cell solution NMR. Chem Rev 122(10):9267–9306. https://doi.org/10.1021/acs.chemrev.1c00790
Majumder S, Xue J, DeMott CM, Reverdatto S, Burz DS, Shekhtman A (2015) Probing protein quinary interactions by in-cell nuclear magnetic resonance spectroscopy. Biochemistry 54(17):2727–2738. https://doi.org/10.1021/acs.biochem.5b00036
Mochizuki A, Saso A, Zhao Q, Kubo S, Nishida N, Shimada I (2018) Balanced regulation of redox status of intracellular thioredoxin revealed by in-cell NMR. J Am Chem Soc 140(10):3784–3790. https://doi.org/10.1021/jacs.8b00426
Monteith WB, Pielak GJ (2014) Residue level quantification of protein stability in living cells. Proc Natl Acad Sci USA 111(31):11335–11340. https://doi.org/10.1073/pnas.1406845111
Monteith WB, Cohen RD, Smith AE, Guzman-Cisneros E, Pielak GJ (2015) Quinary structure modulates protein stability in cells. Proc Natl Acad Sci USA 112(6):1739–1742. https://doi.org/10.1073/pnas.1417415112
Mu X, Choi S, Lang L, Mowray D, Dokholyan NV, Danielsson J, Oliveberg M (2017) Physicochemical code for quinary protein interactions in Escherichia coli. Proc Natl Acad Sci 114(23):E4556–E4563. https://doi.org/10.1073/pnas.1621227114
Narasimhan S, Scherpe S, LuciniPaioni A, van der Zwan J, Folkers GE, Ovaa H, Baldus M (2019) DNP-supported solid-state NMR spectroscopy of proteins inside mammalian cells. Angew Chem Int Ed 58(37):12969–12973. https://doi.org/10.1002/anie.201903246
Ogino S, Kubo S, Umemoto R, Huang S, Nishida N, Shimada I (2009) Observation of NMR signals from proteins introduced into living mammalian cells by reversible membrane permeabilization using a pore-forming toxin, streptolysin O. J Am Chem Soc 131(31):10834–10835. https://doi.org/10.1021/ja904407w
Pham LBT, Costantino A, Barbieri L, Calderone V, Luchinat E, Banci L (2023) Direct expression of fluorinated proteins in human cells for 19F in-cell NMR spectroscopy. J Am Chem Soc 145(2):1389–1399. https://doi.org/10.1021/jacs.2c12086
Prince E, Kumacheva E (2019) Design and applications of man-made biomimetic fibrillar hydrogels. Nat Rev Mater. https://doi.org/10.1038/s41578-018-0077-9
Riek R, Pervushin K, Wüthrich K (2000) TROSY and CRINEPT: NMR with large molecular and supramolecular structures in solution. Trends Biochem Sci 25(10):462–468. https://doi.org/10.1016/S0968-0004(00)01665-0
Rossi G, Manfrin A, Lutolf MP (2018) Progress and potential in organoid research. Nat Rev Genet. https://doi.org/10.1038/s41576-018-0051-9
Sakai T, Tochio H, Tenno T, Ito Y, Kokubo T, Hiroaki H, Shirakawa M (2006) In-cell NMR spectroscopy of proteins inside Xenopus laevis oocytes. J Biomol NMR 36(3):179–188. https://doi.org/10.1007/s10858-006-9079-9
Schütz S, Sprangers R (2020) Methyl TROSY spectroscopy: a versatile NMR approach to study challenging biological systems. Prog Nucl Magn Reson Spectrosc 116:56–84. https://doi.org/10.1016/j.pnmrs.2019.09.004
Selenko P, Serber Z, Gadea B, Ruderman J, Wagner G (2006) Quantitative NMR analysis of the protein G B1 domain in Xenopus laevis egg extracts and intact oocytes. Proc Natl Acad Sci USA 103(32):11904–11909. https://doi.org/10.1073/pnas.0604667103
Serber Z, Keatinge-Clay AT, Ledwidge R, Kelly AE, Miller SM, Dötsch V (2001) High-resolution macromolecular NMR spectroscopy inside living cells. J Am Chem Soc 123(10):2446–2447. https://doi.org/10.1021/ja0057528
Sharaf NG, Barnes CO, Charlton LM, Young GB, Pielak GJ (2010) A bioreactor for in-cell protein NMR. J Magn Reson 202(2):140–146. https://doi.org/10.1016/j.jmr.2009.10.008
Siegal G, Selenko P (2019) Cells, drugs and NMR. J Magn Reson 306:202–212. https://doi.org/10.1016/j.jmr.2019.07.018
Theillet F-X (2022) In-cell structural biology by NMR: the benefits of the atomic scale. Chem Rev 122(10):9497–9570. https://doi.org/10.1021/acs.chemrev.1c00937
Theillet F-X, Luchinat E (2022) In-cell NMR: why and how? Prog Nucl Magn Reson Spectrosc 132–133:1–112. https://doi.org/10.1016/j.pnmrs.2022.04.002
Theillet F-X, Binolfi A, Bekei B, Martorana A, Rose HM, Stuiver M, Verzini S, Lorenz D, van Rossum M, Goldfarb D, Selenko P (2016) Structural disorder of monomeric α-synuclein persists in mammalian cells. Nature 530(7588):45–50. https://doi.org/10.1038/nature16531
Yang H, Li S, Mickles CA, Guzman-Luna V, Sugisaki K, Thompson CM, Dang HH, Cavagnero S (2022) Selective isotope labeling and LC-photo-CIDNP enable NMR spectroscopy at low-nanomolar concentration. J Am Chem Soc 144(26):11608–11619. https://doi.org/10.1021/jacs.2c01809
Ye Y, Liu X, Zhang Z, Wu Q, Jiang B, Jiang L, Zhang X, Liu M, Pielak GJ, Li C (2013) 19F NMR spectroscopy as a probe of cytoplasmic viscosity and weak protein interactions in living cells. Chem Eur J 19(38):12705–12710. https://doi.org/10.1002/chem.201301657
Zanoni M, Cortesi M, Zamagni A, Arienti C, Pignatta S, Tesei A (2020) Modeling neoplastic disease with spheroids and organoids. J Hematol Oncol 13(1):97. https://doi.org/10.1186/s13045-020-00931-0
Zhu W, Guseman AJ, Bhinderwala F, Lu M, Su X-C, Gronenborn AM (2022) Visualizing proteins in mammalian cells by 19F NMR spectroscopy. Angew Chem Int Ed 61(23):e202201097. https://doi.org/10.1002/anie.202201097
Acknowledgements
The authors acknowledge the support by the Italian Ministry for University and Research (FOE funding) to the CERM/CIRMMP Italian Centre of Instruct-ERIC, a landmark ESFRI project.
Funding
Open access funding provided by Università degli Studi di Firenze within the CRUI-CARE Agreement. E.L. contract is supported by Horizon 2020 funds (grant nos. 862658, 817737, 862480) and by the MUR BIOZOOSTAIN project of the University of Bologna.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The manuscript complies to the Ethical Rules applicable for this journal.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This contribution belongs to the topical collection Lincei Prizewinners Author Lucia Banci was the winner of the 2022 International Prize Professor Luigi Tartufari for Physics/Chemistry attributed by the Accademia Nazionale dei Lincei in Rome.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Luchinat, E., Banci, L. In-cell NMR: recent progresses and future challenges. Rend. Fis. Acc. Lincei 34, 653–661 (2023). https://doi.org/10.1007/s12210-023-01168-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12210-023-01168-y