Abstract
Multiple myeloma is divided into two distinct genetic subtypes based on chromosome content. Hyperdiploid myeloma is characterized by multiple trisomies of chromosomes 3, 5, 7, 9 11, 15, 19 and 21, and lacks recurrent immunoglobulin gene translocations. Non-hyperdiploid myeloma in contrast is characterized by chromosome translocations t(4;14), t(14;16), t(14;20), t(6;14) and t(11;14). A unifying event in the pathogenesis of multiple myeloma is the dysregulated expression of a cyclin D gene, either directly by juxtaposition to an immunoglobulin enhancer, as a result of ectopic expression of a MAF family transcription factor, or indirectly by as yet unidentified mechanisms. Secondary genetic events include rearrangements of MYC, activating mutations of NRAS, KRAS or BRAF, a promiscuous array of mutations that activate NFkB and deletions of 17p. Among the poor-risk genetic features are t(4;14), t(14;16), t(14;20), del 17p and gains of 1q. Available evidence supports the use of a risk-stratified approach to the treatment of patients with multiple myeloma, with the early and prolonged use of bortezomib particularly in patients with t(4;14) and del 17p.
Similar content being viewed by others
Introduction
Multiple myeloma (MM) is a monoclonal tumor of antibody-secreting plasma cells (PC) in the bone marrow (BM) that is often diagnosed by the presence of a typical M-spike by serum protein electrophoresis (SPEP) or by free light chains in the urine. In its symptomatic phase is associated with significant end organ damage including lytic bone lesions, anemia, loss of kidney function, immunodeficiency, and amyloid deposits in various tissues [1]. MM incidence is higher in blacks than whites, and in men, than women [2], for a total estimate of 22,350 new cases and 10,710 deaths in the United States in 2013 [3]. Although MM continues to be considered an incurable disease, thanks to the recent therapeutic advances, the 5-year survival rate reported in the SEER database has increased from 28 % (1987–1989) to 43 % (2002–2008) [2]. Notably, a subset of patients with cytogenetically defined low-risk MM, initially treated in 1999 were reported having a 10-year survival rate of 75 % [4], with presumably even better results possible for patients starting treatment today. MM cells are the malignant counterparts of post-germinal center (GC) long-lived PCs, characterized by strong BM dependence, somatic hypermutation (SHM) of immunoglobulin (Ig) genes, and isotype class switch resulting in the absence of IgM expression in all but 1 % of tumors [5]. However, MM cells differ from healthy PCs because they retain the potential for a low rate of proliferation (1–3 % of cycling cells).
Multi-step clinical course of multiple myeloma
Virtually every case of MM is preceded by a pre-malignant PC tumor called monoclonal gammopathy of undetermined significance (MGUS) [6, 7] that, like MM, produces a typical M-spike (almost always non-IgM) by SPEP or free light chain in the urine. It has to be distinguished from a IgM secreting lymphoid MGUS, a precursor phase of chronic lymphocytic leukemia, lymphoplasmacytoma and Waldenstrom’s macroglobulinemia. PC MGUS is age dependent, is present in about 4 % of individuals over the age of 50 [8, 9], and can progress to MM at average rates of 1 % per year. MGUS is distinguished from MM by having a M-spike of <30 g/L, with no more than 10 % of BM mononuclear cells being tumor cells, and no end organ damage or other symptoms. Progression of MGUS to smoldering MM and symptomatic MM is associated with an expanding BM tumor mass and increasingly severe organ impairment or symptoms [1]. Despite the recent advances in the understanding of the MM pathogenesis, it is still largely impossible to predict which MGUS patient will and which one will never progress to MM. Although MM cells are characterized by a strong dependence on the BM tumor microenvironment, at late stages of the disease the more aggressive tumor may sometimes extend to extramedullary locations, such as spleen, liver, and extracellular spaces. Extramedullary MM (EMM) can also present with a leukemic phase, that is classified as secondary or primary plasma cell leukemia (PCL), depending on whether or not a preceding intramedullary MM was recognized. Most of the available human MM cell lines (HMCLs) have been generated from EMM or PCL tumors [10, 11] and represent a renewable repository of the oncogenic events involved in initiation and progression of the most aggressive end-stage MM tumors.
Post-germinal center long-lived plasma cells are the normal counterpart of the malignant cell in multiple myeloma
During a secondary immune response, activated lymphocytes migrate into GCs were they undergo antigen selection by multiple rounds of somatic hypermutation (SHM) and IgH class switch recombination (CSR). Cells whose B cell receptor loses affinity for the antigen are counter-selected and undergo apoptosis, while positively selected cells are rescued from apoptosis by expression of BCL2 and differentiate into either memory cells or plasma blasts (PB) before homing to the BM as long-lived PC. Although pre-GC short-lived PCs can also be generated during primary immune response, the presence of somatic mutations in the immunoglobulin genes without further remodeling clearly indicates a post-GC origin for MM.
Primary IgH translocations are an early oncogenic event shared by MGUS and MM
Translocations involving the IgH locus (14q32) or one of the IgL loci (κ, 2p12 or λ, 22q11) are present in at least half of MM cases and are thought to result from errors during the physiological process of CSR or SHM since the breakpoints are usually located near or within IgH switch regions, but sometimes near VDJ sequences [12]. It is presumed that these translocations represent primary—perhaps initiating—oncogenic events as normal B cells pass through GCs. In fact, although clonal heterogeneity has been identified in MM as in many other cancers, the primary chromosome translocations continue to mark the tumor clone throughout disease progression. As in other B cell tumors, these translocations result in dysregulated expression of an oncogene that is juxtaposed to the strong Ig enhancers. However, translocations involving an IgH switch region uniquely dissociate the intronic (Emu) from one or both 3′ IgH enhancers (3′E), so that two putative oncogenes can become dysregulated on the two derivative chromosomes. This is exemplified by the t(4;14) translocation that simultaneously dysregulate FGFR3 on der(14) and MMSET on der(4) in MM.
These IgH translocations are efficiently detected by fluorescent in situ hybridization (FISH) analyses. Large studies from several groups show that the prevalence of IgH translocations increases with disease stage: about 50 % in MGUS or SMM, 55–70 % for intramedullary MM, 85 % in PCL, and >90 % in HMCL [13, 14]. Limited studies indicate that IgL translocations are present in about 10 % of MGUS/SMM tumors, and about 15–20 % of intramedullary MM tumors and HMCL [11]. Translocations involving an IgK locus are rare, occurring in only 1–2 % of MM tumors and HMCL [11].
Primary IgH translocations dysregulate three gene groups: CCND, MAF and FGFR3/MMSET
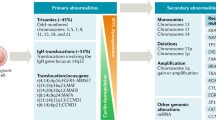
There are three recurrent primary IgH translocation groups, with the chromosomal sites, target oncogenes, and approximate prevalence in MM (~40 % prevalence for all three groups) as follows: CYCLIN D (11q13-CYCLIN D1-15 %; 12p13-CYCLIN D2-<1 %; 6p25-CYCLIN D3-2 %) MAF (16q23-MAF-5 %; 20q12-MAFB-2 %; 8q24.3-MAFA-<1 %; MMSET/(FGFR3)-4p16-(MMSET in all but also FGFR3 in 80 % of these tumors)-15 % (Fig. 1). With the exception perhaps of FGFR3, it is interesting to note that none of the primary translocations causes dysregulation of strong oncogenes, suggesting that perhaps this would be incompatible with terminal differentiation of PCs and their homing to the BM. Also IgH translocation groups are mutually exclusive, although double translocations have been reported in HMCLs (e.g. KMS11 carries both a MAF and FGFR3 translocation on the two IgH alleles).
It is thought that CYCLIN D translocations only dysregulate expression of a CYCLIN D gene. By contrast MAF translocations dysregulate expression of a MAF transcription factor that causes increased expression of many genes, including CYCLIN D2 and adhesion molecules that are thought to enhance the ability of the tumor cell to interact with the BM microenvironment [15, 16]. The contributions of the two genes dysregulated by t(4;14) remain controversial. MMSET is a chromatin-remodeling factor that is over-expressed in all tumors with a t(4;14), whereas about 20 % of tumors lack der(14) and FGFR3 expression. The rare acquisition of FGFR3 activating mutations during progression confirms a role for FGFR3 in MM pathogenesis. Although an activated mutant FGFR3 can be oncogenic, it recently was shown that wild-type FGFR3 (as is found in most t[4; 14]) can contribute to B cell oncogenesis [17]. It remains to be determined if FGFR3 is critical early in pathogenesis but becomes dispensable during progression of t(4;14) MM, especially in the presence of RAS-BRAF activating mutations that, like mutated FGFR3, also lead to constitutive phosphorylation of ERK1-2. Preclinical studies suggest that tyrosine kinase inhibitors are active only against t(4;14) HMCL with activating mutations of FGFR3, whereas anti-FGFR3 monoclonal antibodies that inhibit FGFR3 signaling but also elicit antibody-dependent cell-mediated cytotoxicity are active against HMCLs expressing wild-type FGFR3 [18, 19]. Definitive results about the clinical activity of FGFR3 targeted therapy have not been reported yet. Despite an apparently indispensable role in t(4;14) MM, it remains to be determined how MMSET contributes to MM pathogenesis. There are some clues. It is a histone methyltransferase for H3K36me2, and when over-expressed results in a global increase in H3K36me2 methylation, and a decrease in H3K27me3 methylation, which most likely is the cause of the many changes in gene expression observed in t(4;14) tumors [15, 20–22]. In addition, it recently has been determined that MMSET has a role in DNA repair. Following DNA damage MMSET is phosphorylated on Ser102 by ATM and is recruited to sites of double strand breaks (DSB) where it results in methylation of H4K20 that is required for recruitment of p53-binding protein (53BP1). 53BP1 is required for p53 accumulation, G2/M checkpoint arrest, and the intra-S-phase checkpoint in response to ionizing radiation. Approximately half of the translocation breakpoints in t(4;14) MM result in a truncated MMSET that lacks Ser102 and cannot be recruited to DSBs, resulting in a failure to recruit 53BP1 and a loss of the normal DNA damage response pathway. It is not known whether this biologic difference results in a different clinical outcome for t(4;14) MM patients with a truncated versus full-length MMSET [23]. Importantly, loss of MMSET expression alters adhesion, suppresses growth, and results in apoptosis of HMCLs, suggesting that it is an attractive therapeutic target [21]. MMSET has been shown to post-transcriptionally enhance the expression of MYC by repressing miR-126, which targets the 3′ untranslated region of MYC, inhibiting translation [24]. MMSET has been found over-expressed in a subset of many different cancers, where its over-expression correlates with tumor aggressiveness and prognosis [25, 26]. It has been shown to be a required effector of EZH2, and in prostate cancer to mediate constitutive NF-kB activation, and, by activation of TWIST, epithelial to mesenchymal transition [25, 27, 28].
Multiple trisomies is an alternative pathogenetic pathway
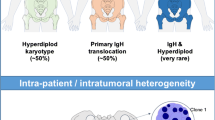
There is a consensus that chromosome content reflects at least two pathways of pathogenesis. Nearly half of MGUS and MM tumors are hyperdiploid (HRD), with 48–75 (mostly 49–56) chromosomes, usually with extra copies of three or more specific chromosomes (3, 5, 7, 9, 11, 15, 19, 21). Non-hyperdiploid (NHRD) tumors have <48 and/or >75 chromosomes. Strikingly, HRD tumors rarely (~10 %) have a primary IgH translocation, whereas NHRD tumors usually (~70 %) have an IgH translocation [29] (Fig. 2). Although it has been proposed that NHRD and HRD tumors represent different pathways of pathogenesis, the timing, mechanism, and molecular consequences of hyperdiploidy are unknown. In any case, HRD patients seem to have a better prognosis than NHRD patients. Curiously, EMM tumors and HMCLs nearly always have a NHRD genotype, suggesting that HRD tumors are more stromal cell dependent than NHRD tumors. Alternatively it is possible that HRD is selected in proliferating cells. In fact, a few cell lines derived from HRD patients have lost the extra chromosomes (unpublished observation). Interestingly, in patients with t(4;14) or t(14;16) or t(14;20) or del17p the presence of one or more trisomies is associated with a substantially better prognosis than absence of trisomies. This suggests that the phenotype associated with trisomies may be dominant [30].
Cyclin Ds dysregulation in MM. MGUS and MM karyotypes can be divided into hyperdiploid and non-hyperdiploid based on chromosomal content. Almost all hyperdiploid tumors have biallelic cyclin D1 trans-dysregulation. Non-hyperdiploid tumors often have t(14q32) translocations affecting the indicated loci (frequency is shown). In about 25 % of them, one of the D type cyclin is cis-dysregulated by a 14q32 translocation, in the other non-hyperdiploid tumors cyclin D2 expression is trans-dysregulated
Cyclin Ds are ectopically expressed throughout MM progression
Almost all cases of plasma cell neoplasm starting from the MGUS stage and independently on the chromosome content aberrantly express one or more of the CYCLIN D genes and it has been proposed that dysregulation of a CYCLIN D gene provides a unifying, early oncogenic event in MGUS and MM (Fig. 2). Remarkably though this is not associated with increased proliferation, as the PC labeling index in MGUS, like in normal PCs, remain virtually = 0. Yet the expression level of cyclin D1, cyclin D2 or cyclin D3 mRNA in MM and MGUS is distinctly higher than in normal PCs. This results from several mechanisms including a direct cis-dysregulation in MM tumors with a CYCLIN D gene translocation [i.e. t(11;14) t(6;14) or t(12;14)] or a trans-dysregulation in tumors with a translocation of MAF [t(14;16)], encoding a transcription factor that directly binds to the CYCLIN D2 promoter. Although MMSET/FGFR3 tumors express moderately high levels of CYCLIN D2, the cause of increased CYCLIN D2 expression remains unknown. The majority of HRD tumors express CYCLIN D1 bi-allelically, perhaps because they contain a trisomic chromosome 11, whereas most other tumors express increased levels of CYCLIN D2 by unknown mechanism. Only a few percent of MM tumors do not express any CYCLIN D gene, but have been shown to contain a high level of contamination with normal cells. Another fraction of cyclin D negative samples shows bi-allelic deletion of RB1, the cell cycle inhibitor directly targeted by CYCLIN D, therefore bypassing the need for CYCLIN D gene.
Molecular classification of MM
The patterns of spiked expression of genes deregulated by primary IgH translocations and the universal over-expression of CCNDs genes led to the translocations and cyclin D (TC) classification that includes eight groups: those with primary translocations (designated 4p16, 11q13, 6p21, MAF), those that over-expressed CCND1 and CCND2 either alone or in combination (D1, D1&D2, D2), and the rare cases that do not over-express any CCND genes (‘none’) [15]. Greater than 95 % of tumors in the D1 group are HRD. In addition, most of the patients with HRD MM and trisomy 11 fall within the D1 and D1&D2 groups, while those without trisomy 11 fall within the D2 group, although a majority of the D2 group are NHRD. This classification system is derived from a supervised analysis of gene expression data based on the different mechanisms that dysregulate a CCND gene as an early and unifying event in pathogenesis.
An MM classification based on an unsupervised analysis of microarray gene expression profiling from the UAMS identified 7 tumor groups characterized by the co-expression of unique gene clusters [31]. This classification was partially replicated in an independent unsupervised analysis of a combined HOVON-GMMG dataset that identified 10 tumor groups with considerable overlap with the UAMS groups [32]. Interestingly, these clusters partially overlap with the subgroups of the TC classification corresponding to the different primary translocations and HRD. Importantly, however, they also highlight other secondary events that become dominant during MM progression that can occur independently in each subtype of MM: proliferation (PR), expression of NFkB target genes (NFkB), cancer-testis antigens (CTA), and the phosphatase PTP4A3/PRL3 (PRL3). In addition to insights into the molecular biology of the disease, these classifications are prognostically relevant because, together with other cytogenetic markers (i.e. 17p deletion) they help stratifying patients into high and low risk. The CD-1 and CD-2 groups represent subgroups of patients with t(11;14) and t(6;14), with the former characterized by argininosuccinate synthetase 1 expression, and the later by expression of B cell antigens (CD20, VPREB, CD79A). Interestingly they identify patients with markedly different clinical outcomes. Of all the molecular subgroups, CD-1 has the quickest onset and highest frequency of CR (90 %), whereas CD-2 has the slowest onset, and lowest frequency of CR (45 %), when treated with Total Therapy 3. However, after the MF, the CD-1 has the shortest CR duration (77 % at 2 years), whereas the CD-2 has the longest (100 % at 2 years) [33].
Secondary oncogenic events drive MGUS and MM progression
A plethora of mutations have been identified in MM patients, which can occur at different frequencies independently in the different disease groups and are thought to promote disease progression.
MYC dysregulation
There is increased expression of c-MYC in most newly diagnosed MM tumors compared to MGUS tumors [34]. Recently, it was shown that sporadic activation of a MYC transgene in GC B cells in an MGUS prone mouse strain led to the universal development of MM tumors [35, 36]. Hence, increased MYC expression seems to be responsible for progression from MGUS to MM. Complex translocations involving MYC (c-MYC ≫ N-MYC > L-MYC) appear to be secondary progression events that often do not involve Ig loci [37]. They are rare or absent in MGUS, but occur in 15 % of newly diagnosed tumors, 50 % of advanced tumors, and 90 % of HMCLs [11, 38]. A recent report suggests that a small molecule inhibitor of BRD4 can inhibit MYC RNA expression in MM, with therapeutic effect [39].
Chromosome 13 deletion
A recent study concludes that chromosome 13 deletion can be an early event in MGUS (e.g., in MAF, MMSET tumors) or a progression event (e.g., in t(11;14) tumors) [40]. The pathogenic effect of this chromosome deletion is unknown, though it is possible that haploinsufficiency of RB1 promotes tumorigenesis [13]. A recent genome wide sequencing study identified mutations of DIS3, a gene of unknown function on 13q, in about 10 % of MM. Although only very few mutations have been reported to date, it has been suggested that DIS3 mutation occurs in parallel with deletions of RB1 [41], suggesting a possible dependence between these two events. Although del13 was initially reported to be an independent prognostic factor, it is now accepted only when detected by conventional cytogenetics in the more proliferative cells.
Activating mutations of RAS and BRAF
The prevalence of activating NRAS or KRAS mutations is about 15–18 % each in newly diagnosed and relapsed MM tumors [13, 42], but substantially higher in tumors that express CCND1 compared to tumors that express CCND2. For MGUS tumors, the prevalence of NRAS mutations is 7 %, but KRAS mutations have not been described [43]. This is consistent with increasing evidence that NRAS and KRAS mutations have overlapping but non-identical effects [44], and also the hypothesis that KRAS mutations provide a molecular mark of the transition of MGUS to MM [45, 46]. MM tumors depend on the continued expression of activated but not wild-type RAS [47]. Recently, BRAF mutations were described in 4 % of MM tumors, suggesting a possible role for BRAF inhibitors in these cases [48].
Activating mutations of NFkappaB pathway
Extrinsic ligands (APRIL and BAFF) produced by BM stromal cells provide critical survival signals to long-lived PCs by stimulating TACI, BCMA, and BAFF receptors to activate the NFKB pathways [49]. Most MGUS and MM tumors highly express NFKB target genes, suggesting a continued role of extrinsic signaling in PC tumors [50, 51] and at least in part explaining the constant dependency of MM cells on the BM microenvironment. Activating mutations in positive regulators and inactivating mutations in negative regulators of the NFKB pathway have been identified in at least 20 % of untreated MM tumors and ~50 % of HMCLs, rendering the cells less dependent on ligand-mediated NFKB activation [48] and most likely contributing to extra-medullary spread of the disease. Interestingly, the NFKB negative regulator TRAF3 located on 14q32 is inactivated in >10 % MM tumors, suggesting that at least in the presence of RAS/BRAF compensating mutation there may be an advantage for t(4;14) MM to lose the der(14) containing FGFR3 in favor of activating the NFKB pathway. Small molecules that inhibit extrinsic signaling [including TACI.Fc, IKKβ, and NIK (MAP3K14)] are being developed as potential therapeutic agents [52, 53]. There also is some evidence suggesting that cells addicted to constitutive NFKB activation may be particularly sensitive to proteasome inhibition [51].
Chromosome 17p loss and abnormalities of TP53
Deletions that include the TP53 locus occur in ~10 % of untreated MM tumors, and the prevalence increases with disease stage [13, 54]. TP53 mutations were present in 37 % of untreated MM tumors with del17p, but not in patients without del17p [55]. Even in the absence of TP53 mutations, del17p remains a strong independent negative predictor for survival of MM patients, although it remains to be determined if the poor prognosis is due to haploinsufficiency or to predisposition to complete inactivation of TP53 eventually occurring with tumor progression. Recently, decreased expression of microRNAs miR-199, -192, and -215 in MM was reported to increase MDM2, an inhibitor of TP53 [56], contributing to loss of p53 activity.
Gain of chromosome 1q and loss of chromosome 1p
These genomic events frequently occur together in MM, and each of them is associated with a poor prognosis [13, 57]. The relevant genes on 1q are unclear at this time although the anti-apoptotic gene MCL1 has been suggested as a potential driver of the adverse survival. By contrast, there are potential targets on two regions of 1p that are associated with a poor prognosis: CDKN2C (p18INK4c) at 1p32.3 and FAM46C at 1p12 [58, 59]. Homozygous deletion of the cell cycle regulator CDKN2C, which is present in about 30 % of HMCL and about 5 % of untreated MM tumors, is associated with increased proliferation and a poor prognosis, whereas monoallelic deletion is not. Mutations of FAM46C—often with hemizygous deletion—were identified in 3.4 and 13 % of MM tumors in two studies, and in 25 % of 16 HMCL, although the function of this gene is still unknown [48, 58].
Other pathogenic events
Secondary Ig translocations, including most IgK and IgL translocations and IgH translocations not involving one of the seven primary partners, can occur at all stages of disease, and with a similar frequency in HRD and NHRD tumors, but apart from MYC, few partner loci have been identified [11]. Other genomic rearrangements are frequent, but only a few specific target genes have been identified [57, 60, 61]. Changes in DNA methylation are frequent, with one study suggesting that a marked increase in hypomethylation is associated with the MGUS to MM transition [62], whereas a second study suggests only a small increase in hypomethylation for MM compared to MGUS [63]. Mutations in seven genes regulating RNA metabolism, protein translation and homeostasis were identified in 16 of 38 patients [48]. In addition to previous studies implicating roles for MMSET and KDM6A (UTX), genomic sequencing studies found that other histone modifying enzymes are frequent targets of mutation, although the epigenetic consequences are unknown [48]. Similarly, changes in microRNA expression at different stages have been identified, but more extensive studies are needed [56, 64].
High-risk MM is associated to intra-clonal tumor heterogeneity
Recent evidences suggest that tumor heterogeneity is prevalent in MM, as in many other cancers, and that different subclones are present within the tumor population, characterized by distinct genetic mutations that contributed independently to the tumor progression [41, 61, 65]. Recently a high level of intra-clonal tumor heterogeneity has been described in some patients with high-risk MM [41, 61, 65] associated in one case with alternating clonal dominance under therapeutic selective pressure, observations with important clinical implications. The findings suggest a competition between subclones for limited resources and raise the possibility that early, suboptimal treatment may eradicate the “good” drug-sensitive clone, making room for the “bad” drug-resistant clone to expand. They support the use of aggressive multi-drug combination approaches for high-risk disease with unstable genomes and clonal heterogeneity, and sequential one- or two-drug approaches for low-risk disease with stable genomes and lacking clonal heterogeneity.
Clinical implications of the molecular classification of multiple myeloma
The t(4;14) chromosome translocation is the genetic event in MM with the most important clinical significance. It is a poor prognostic factor for patients treated with alkylating agents, IMiDs, and bortezomib [66–69]. However, there is a survival advantage to the upfront use of bortezomib versus control in these patients [68, 70, 71], with a suggestion that prolonged use may totally overcome the adverse prognosis [71]. Despite numerous randomized clinical trials of IMiDs compared to control in the treatment of thousands of MM patients in which several studies showed improvements in overall survival (OS) for the cohort as a whole, we do not know which molecular subgroups received the maximum benefit from IMiDs versus those that received no benefit, or those that may have been harmed. From all of these studies there are a few reports of the effects of IMiDs versus control on the survival of a molecular subgroup (Table 1).
In TT2 the OS advantage of thalidomide versus placebo appeared confined to the 23 % of patients with both GEP-defined low-risk disease and metaphase cytogenetic abnormalities [72]. In contrast in the MRC-IX study the 44 % of patients with unfavorable cytogenetics [t(4;14), t(14;16), t(14;20), gain(1q21), del(1p32), del(17p)] randomized to thalidomide maintenance saw no prolongation of PFS, and the OS was significantly shorter than those randomized to placebo [73]. In the IFM 99-02 trial, the patients with del13 randomized to thalidomide maintenance saw no prolongation of EFS, but the OS was not reported [74]. In the non-intensive pathway of MRC-IX, CTDa vs MP, a trend to improved OS with thalidomide induction was noted only in the favorable cytogenetics group, but the OS for the unfavorable cytogenetic group was not reported [75]. In the Spanish study of VTD vs VMP, the non-hyperdiploid [which includes the high-risk t(4;14), t(14;16) and t(14;20)] patients randomized to thalidomide induction had significantly shorter 3-year OS (53 vs 72 %, p = 0.02) [76]. The HOVON-65/GMMG-HD4 trial randomized patients to one of two pathways: vincristine–adriamycin–dexamethasone induction, followed by high-dose melphalan and thalidomide maintenance versus bortezomib–doxorubicin–dexamethasone induction followed by high-dose melphalan and bortezomib maintenance. They noted a shorter 3-year OS for the patients randomized to the thalidomide arm with del13 (61 vs 81 %, p = 0.07), t(4;14) (44 vs 66 %, p = 0.37) and del17p (17 vs 69 %, p = 0.28) [77]. There are no data regarding the OS of different cytogenetic subgroups randomized to lenalidomide vs placebo. In the IFM 2005 study both the del13 and del17p patients randomized to lenalidomide maintenance had a significant improvement in PFS, but only a very minimal effect was seen in the t(4;14) [78, 79]. In summary therefore it appears that the maximum benefit of thalidomide is seen in the good-risk patients, whereas no benefit and sometimes worse outcomes are seen with its use in poor-risk patients. Further studies are urgently required to define the utility and safety of IMiDs in the various molecular subtypes of MM.
The MF molecular subgroups, t(14;16) and t(14;20), have each individually been associated with a poor prognosis [33, 80], although not seen for the t(14;16) in one study [81]. In addition del17p is universally associated with poor prognosis [57, 68]. Finally patients defined as high-risk by a GEP index of proliferation [82] or other GEP-defined risk scores [83, 84] (which all appear to discriminate prognosis equally in an independent dataset [82]) do poorly. Unlike the t(4;14), for these latter subgroups neither bortezomib nor any other intervention has been shown to offer a survival advantage, although the data are unfortunately very limited. These patients should be considered for clinical trials exploring innovative approaches.
Based on all of these considerations, the hematologists at the Mayo Clinic have proposed a risk-adapted strategy for the treatment of patients that cannot be enrolled on clinical trials (Fig. 3) [85]. The standard-risk patients can be treated with lenalidomide and low dose dexamethasone, postponing the toxicity and inconvenience associated with bortezomib. In contrast the t(4;14) receives bortezomib as part of induction and maintenance for at least 1 year. Finally a combination of lenalidomide, bortezomib and dexamethasone with a goal of CR is recommended for the high-risk patients.
mSMART recommendations for a risk-adapted approach to therapy. Clinical trials strongly recommended as the first option. *Note that a subset of patients with these factors will be classified as high-risk by GEP, LDH > ULN and beta-2 M > 5.5 also may indicate worse prognosis, and prognosis is worse when associated with high beta-2 microglobulin and anemia. a Bortezomib containing regimens preferred in renal failure or if rapid response needed. b If age >65 or >4 cycles of Rd Consider G-CSF plus cytoxan or plerixafor. c Continuing Rd is option for patients responding to Rd and with low toxicities; Dex is usually discontinued after first year. d In patients treated with Rd, continuing treatment is an option for patients responding well with low toxicities; Dex is usually discontinued after first year
Conclusion
Significant progress has been made is understanding the molecular pathogenesis and biology of MM. Oncogenic pathways can be activated through cell intrinsic or extrinsic mechanisms. Similar to other cancers, MM is characterized by multi-stage accumulation of genetic abnormalities deregulating different pathways. Much of this knowledge is already being utilized for diagnosis, prognosis and risk-stratification of patients. Importantly, from a clinical standpoint, this knowledge has led to development of novel therapeutic strategies, some of which are already in clinical use, and many others showing promise in pre-clinical and early clinical studies.
References
Malpas JS, Bergsagel DE, Kyle R, Anderson K. Multiple myeloma: biology and management. Oxford: Oxford University Press; 2004.
Howlader N, Noone AM, Krapcho M, Neyman N, Aminou R, Altekruse SF, Kosary CL, Ruhl J, Tatalovich Z, Cho H, Mariotto A, Eisner MP, Lewis DR, Chen HS, Feuer EJ, Cronin KA (eds). SEER cancer statistics review 1975–2009 (Vintage 2009 Populations). Bethesda: National Cancer Institute; 2012. http://seer.cancer.gov/csr/1975_2009_pops09/.
Siegel R, Naishadham D, Jemal A. Cancer statistics. CA Cancer J Clin. 2013;63(1):11–30.
Avet-Loiseau H, Attal M, Campion L, Caillot D, Hulin C, Marit G, et al. Long-term analysis of the IFM 99 trials for myeloma: cytogenetic abnormalities [t(4;14), del(17p), 1q gains] play a major role in defining long-term survival. J Clin Oncol. 2012;30(16):1949–52.
Kuehl WM, Bergsagel PL. Multiple myeloma: evolving genetic events and host interactions. Nat Rev Cancer. 2002;2(3):175–87.
Landgren O, Kyle RA, Pfeiffer RM, Katzmann JA, Caporaso NE, Hayes RB, et al. Monoclonal gammopathy of undetermined significance (MGUS) consistently precedes multiple myeloma: a prospective study. Blood. 2009;113(22):5412–7.
Weiss BM, Abadie J, Verma P, Howard RS, Kuehl WM. A monoclonal gammopathy precedes multiple myeloma in most patients. Blood. 2009;113(22):5418–22.
Dispenzieri A, Katzmann JA, Kyle RA, Larson DR, Melton LJ, Colby CL, et al. Prevalence and risk of progression of light-chain monoclonal gammopathy of undetermined significance: a retrospective population-based cohort study. Lancet. 2010;375(9727):1721–8.
Kyle RA, Therneau TM, Rajkumar SV, Larson DR, Plevak MF, Offord JR, et al. Prevalence of monoclonal gammopathy of undetermined significance. N Engl J Med. 2006;354(13):1362–9.
Drexler HG, Matsuo Y. Malignant hematopoietic cell lines: in vitro models for the study of multiple myeloma and plasma cell leukemia. Leuk Res. 2000;24(8):681–703.
Gabrea A, Martelli ML, Qi Y, Roschke A, Barlogie B, Shaughnessy JD Jr, et al. Secondary genomic rearrangements involving immunoglobulin or MYC loci show similar prevalences in hyperdiploid and nonhyperdiploid myeloma tumors. Genes Chromosom Cancer. 2008;47(7):573–90.
Bergsagel PL, Kuehl WM. Chromosome translocations in multiple myeloma. Oncogene. 2001;20(40):5611–22.
Fonseca R, Bergsagel PL, Drach J, Shaughnessy J, Gutierrez N, Stewart AK, et al. International myeloma working group molecular classification of multiple myeloma: spotlight review. Leukemia. 2009;23(12):2210–21.
Avet-Loiseau H, Facon T, Daviet A, Godon C, Rapp MJ, Harousseau JL, et al. 14q32 translocations and monosomy 13 observed in monoclonal gammopathy of undetermined significance delineate a multistep process for the oncogenesis of multiple myeloma. Intergroupe Francophone du Myelome. Cancer Res. 1999;59(18):4546–50.
Bergsagel PL, Kuehl WM, Zhan F, Sawyer J, Barlogie B, Shaughnessy J Jr. Cyclin D dysregulation: an early and unifying pathogenic event in multiple myeloma. Blood. 2005;106(1):296–303.
Hurt EM, Wiestner A, Rosenwald A, Shaffer AL, Campo E, Grogan T, et al. Overexpression of c-maf is a frequent oncogenic event in multiple myeloma that promotes proliferation and pathological interactions with bone marrow stroma. Cancer Cell. 2004;5(2):191–9.
Zingone A, Cultraro CM, Shin D-M, Bean CM, Morse HC, Janz S, et al. Ectopic expression of wild-type FGFR3 cooperates with MYC to accelerate development of B-cell lineage neoplasms. Leukemia. 2010;24(6):1171–8.
Qing J, Du X, Chen Y, Chan P, Li H, Wu P, et al. Antibody-based targeting of FGFR3 in bladder carcinoma and t(4;14)-positive multiple myeloma in mice. J Clin Invest. 2009;119(5):1216–29.
Trudel S, Li ZH, Wei E, Wiesmann M, Chang H, Chen C, et al. CHIR-258, a novel, multitargeted tyrosine kinase inhibitor for the potential treatment of t(4;14) multiple myeloma. Blood. 2005;105(7):2941–8.
Marango J, Shimoyama M, Nishio H, Meyer JA, Min D-J, Sirulnik A, et al. The MMSET protein is a histone methyltransferase with characteristics of a transcriptional corepressor. Blood. 2008;111(6):3145–54.
Martinez-Garcia E, Popovic R, Min D-J, Sweet SMM, Thomas PM, Zamdborg L, et al. The MMSET histone methyl transferase switches global histone methylation and alters gene expression in t(4;14) multiple myeloma cells. Blood. 2011;117(1):211–20.
Kuo AJ, Cheung P, Chen K, Zee BM, Kioi M, Lauring J, et al. NSD2 links dimethylation of histone H3 at lysine 36 to oncogenic programming. Mol Cell. 2011;44(4):609–20.
Pei H, Zhang L, Luo K, Qin Y, Chesi M, Fei F, et al. MMSET regulates histone H4K20 methylation and 53BP1 accumulation at DNA damage sites. Nature. 2011;470(7332):124–8.
Min DJ, Ezponda T, Kim MK, Will CM, Martinez-Garcia E, Popovic R, et al. MMSET stimulates myeloma cell growth through microRNA-mediated modulation of c-MYC. Leukemia. 2012. doi:10.1038/leu.2012.269.
Asangani IA, Ateeq B, Cao Q, Dodson L, Pandhi M, Kunju LP, et al. Characterization of the EZH2-MMSET histone methyltransferase regulatory axis in cancer. Mol Cell. 2013;49(1):80–93.
Hudlebusch HR, Santoni-Rugiu E, Simon R, Ralfkiaer E, Rossing HH, Johansen JV, et al. The histone methyltransferase and putative oncoprotein MMSET is overexpressed in a large variety of human tumors. Clin Cancer Res. 2011;17(9):2919–33.
Ezponda T, Popovic R, Shah MY, Martinez-Garcia E, Zheng Y, Min DJ, et al. The histone methyltransferase MMSET/WHSC1 activates TWIST1 to promote an epithelial-mesenchymal transition and invasive properties of prostate cancer. Oncogene. 2012.
Yang P, Guo L, Duan ZJ, Tepper CG, Xue L, Chen X, et al. Histone methyltransferase NSD2/MMSET mediates constitutive NF-kappaB signaling for cancer cell proliferation, survival, and tumor growth via a feed-forward loop. Mol Cell Biol. 2012;32(15):3121–31.
Fonseca R, Debes-Marun CS, Picken EB, Dewald GW, Bryant SC, Winkler JM, et al. The recurrent IgH translocations are highly associated with nonhyperdiploid variant multiple myeloma. Blood. 2003;102(7):2562–7.
Kumar S, Fonseca R, Ketterling RP, Dispenzieri A, Lacy MQ, Gertz MA, et al. Trisomies in multiple myeloma: impact on survival in patients with high-risk cytogenetics. Blood. 2012;119(9):2100–5.
Zhan F, Huang Y, Colla S, Stewart JP, Hanamura I, Gupta S, et al. The molecular classification of multiple myeloma. Blood. 2006;108(6):2020–8.
Broyl A, Hose D, Lokhorst H, de Knegt Y, Peeters J, Jauch A, et al. Gene expression profiling for molecular classification of multiple myeloma in newly diagnosed patients. Blood. 2010;116(14):2543–53.
Nair B, van Rhee F, Shaughnessy JD, Anaissie E, Szymonifka J, Hoering A, et al. Superior results of total therapy 3 (2003–33) in gene expression profiling-defined low-risk multiple myeloma confirmed in subsequent trial 2006–66 with VRD maintenance. Blood. 2010;115(21):4168–73.
Chng WJ, Huang GF, Chung T-H, Ng SB, Gonzalez-Paz N, Troska-Price T, et al. Clinical and biological implications of MYC activation: a common difference between MGUS and newly diagnosed multiple myeloma. Leukemia. 2011;25(6):1026–35.
Chesi M, Robbiani DF, Sebag M, Chng WJ, Affer M, Tiedemann R, et al. AID-dependent activation of a MYC transgene induces multiple myeloma in a conditional mouse model of post-germinal center malignancies. Cancer Cell. 2008;13(2):167–80.
Chesi M, Matthews GM, Garbitt VM, Palmer SE, Shortt J, Lefebure M, et al. Drug response in a genetically engineered mouse model of multiple myeloma is predictive of clinical efficacy. Blood. 2012;120(2):376–85.
Dib A, Gabrea A, Glebov OK, Bergsagel PL, Kuehl WM. Characterization of MYC translocations in multiple myeloma cell lines. J Natl Cancer Inst. 2008;39:25–31.
Avet-Loiseau H, Gerson F, Magrangeas F, Minvielle S, Harousseau JL, Bataille R. Rearrangements of the c-myc oncogene are present in 15 % of primary human multiple myeloma tumors. Blood. 2001;98(10):3082–6.
Delmore JE, Issa GC, Lemieux ME, Rahl PB, Shi J, Jacobs HM, et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell. 2011;146(6):904–17.
Chiecchio L, Dagrada GP, Ibrahim AH, Dachs Cabanas E, Protheroe RK, Stockley DM, et al. Timing of acquisition of deletion 13 in plasma cell dyscrasias is dependent on genetic context. Haematologica. 2009;94(12):1708–13.
Walker BA, Wardell CP, Melchor L, Hulkki S, Potter NE, Johnson DC, et al. Intraclonal heterogeneity and distinct molecular mechanisms characterize the development of t(4;14) and t(11;14) myeloma. Blood. 2012;120(5):1077–86.
Chng WJ, Gonzalez-Paz N, Price-Troska T, Jacobus S, Rajkumar SV, Oken MM, et al. Clinical and biological significance of RAS mutations in multiple myeloma. Leukemia. 2008;22(12):2280–4.
Zingone A, Kuehl WM. Pathogenesis of monoclonal gammopathy of undetermined significance and progression to multiple myeloma. Semin Hematol. 2011;48(1):4–12.
Haigis KM, Kendall KR, Wang Y, Cheung A, Haigis MC, Glickman JN, et al. Differential effects of oncogenic K-Ras and N-Ras on proliferation, differentiation and tumor progression in the colon. Nat Genet. 2008;40(5):600–8.
Rasmussen T, Haaber J, Dahl IM, Knudsen LM, Kerndrup GB, Lodahl M, et al. Identification of translocation products but not K-RAS mutations in memory B cells from patients with multiple myeloma. Haematologica. 2010;95(10):1730–7.
Rasmussen T, Kuehl M, Lodahl M, Johnsen HE, Dahl IMS. Possible roles for activating RAS mutations in the MGUS to MM transition and in the intramedullary to extramedullary transition in some plasma cell tumors. Blood. 2005;105(1):317–23.
Steinbrunn T, Stühmer T, Gattenlöhner S, Rosenwald A, Mottok A, Unzicker C, et al. Mutated RAS and constitutively activated Akt delineate distinct oncogenic pathways, which independently contribute to multiple myeloma cell survival. Blood. 2011;117(6):1998–2004.
Chapman MA, Lawrence MS, Keats JJ, Cibulskis K, Sougnez C, Schinzel AC, et al. Initial genome sequencing and analysis of multiple myeloma. Nature. 2011;471(7339):467–72.
Elgueta R, de Vries VC, Noelle RJ. The immortality of humoral immunity. Immunol Rev. 2011;236:139–50.
Annunziata CM, Davis RE, Demchenko Y, Bellamy W, Gabrea A, Zhan F, et al. Frequent engagement of the classical and alternative NF-kappaB pathways by diverse genetic abnormalities in multiple myeloma. Cancer Cell. 2007;12(2):115–30.
Keats JJ, Fonseca R, Chesi M, Schop R, Baker A, Chng WJ, et al. Promiscuous mutations activate the noncanonical NF-kappaB pathway in multiple myeloma. Cancer Cell. 2007;12(2):131–44.
Demchenko YN, Glebov OK, Zingone A, Keats JJ, Bergsagel PL, Kuehl WM. Classical and/or alternative NF-kappaB pathway activation in multiple myeloma. Blood. 2010;115(17):3541–52.
Rossi JF, Moreaux J, Hose D, Requirand G, Rose M, Rouillé V, et al. Atacicept in relapsed/refractory multiple myeloma or active Waldenström’s macroglobulinemia: a phase I study. Br J Cancer. 2009;101(7):1051–8.
Tiedemann RE, Gonzalez-Paz N, Kyle RA, Santana-Davila R, Price-Troska T, Van Wier SA, et al. Genetic aberrations and survival in plasma cell leukemia. Leukemia. 2008;22(5):1044–52.
Lode L, Eveillard M, Trichet V, Soussi T, Wuilleme S, Richebourg S, et al. Mutations in TP53 are exclusively associated with del(17p) in multiple myeloma. Haematologica. 2010;95(11):1973–6.
Pichiorri F, Suh S–S, Rocci A, De Luca L, Taccioli C, Santhanam R, et al. Downregulation of p53-inducible microRNAs 192, 194, and 215 impairs the p53/MDM2 autoregulatory loop in multiple myeloma development. Cancer Cell. 2010;18(4):367–81.
Walker BA, Leone PE, Chiecchio L, Dickens NJ, Jenner MW, Boyd KD, et al. A compendium of myeloma-associated chromosomal copy number abnormalities and their prognostic value. Blood. 2010;116(15):e56–65.
Boyd KD, Ross FM, Walker BA, Wardell CP, Tapper WJ, Chiecchio L, et al. Mapping of chromosome 1p deletions in myeloma identifies FAM46C at 1p12 and CDKN2C at 1p32.3 as being genes in regions associated with adverse survival. Clin Cancer Res. 2011;17(24):7776–84.
Dib A, Peterson TR, Raducha-Grace L, Zingone A, Zhan F, Hanamura I, et al. Paradoxical expression of INK4c in proliferative multiple myeloma tumors: bi-allelic deletion vs increased expression. Cell Div. 2006;1:23.
Avet-Loiseau H, Li C, Magrangeas F, Gouraud W, Charbonnel C, Harousseau J-L, et al. Prognostic significance of copy-number alterations in multiple myeloma. J Clin Oncol. 2009;27(27):4585–90.
Keats JJ, Chesi M, Egan JB, Garbitt VM, Palmer SE, Braggio E, et al. Clonal competition with alternating dominance in multiple myeloma. Blood. 2012;120(5):1067–76.
Walker BA, Wardell CP, Chiecchio L, Smith EM, Boyd KD, Neri A, et al. Aberrant global methylation patterns affect the molecular pathogenesis and prognosis of multiple myeloma. Blood. 2011;117(2):553–62.
Salhia B, Baker A, Ahmann G, Auclair D, Fonseca R, Carpten J. DNA methylation analysis determines the high frequency of genic hypomethylation and low frequency of hypermethylation events in plasma cell tumors. Cancer Res. 2010;70(17):6934–44.
Pichiorri F, Suh SS, Ladetto M, Kuehl M, Palumbo T, Drandi D, et al. MicroRNAs regulate critical genes associated with multiple myeloma pathogenesis. Proc Natl Acad Sci USA. 2008;105(35):12885–90.
Magrangeas F, Avet-Loiseau H, Gouraud W, Lode L, Decaux O, Godmer P, et al. Minor clone provides a reservoir for relapse in multiple myeloma. Leukemia. 2013;27:473–81.
Avet-Loiseau H, Attal M, Moreau P, Charbonnel C, Garban F, Hulin C, et al. Genetic abnormalities and survival in multiple myeloma: the experience of the Intergroupe Francophone du Myélome. Blood. 2007;109(8):3489–95.
Kapoor P, Kumar S, Fonseca R, Lacy MQ, Witzig TE, Hayman SR, et al. Impact of risk stratification on outcome among patients with multiple myeloma receiving initial therapy with lenalidomide and dexamethasone. Blood. 2009;114(3):518–21.
Avet-Loiseau H, Leleu X, Roussel M, Moreau P, Guerin-Charbonnel C, Caillot D, et al. Bortezomib plus dexamethasone induction improves outcome of patients with t(4;14) myeloma but not outcome of patients with del(17p). J Clin Oncol. 2010;28(30):4630–4.
Sonneveld P, Schmidt-Wolf IG, van der Holt B, El Jarari L, Bertsch U, Salwender H, et al. Bortezomib induction and maintenance treatment in patients with newly diagnosed multiple myeloma: results of the randomized phase III HOVON-65/GMMG-HD4 trial. J Clin Oncol. 2012;30(24):2946–55.
San-Miguel JF, Schlag R, Khuageva NK, Dimopoulos MA, Shpilberg O, Kropff M, et al. Bortezomib plus melphalan and prednisone for initial treatment of multiple myeloma. N Engl J Med. 2008;359(9):906–17.
Pineda-Roman M, Zangari M, Haessler J, Anaissie E, Tricot G, van Rhee F, et al. Sustained complete remissions in multiple myeloma linked to bortezomib in total therapy 3: comparison with total therapy 2. Br J Haematol. 2008;140(6):625–34.
Barlogie B, Pineda-Roman M, van Rhee F, Haessler J, Anaissie E, Hollmig K, et al. Thalidomide arm of total therapy 2 improves complete remission duration and survival in myeloma patients with metaphase cytogenetic abnormalities. Blood. 2008;112(8):3115–21.
Morgan GJ, Gregory WM, Davies FE, Bell SE, Szubert AJ, Brown JM, et al. The role of maintenance thalidomide therapy in multiple myeloma: MRC Myeloma IX results and meta-analysis. Blood. 2012;119(1):7–15.
Attal M, Harousseau JL, Leyvraz S, Doyen C, Hulin C, Benboubker L, et al. Maintenance therapy with thalidomide improves survival in patients with multiple myeloma. Blood. 2006;108(10):3289–94.
Morgan GJ, Davies FE, Gregory WM, Russell NH, Bell SE, Szubert AJ, et al. Cyclophosphamide, thalidomide, and dexamethasone (CTD) as initial therapy for patients with multiple myeloma unsuitable for autologous transplantation. Blood. 2011;118(5):1231–8.
Mateos M-V, Oriol A, Martínez-López J, Gutiérrez N, Teruel A-I, de Paz R, et al. Bortezomib, melphalan, and prednisone versus bortezomib, thalidomide, and prednisone as induction therapy followed by maintenance treatment with bortezomib and thalidomide versus bortezomib and prednisone in elderly patients with untreated multiple myeloma: a randomised trial. Lancet Oncol. 2010;11(10):934–41.
Neben K, Lokhorst HM, Jauch A, Bertsch U, Hielscher T, van der Holt B, et al. Administration of bortezomib before and after autologous stem-cell transplantation improves outcome in multiple myeloma patients with deletion 17p. Blood. 2012;119(4):940–8.
Avet-Loiseau H, Caillot D, Marit G, Lauwers-Cances V, Roussel M, Facon T, et al. Long-term maintenance with lenalidomide improves progression free survival in myeloma patients with high-risk cytogenetics: An IFM study. ASH Annual Meeting Abstracts. 2010;116(21):1944.
Attal M, Lauwers-Cances V, Marit G, Caillot D, Moreau P, Facon T, et al. Lenalidomide maintenance after stem-cell transplantation for multiple myeloma. N Engl J Med. 2012;366(19):1782–91.
Ross FM, Chiecchio L, Dagrada G, Protheroe RKM, Stockley DM, Harrison CJ, et al. The t(14;20) is a poor prognostic factor in myeloma but is associated with long-term stable disease in monoclonal gammopathies of undetermined significance. Haematologica. 2010;95(7):1221–5.
Avet-Loiseau H, Malard F, Campion L, Magrangeas F, Sebban C, Lioure B, et al. Translocation t(14;16) and multiple myeloma: is it really an independent prognostic factor? Blood. 2011;117(6):2009–11.
Hose D, Reme T, Hielscher T, Moreaux J, Messner T, Seckinger A, et al. Proliferation is a central independent prognostic factor and target for personalized and risk-adapted treatment in multiple myeloma. Haematologica. 2011;96(1):87–95.
Shaughnessy JD, Zhan F Jr, Burington BE, Huang Y, Colla S, Hanamura I, et al. A validated gene expression model of high-risk multiple myeloma is defined by deregulated expression of genes mapping to chromosome 1. Blood. 2007;109(6):2276–84.
Decaux O, Lode L, Magrangeas F, Charbonnel C, Gouraud W, Jézéquel P, et al. Prediction of survival in multiple myeloma based on gene expression profiles reveals cell cycle and chromosomal instability signatures in high-risk patients and hyperdiploid signatures in low-risk patients: a study of the Intergroupe Francophone du Myélome. J Clin Oncol. 2008;26(29):4798–805.
Kumar SK, Mikhael JR, Buadi FK, Dingli D, Dispenzieri A, Fonseca R, et al. Management of newly diagnosed symptomatic multiple myeloma: updated Mayo Stratification of Myeloma and Risk-Adapted Therapy (mSMART) consensus guidelines. Mayo Clin Proc. 2009;84(12):1095–110.
Morgan GJ, Davies FE, Gregory WM, Bell SE, Szubert AJ, Navarro-Coy N, et al. Cyclophosphamide, thalidomide, and dexamethasone as induction therapy for newly diagnosed multiple myeloma patients destined for autologous stem-cell transplantation: MRC Myeloma IX randomized trial results. Haematologica. 2012;97(3):442–50.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Chesi, M., Bergsagel, P.L. Molecular pathogenesis of multiple myeloma: basic and clinical updates. Int J Hematol 97, 313–323 (2013). https://doi.org/10.1007/s12185-013-1291-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12185-013-1291-2