Abstract
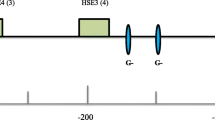
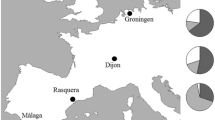
The small heat shock gene (shsp) cluster of Drosophila buzzatii was sequenced and the gene order and DNA sequence were compared with those of the shsps in Drosophila melanogaster. The D. buzzatii shsp cluster contains an inversion and a duplication of hsp26. A phylogenetic tree was constructed based on hsp26 genes from several Drosophila species of the Sophophora and Drosophila subgenera. The tree shows first a separation of the Sophophora and the Drosophila subgenera and then the Drosophila subgenus is divided into the Hawaiian Drosophila and the repleta/virilis groups. Only the latter contain a duplicated hsp26. Comparing the gene organisation of the shsp cluster shows that all the Drosophila subgenus species contain the inversion. Putative heat shock elements (HSE) were found in the promoters of all the shsp and putative regulator elements for tissue specific expression were found in the promoter of hsp23, hsp27 and one of the hsp26 genes. hsp23 was found to be polymorphic for four non-synonymous changes that all lead to exchange of a Valine. The duplicated hsp26 gene in D. buzzatii (phsp26) was polymorphic for two non-synonymous changes. The allele frequencies of these variants were determined in nine D. buzzatii populations covering most of its distribution in Australia using high-resolution melting curves. The allele frequencies of one of the hsp23 variants showed a significant linear regression with longitude and the pooled frequency of the four Valine changes of hsp23 in the nine populations showed a significant linear regression with longitude and with a composite measure of climatic variables.




Similar content being viewed by others
Abbreviations
- shsp :
-
small heat shock protein gene
- HSE:
-
heat shock element
- EcRE:
-
ecdysone response element
- CTAB:
-
hexadecyltrimethylammonium bromide
- CNS:
-
conserved non-coding sequence
References
Adams MD, Celniker SE, Holt RA, Evans CA, Gocayne JD, Amanatides PG et al (2000) The genome sequence of Drosophila melanogaster. Science 287:2185–2195
Alonso J, Rodriguez JM, Baena-López LA, Alonso MT, Santarén JF (2005) Constitutive expression of heat shock protein p23 correlates with proneural territories in imaginal discs of Drosophila melanogaster. Proteomics 5:1–10
Andolfatto P (2001) Contrasting pattern of X-linked and autosomal nucleotide variation in Drosophila melanogaster and Drosophila simulans. Mol Biol Evol 18:279–290
Ayme A, Tissières A (1985) Locus 67B of Drosophila melanogaster contains seven, not four, closely related heat shock genes. EMBO J 4:2949–2954
Barker JSF, Sene FD, East PD, Pereira MAQR (1985) Allozyme and chromosomal polymorphism of Drosophila buzzatii in Brazil and Argentina. Genetica 67:161–170
Barker JSF, Krebs RA, Davies HI (2005) Geographical distributions, relative abundance and coexistence of Drosophila aldrichi and Drosophila buzzatii in Australia. Austral Ecol 30:546–557
Barker JSF, Frydenberg J, González J, Davies HI, Ruiz A, Sørensen JG, Loeschcke V (2009) Bottlenecks, population differentiation and apparent selection at microsatellite loci in Australian Drosophila buzzatii. Heredity 102:389–401
Birney E, Durbin R (1997) Dynamite: a flexible code generating language for dynamic programming methods used in sequence comparison. Proc Int Conf Intell Syst Mol Biol 5:56–64
Brudno M, Steinkamp R, Morgenstern B (2004) The CHAOS/DIALIGN WWW server for multiple alignment of genomic sequences. Nucleic Acids Res 32:W41–W44
Cohen RS, Meselson M (1985) Separate regulatory elements for the heat-inducible and ovarian expression of the Drosophila hsp26 gene. Cell 43:737–746
Collinge JE, Anderson AR, Weeks AR, Johnson TK, McKechnies (2008) Latitudinal and cold-tolerance variation associate with DNA repeat-number variation in the hsr-omega RNA gene of Drosophila melanogaster. Heredity 101:260–270
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Drosophila 12 Genomes Consortium (2007) Evolution of genes and genomes on the Drosophila phylogeny. Nature 450:203–218
Dubrovsky EB, Dubrovskaya VA, Berger EM (2001) Selective binding of Drosophila BR-C isoforms to a distal regulatory element in the hsp23 promoter. Insect Biochem Mol Biol 31:1231–1239
Flybase Consortium (2002) The flybase database of the Drosophila genome projects and community literature. Nucleic Acids Res. 30:106–108
Force A, Cresko WA, Pickett FB, Proulx SR, Amemiya C, Lynch M (2005) The origin of subfunctions and modular gene regulation. Genetics 170:433–446
Frank LH, Cheung HK, Cohen RS (1992) Identification and characterization of Drosophila female germ line transcriptional control elements. Development 114:481–491
Fry JD, Donlon K, Saweikis M (2008) A worldwide polymorphism in aldehyde dehydrogenase in Drosophila melanogaster: evidence for selection mediated by dietary ethanol. Evolution 62:66–75
Frydenberg J, Pertoldi C, Dahlgaard J, Loeschcke V (2002) Genetic variation in original and colonizing Drosophila buzzatii populations analysed by microsatellite loci isolated with a new PCR screening method. Mol Ecol 11:181–190
Frydenberg J, Hoffmann AA, Loeschcke V (2003) DNA sequence variation and latitude associations in hsp23, hsp26 and hsp27 from populations of Drosophila melanogaster. Mol Ecol 12:2025–2032
Gómez GA, Hasson E (2003) Transpecific polymorphisms in an inversion linked esterase locus in Drosophila buzzatii. Mol Biol Evol 20:410–423
González J, Casals F, Ruiz A (2007) Testing chromosomal phylogenies and inversion breakpoint reuse in Drosophila. Genetics 175:167–177
Gottler LM, Salud Bea R, Shelburne CE, Ramamoorthy A, Marsh ENG (2008) Using fluorous amino acids to probe the effects of changing hydrophobicity on the physical and biological properties of β-Hairpin antimicrobial peptide protegrin-1. Biochem 47:9243–9250
Hasson E, Rodriguez C, Fanara JJ, Naveira H, Reig OA, Fontdevila A (1985) The evolutionary history of Drosophila buzzatii XXVI. Macrogeographic patterns in the inversion polymorphism in New World populations. J Evol Biol 8:369–384
Houlder DJ, Hutchinson MF, Nix HA, McMahon JP (2000) Anuclim User Guide, Version 5.1. Centre for Ressource and Environmental Studies, Australian National University, Canberra
JMP (2007) SAS Institut. Inc. Cary, NC. www.JMP.com
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Laayouni H, Santos M, Fontdevila A (2000) Toward a physical map of Drosophila buzzatii: use of randomly amplified polymorphic DNA polymorphisms and sequence-tagged site landmarks. Genetics 156:1797–1816
Matzkin LM, Merritt TJS, Zhu C-T, Eanes WF (2005) The structure and population genetics of the breakpoints associated with the cosmopolitan chromosomal inversion In(3R)Payne in Drosophila melanogaster. Genetics 170:1143–1152
Mayor C, Brudno M, Schwartz JR, Poliakov A, Rubin EM, Frazer KA, Pachter LS, Dubchak I (2000) VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 16:1046–1047
Mestril R, Schiller P, Amin J, Klapper H, Ananthan J, Voellmy R (1986) Heat shock and ecdysterone activation of Drosophila melanogaster hsp23 gene; a sequence element implied in developmental regulation. EMBO J 5:1667–1673
Michaud S, Tanguay RM (2003) Expression of the Hsp23 chaperone during Drosophila embryogenesis: association to distinct neural and glial lineages. BMC Dev Biol 3:9–20
Michaud S, Marin R, Tanguay RM (1997) Regulation of heat shock gene induction and expression during Drosophila development. Cell Mol Life Sci 53:104–113
Moriyama EN, Powell JR (1996) Intraspecific nuclear DNA variation in Drosophila. Mol Biol Evol 13:261–277
Morrow G, Heikkile JJ, Tanguay RM (2006) Differences in the chaperone-like activities of the four main small heat shock proteins of Drosophila melanogaster. Cell Stress Chaperones 11:51–60
Norry FM, Sambucetti P, Scannapieco AC, Loeschcke V (2006) Altitudinal patterns for longevity, fecundity and senescence in Drosophila buzzatii. Genetica 128:81–93
Piccinali R, Aguadé M, Hasson E (2004) Comparative molecular population genetics of the Xdh locus in the cactophilic sibling species Drosophila buzzatii and D. koepfeae. Mol Biol Evol 21:141–152
Piccinali RV, Mascord LJ, Barker JSF, Oakeshott JG, Hasson E (2007) Molecular population genetics of the α-Esterase5 gene locus in original and colonized populations of Drosophila buzzatii and its sibling Drosophila koepferae. J Mol Evol 64:158–170
Powell LM, zur Lage PI, Prentice DRA, Senthinathan B, Jarman AP (2004) The proneutral proteins Atonal and Scute regulate neural target genes through different E-box binding sites. Mol Cell Biol 24:9517–9526
Richards S, Liu Y, Bettencourt BR, Hradecky P, Letovsky S, Nielsen R et al (2005) Comparative genome sequencing of Drosophila pseudoobscura: chromosomal, gene, and cis-element evolution. Genome Res 15:1–18
Riddihough G, Pelham HRB (1986) Activation of the Drosophila hsp27 promoter by heat shock and by ecdysone involves independent and remote regulatory sequences. EMBO J 5:1653–1658
Rogulski KR, Cartwright IL (1995) Multiple interacting elements delineate an Ecdysone-dependent regulatory region with secondary responsive character. J Mol Biol 249:298–318
Rozas J, Sánchez-DelBarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Sandaltzopoulos R, Mitchelmore C, Bonte E, Wall G, Becker PB (1995) Dual regulation of the Drosophila hsp26 promoter in vitro. Nucleic Acids Res 23:2479–2487
Sharakhov IV, White BJ, Sharakhova MV, Kayondo J, Lobo NF, Santolamazza F, Torre AD, Simard F, Collins FH, Besanaky NJ (2006) Breakpoint structure reveals the unique origin of an interspecific chromosomal inversion (2La) in the Anopheles gambiae complex. Proc Natl Acad Sci USA 103:6258–6262
Sorensen JG, Norry FM, Scannapieco AC, Loeschcke V (2005) Altitude variation for stress resistance traits and thermal adaptation in adult Drosophila buzzatii from the New World. J Evol Biol 18:829–837
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Acknowledgments
The authors thank Jesper G. Sørensen for help catching some of the Australian fly samples and Fabian Norry for some of the Argentinean fly samples, Jürgen Tomiuk and Cino Pertoldi for critical reading of the manuscript and the Novo Nordisk Foundation for financial support.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Homologous sequences in D. buzzatii and D. melanogaster hsp23 promoters (DOC 33 kb)
Table S2
Homologous sequences in D. buzzatii phsp26 and D. melanogaster hsp26 promoters (DOC 31 kb)
Table S3
Homologous sequences in D. buzzatii hsp26 and D. melanogaster hsp26 promoters (DOC 29 kb)
Table S4
Homologous sequences in D. buzzatii and D. melanogaster hsp27 promoters (DOC 31 kb)
Rights and permissions
About this article
Cite this article
Frydenberg, J., Barker, J.S.F. & Loeschcke, V. Characterization of the shsp genes in Drosophila buzzatii and association between the frequency of Valine mutations in hsp23 and climatic variables along a longitudinal gradient in Australia. Cell Stress and Chaperones 15, 271–280 (2010). https://doi.org/10.1007/s12192-009-0140-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12192-009-0140-y