Abstract
Bee pollen is one of the hive products that is of most interest today due to its multiple beneficial health properties, making it an increasingly popular food supplement. Bee pollen contains many bioactive compounds, such as fatty acids, vitamins, minerals, proteins, and amino acids, among others. In the present study, the free amino acid content was determined in bee pollen by using liquid chromatography coupled to a fluorescence detector. Sample treatment consisted of a solvent extraction of the free amino acids with ultrapure water and a further centrifugation of the extract, which was repeated twice. After that, it was necessary to perform a pre-column derivatization of the amino acids using a combination of two reagents (o-phthalaldehyde and 9-fluorenylmethyl chloroformate) prior to their separation in a Gemini® C18 reverse phase column in gradient elution mode. The analytical performance was evaluated, and several commercial bee pollen samples were analyzed. Significant differences in the free amino acid profile and concentration, which ranged between 19 and 192 mg/g, were observed depending on the botanical origin of the samples.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bee pollen is one of the products of the hive, along with others such as royal jelly or propolis, which are gaining ground for their progressive incorporation into the daily human diet as food supplements. This is a mixture of flower pollen residues together with nectar or honey, enzymes, wax, and salivary substances from bees, creating small grains (Ares et al. 2018; Themelis et al. 2019). Bee pollen is considered one of the few foods that are perfectly complete, with at least 250 substances present in its composition; among these are sugars, lipids (triglycerides, phospholipids), carbohydrates, proteins, amino acids, vitamins A, C, E and D, minerals such as zinc, copper and iron, carotenoids, flavonoids and both macro- and micronutrients (Campos et al. 2021; Chantarudee et al. 2012; Kafantaris et al. 2021). Some of these compounds are bioactive, and in the last years many studies have described the potential benefits of bee pollen for health (Ares et al. 2018). It has been described that the consumption of bee pollen helps improve the cardiovascular system, stimulate body immunity, promote antitumor effects, delay aging, eliminate free radicals, regulate digestive functions, and treat prostate problems (Li et al. 2019). Thus, it can be used therapeutically via its consumption, but always under the supervision of a doctor or nutritionist. However, it should be mentioned that the specific composition of bee pollen is greatly dependent on its botanical and geographical origin (Prđun et al. 2021), and consequently determining bee pollen constituents could be used to ascertain its origin, nutritional value, and bioactivity. Bee pollen contains all the essential amino acids needed by humans (Ares et al. 2018; Li et al. 2019), which are those that the human body is not capable of producing autonomously. Meanwhile, non-essential amino acids are those that can be re-synthesized by animal organisms in sufficient amounts for them not having to require incorporation in the diet (Wu et al. 2013). The content of amino acids that this food supplement can provide is variable, and depends mainly, as was previously mentioned, on the botanical and geographical origin of the pollen. For this reason, and due to the current heightened interest in this product in the European socio-economic market, it is considerably important to study its profile in different bee pollens to ascertain the relationship with its floral origin, in view of the preferences that exist among sources. This can also be used to demonstrate possible adulterations in the composition of the pollen, a matter of relevance for the food industry and, in turn, to give added value to this product (Ares et al. 2018; Taha et al 2019). Amino acids have usually been determined in bee pollen by liquid chromatography (LC) in reverse phase mode (C18-based columns), with mobile phases that consisted of combinations of buffer solutions at basic pH with organic solvents (González-Paramás et al 2006; Lilek et al. 2015; Stabler et al. 2018; Themelis et al. 2019; You et al. 2007; Zhang et al. 2009); meanwhile, gas chromatography (Omar et al. 2018), ion exchange chromatography (Taha et al. 2019), colorimetric methods employing ninhydrin (Canale et al. 2016), and amino acids analyzers (Nikkeshi et al. 2021; Yang et al. 2013) have also been employed in some cases. Fluorescence detectors (FLD) have been the predominant choice for determining amino acids in bee pollen when LC is used, and in this case it was necessary to derivatize these prior to their detection. The most common derivatizing reagents are 9-fluorenylmethyl chloroformate (FMOC-Cl; Stabler et al. 2018; Themelis et al. 2019) and its analog 2-(11Hbenzo[α]-carbazol-11-yl) ethyl chloroformate (BCEC–Cl; You et al. 2007) or o-phthalaldehyde (OPA; González-Paramás et al. 2006; Stabler et al. 2018). FMOC-Cl reacts with primary and secondary amines, forming very stable derivatives with high fluorescence. This reagent exhibits natural fluorescence and may interfere with detection if the chromatographic conditions are not adequate. On the other hand, OPA reacts only with primary amines, making it impossible to detect amino acids such as proline (Pro). Regarding the sample treatment used for extracting amino acids from bee pollen, this varies depending on the studies consulted. Some authors have proposed an extraction with ultrapure water and different organic solvents, such as ethanol (Canale et al. 2016) González-Paramás et al. 2006; Nikkeshi et al. 2021), methanol (Lilek et al. 2015; Stabler et al. 2018), and acetonitrile (Zhang et al. 2009). However, when total amino acid content (not only free amino acids) was determined, it was also usually necessary to perform acid hydrolysis with hydrochloric acid so as to release the protein-bound amino acids (Stabler et al. Taha et al. 2019; Themelis et al. 2019; Yang et al. 2013; You et al. 2007). It should be noted that during acid hydrolysis, some amino acids, such as tryptophan (Trp) and methionine (Met), can be destroyed or altered (Themelis et al. 2019).
Therefore, the main goal of this paper is to propose an alternative LC-FLD method for determining 20 free amino acids in bee pollen. Specific and efficient extraction, derivatization, and determination procedures were optimized to ensure good recovery, rapidness, and maximum respect for the principles of green analytical chemistry (Gałuszka et al. 2013). This not only reduced costs but also the number and toxicity of reagents, as well as the time employed. Further aims of this work focused on evaluating the performance of the method, and free amino acid content in bee pollen samples from different botanical origins.
Materials and Methods
Chemicals and Standards
FMOC-Cl, OPA, and mercaptopropionic acid were supplied by Sigma–Aldrich Chemie Gbmh (Steinheim, Germany). LC grade methanol and acetonitrile were both obtained from Lab-Scan Ltd. (Dublin, Ireland), while sodium acetate was purchased from Carlo Erba (Barcelona, Spain). Boric acid and sodium hydroxide were provided by Merck (Darmstadt, Germany), and ultrapure water was obtained from Millipore Milli-RO plus and Milli-Q systems (Bedford, MA, USA). An Eppendorf Centrifuge 5810R (Hamburg, Germany), a Moulinette chopper device (Moulinex. Paris, France), IKA® Ultra-Turrax® T18 basic disperser (IKA®-Werke GmbH & Co. KG, Staufen, Germany), syringe filters (17 mm, Nylon 0.45 μm; Nalgene, Rochester, NY, USA), a drying oven, and a vibromatic system (J.P. Selecta S.A., Barcelona, Spain) were used for the sample treatment. Analytical standards of the investigated amino acids (alanine, ALA; arginine, ARG; asparagine, ASN; aspartic acid, ASP; glutamic acid, GLU; glutamine, GLN; glycine, GLY; histidine, HIS; isoleucine, ISO; leucine, LEU; lysine, LYS; methionine, MET; phenylalanine, PHE; proline, PRO: serine, SER; threonine, THR; tryptophan, TRP; tyrosine, TYR; and valine, VAL) as well and the internal standard (IS), ү-aminobutyric acid (GABA) were purchased from Sigma–Aldrich (St. Louis, MO, USA). Gamma-aminobutyric acid (GABA) was selected as internal standard (IS) because it is not present in bee pollen, and it forms easily detectable derivatives and does not cause interference with the rest of the amino acids. Standard stock solution was prepared by dissolving approximately 10 mg of the accurately weighed compounds in 10 mL of HCl (0.1 mol/L), and a final concentration of approximately 1000 mg/L was obtained. Standard in solvent calibration curves were employed for performing the quantification within the range of LOQ (see Table 1) to 30 mg/L (LOQ, 1.0, 5.0, 10.0, 15.0, 20.0, 25.0, and 30 mg/L). Stock, working, and calibration solutions were stored in glass containers in darkness at + 4 °C. All solutions remained stable for over 2 weeks.
Sample Procurement and Treatment
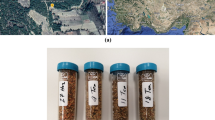
Bee pollen samples (n = 6) were kindly donated by the Center for Agroenvironmetal and Apicultural Investigation-CIAPA (Marchamalo, Guadalajara, Spain), and they were from different plant origins (multifloral, n = 4; sunflower, n = 1; maize, n = 1). Bee pollen samples were individually mixed, ground, and pooled for optimum sample homogeneity. Next, bee pollen was dried until the mass stabilized (humidity was between 9 and 12%), and subsequently it was stored in the dark at – 20 °C until analysis. Then, 0.5 g of bee pollen and 20 mL of ultrapure water were transferred to a centrifuge tube. The mixture was shaken for 90 s in the Ultra-Turrax®, and then it was centrifuged for 10 min at 11,000 rpm at 5 °C. The supernatant was collected, and the remaining solid was again extracted with other 20 mL of ultrapure water, and the procedure above-mentioned was repeated. The obtained supernatants were combined, and it was taken 1 mL that was diluted with ultrapure water (1/10; v/v). At this stage, it was added the IS at a concentration of 10 mg/L. The resulting extract was filtered through a 0.45 μm nylon filter and transferred to a 2 mL vial, which will be placed in the automatic injector for performing the derivatization procedure. OPA and FMOC-Cl were the derivatization reagents, and they were prepared as follows: (i) OPA was prepared by weighing 50 mg of reagent and dissolving it in 4.5 mL of methanol. Then, 41 μL of 3-mercaptopropionic acid were added, making up to a volume of 5 mL with 0.4 M borate buffer at pH 10.2; (ii) FMOC-Cl reagent was prepared by dissolving 2.5 mg in 5 mL of acetonitrile, with a concentration of 9.6 mM. It was also required borate buffer at pH 10.2 for the derivatization procedure, which was prepared from boric acid in ultrapure water and adjusted to pH with a sodium hydroxide solution (5 mol/L). Then, all the reagents were transferred to different 2 mL vials, in order to begin with the online derivatization procedure that is summarized in Table S1 (see Supplementary Material).
LC-FLD Conditions
An Agilent Technologies (Palo Alto, CA, USA) 1100 LC system, equipped with a vacuum degasser, a quaternary solvent pump, an autosampler, a column oven, and a FLD was employed in this study. All were controlled by an Agilent ChemStation software. A Gemini® C18 (150 × 4.6 mm; 5 μm) analytical column (Phenomenex, Torrance, CA, USA) was used for the HPLCFLD analyses and was protected with a C18 security guard cartridge (4 × 3.0 mm i.d.; Phenomenex). As a result of the findings of the optimization study, the mobile phase selected was a mixture of sodium acetate (25 mmol/L at pH 8.0), acetonitrile, and methanol, which was applied at a flow rate of 1.0 mL/min in gradient elution mode (see Table 2). The injection volume and temperature were set at 1 μL and 40 °C, respectively. It must be specified that it was employed a detection program with different wavelengths depending on the free amino acids derivatives that what formed with the derivatization reagents. Therefore, OPA derivatives were detected at 240 nm (excitation) and 450 nm (emission) from 0 to 28.5 min, while FMOC derivatives were monitored from 28.5 min at 266 nm (excitation) and 313 nm (emission).
Results and Discussion
Optimization of the Sample Treatment
As mentioned in the “Introduction” section, solvent extraction has normally been employed as a sample treatment when determining free amino acids in bee pollen. Therefore, we decided to test the suitability of three of the reagents that are more environmentally friendly, such as ultrapure water, acidified ultrapure water (HCl 0.1 mol/L), and an ethanol and ultrapure water mixture (80:20; v/v), which were chosen after performing some preliminary tests (data not shown). Moreover, the performance of two different agitation sources, namely ultrasound (15 min) and Ultra-Turrax® (90 s), was also investigated. The following conditions were employed in the previously mentioned tests: (i) 0.5 g bee pollen (multifloral); (ii) 10 mL solvent; (iii) 90 s shaking time (Ultra-Turrax®); (iv) 10 min centrifugation at 11,000 rpm and 5 °C. The criterion for establishing the optimal conditions was determined by the strongest (highest) signal, which was obtained by comparing the areas of the individual amino acids and that of the IS. All the experiments were performed in triplicate. The results showed (see Fig. 1) that the largest number of amino acids was extracted by using ultrapure water and Ultra-Turrax®, which was followed by the ethanol and water mixture with the same agitation source; meanwhile, the use of acidified ultrapure water provided the worst results. In the case of TRP, the amount extracted was significantly larger with the ethanol and water mixture (data not shown), which may be due to the less polar nature of this amino acid. Thus, ultrapure water and Ultra-Turrax® were selected for continuing with the optimization procedure, as not only did they provide the best extraction, but also because ultrapure water is the most environmentally friendly solvent that can be used. An optimization study considered the most relevant factors that could influence the amount of amino acids extracted from bee pollen samples, namely shaking time, amount of solvent, and temperature (see Supplementary Information, Table S2). In addition, a factorial design (23+1) at 2 levels was devised, the response being the ratio of areas obtained from the chromatograms in order to optimize the above-mentioned parameters. The experimental matrix for this complete factorial design was then made; this consisted of 16 experiments, although a central point with pseudo-intermediate conditions was added, leaving a total of 18 experiments (see Supplementary Information, Table S3). The results for the complete factorial design that were obtained after performing the 18 experiments are summarized in Table S4 (see Supplementary Information) and in Fig. 2, as an analysis of variance (ANOVA) and a Pareto chart, providing a graphic indication of how the factors and the interactions affect both positively and negatively the response to maximize (ratio of areas). The ANOVA and Pareto chart were obtained by means of basic but efficient statistical tools such as Microsoft Excel (Microsoft Office 2010, Microsoft Corporation, Redmond, WA, USA) and Statgraphics Centurion 19-X64 (Royal Technologies S.A., Cundinamarca, Colombia). It can be concluded that for the individual factors A (shaking time), B (solvent volume), and for the AB interaction, the value for p is less than 0.05, which translates into non-compliance with the null hypothesis, significantly influencing the measurement. These factors also have a positive influence on the response since the proposed design aims to optimize the response as far as possible. Thus, the optimal conditions of the parameters considered were 120 s shaking time and two extractions with 20 mL at 20 °C. To test whether it was necessary to carry out a specific number of successive extractions to increase the extraction yield, we assessed the performance of two, three, and four consecutive extractions on the same bee pollen sample by using the previously optimized sample treatment. The recoveries were calculated based on the area ratios obtained, after performing each set of experiments, in relation to their overall total. The results showed (see Supplementary Information, Table S5) that two extractions with 20 mL were sufficiently acceptable, as recoveries in all cases recoveries ranged between 94 and 99%; however, recoveries lower than 5% or 1% were obtained when performing a third and fourth extraction, respectively. To summarize, an alternative sample treatment has been proposed for determining free amino acids in bee pollen. It is fast (less than 25 min), simple (number of steps and reagents), and environmentally friendly (nature of reagents, ultrapure water). Nevertheless, a real comparison could not be made with most of the related studies, as the amounts of bee pollen were much lower (< 50 mg) and an acid hydrolysis was usually performed for analyzing total amino acid content.
Optimization of the Derivatization Procedure
Amino acids are non-fluorescent compounds, and, with the exception of tryptophan, they do not display a significant native fluorescence (Zhang et al. 2009). Thus, a derivatization procedure is necessary to obtain products that are stable and fluorescent for detection by FLD (You et al. 2007). The derivatization process can be carried out in two different ways: in the chromatographic equipment (pre- or post-column) or by inclusion in the sample treatment stage. In this study, we decided to perform an online pre-column derivatization with two of the most employed reagents, FMOC-Cl and OPA. This was because of a recent study by our research group (Biluca et al. 2019) and on the strength of related literature (Stabler et al. 2018). As previously mentioned, OPA is a derivatizing agent that reacts only with primary amines; hence, certain amino acids present in the samples, such as proline and cysteine, will not react with this reagent. In order to obtain the fluorescent derivatives, the reaction was carried out in an alkaline aqueous medium, and, with the presence of 3-mercaptopropionic acid, mercaptoethanol was formed; this is a reducing agent with a thiol group that displays very stable fluorescent derivatives with the AAs (González-Paramás et al. 2006). The formation of the fluorescent derivatives with this reagent is very fast and of great sensitivity. The excitation and emission wavelengths of the OPA are usually 240 nm and 450 nm, respectively (Perucho et al. 2015). Meanwhile, FMOC-CI reacts with both primary and secondary amines, obtaining stable and fluorescent derivatives (Ziegler and Abel 2014); however, unlike OPA, this reagent exhibits natural fluorescence, which can be a problem, as its excess can interfere with the analysis and quantification of amino acids. Moreover, FMOC-CI can form di-substituted derivatives with some amino acids, such as HIS and TYR, obtaining different retention times than monosubstituted ones (Walker and Mills 1995). The excitation and emission wavelengths of the FMOC derivatives are usually close to 265 nm and 310 nm, respectively (Biluca et al. 2019; Themelis et al. 2019). The online derivatization procedure was carried out immediately before injection, using an automated pre-column derivatization programme for the autosampler. Optimization of the derivatization procedure was by means of examining previous studies (Biluca et al. 2019; Stabler et al. 2018) and adapting the proposed conditions to our goals. Different tests were made to obtain the best derivatization conditions, and the final program is summarized in Table S1 (see Supplementary Information). As can be seen, the program includes some final cleaning steps to avoid a carry-over between the samples; this consisted of washing the needle several times with acetonitrile and ultrapure water.
Optimization of the LC-FLD Conditions
Firstly, eight different reverse phase LC columns with the same length and internal diameter were tested in order to select the most suitable for determining amino acids in bee pollen (see Table 3). The ones decided on as initial conditions were those proposed in our recent study (Biluca et al. 2019), in which a Gemini® C18 (150 × 4.6 mm; 5 μm) column was employed. No significant advantages in terms of amino acid separation were obtained with the other column tested compared with the latter (data not shown), and therefore we decided to continue the optimization procedure with this column. Next, several tests were conducted employing different mobile phase constituents (sodium acetate 25 mmol/L (pH 8), ammonium acetate 25 mmol/L (pH 8), acetonitrile, methanol, water), gradient elution programs, temperatures (20–45 °C), and flow rates (0.8–1.2 mL/min), with the aim of obtaining the best separation of amino acids in a shorter analysis time (data not shown). The best conditions are summarized in Table 2 and subsection LC-FLD conditions. As can be seen in Fig. 3, separation of the amino acids was achieved in less than 30 min, which is faster than previous LC methods (González-Paramás et al. 2006; Themelis et al. 2019); it should be mentioned, however, that the number of compounds under study was not the same. In addition, a post-time was not required, as the column was equilibrated with the initial mobile phase conditions, during the online derivatization procedure before the following injection.
Representative LC-FLD chromatograms obtained from multifloral bee pollen sample: A normal; B enlarged chromatogram without PRO (1. ASP; 2. GLU; 3. ASN; 4. SER; 5. GLN; 6. HIS; 7. GLY I; 8. THR; 9. ARG; 10. ALA; 11. GABA I; 12. TYR; 13. GLY II; 14. VAL; 15. MET; 16. GABA II; 17. TRIP; 18. PHE; 19. ILE; 20. LEU; 21. LYS; 22. PRO). The LC-FLD conditions are summarized in subsection LC-FLD conditions and Table 2
Analytical Performance of the Proposed Method
To determine the selectivity of the proposed method, a set of extracts of bee pollen samples (n = 3) was injected onto the chromatographic system, and the results were compared with those obtained for the individual standards of the amino acids under study. It was observed that the retention times matched perfectly in all cases and a great similarity in the FLD spectra in standard and bee pollen samples (data not shown). The limits of detection (LODs) and quantification (LOQs) were experimentally determined, and they were estimated to be three and ten times the signal-to-noise (S/N) ratio, respectively (see Table 1). To this regard, the noise was assessed as the distribution of the response at zero analytes concentration. The obtained LODs and LOQs values are similar to those reported in previous works (González-Paramás et al. 2006; Themelis et al. 2019; You et al. 2007). Calibration curves were constructed by plotting the signal on the y-axis (analyte peak area/IS area) against the analyte concentration on the x-axis. The graphs obtained in all the calibration curves, which had a wide calibration range (LOQ-30 mg/L) were straight lines, with coefficient of the determination values (R2) higher than 0.99 in all cases (see Table 1), and the residual analysis revealed a random scatter with no systematic trend (data not shown). Precision, which was expressed as relative standard deviation (%RSD), experiments were performed concurrently by repeated analysis using a standard solution of the mixture of all the amino acids (5 mg/L) and bee pollen samples (n = 6; intra-day precision), or over three consecutive days (n = 6; inter-day precision). The obtained %RSD values for the areas and retention times were lower or equal than 8% in all cases (data not shown).
Application of the Method
Several bee pollen samples were analyzed (see subsection “Sample procurement and treatment”), and the results are summarized in Table 4. Significant differences can be observed in amino acid content and profile depending on the origin and sample, although a common finding was that PRO was always detected at the highest concentration rate in all the samples, but in a wide concentration range (3–55 mg/g). The prevalence of PRO as a major amino acid and its variable concentrations have generally been reported (González-Paramás et al. 2006; Themelis et al. 2019; You et al. 2007), and this can be explained by the fact that PRO is also contributed by bees, and not only from the original pollen from the plant (González-Paramás et al. 2006). Yet in two samples from different botanical origins (maize and multifloral-4), six amino acids could not be quantified, while in only one sample, multifloral-1, all the amino acids studied were detected and quantified. The total amount of amino acids (the sum of each of the compounds) in the samples was in the range of 19–192 mg/g. The highest level of amino acids was found in multifloral-3 sample (192 mg/g), whose content of histidine (22.2 mg/g) and glycine (18.0 mg/g) was particularly high; meanwhile, multifloral-2 sample exhibited the lowest total amount of amino acids (19.0 mg/g). These findings should only be considered tentative ones, as the number of samples studied was insufficient, and the specific origin of the multifloral samples was unknown. Finally, the amino acid concentrations and their differences depending on the origin are in good agreement with existing data. This can be explained by the fact that amino acid composition is strongly dependent on the plant’s origin, among several other factors (Themelis et al. 2019). Therefore, these results reinforce the potential role of amino acids as biomarkers of bee pollen plant origin.
Conclusions
A novel LC-FLD method was developed for determining free amino acids in bee pollen. This is one the fastest proposals ever published in terms of separation time, as 19 free amino acids were separated in less than 35 min. Extraction was carried out with ultrapure water as the extractant, a stirring time of 90 s, by means of an Ultra-Turrax® homogenizer, followed by a centrifugation step. The total amount of extractant was 40 mL, divided into two consecutive extractions with 20 mL of ultrapure water. The proposed conditions are in good agreement with some of the principles of green Analytical Chemistry. Two derivatizing agents were used for the pre-column derivatization, OPA and FMOC-Cl, which was performed on-line with the use of an injection program. Eight different reverse phase columns, with different physico-chemical characteristics, were evaluated; the best results in terms of chromatographic run time and separation were achieved with the Gemini® C18 (150 × 4.6; 5 µm). The proposed method has been shown to be sufficiently sensitive, selective, and precise, while permitting a wide operational range. Several commercial bee pollen samples were analyzed with the proposed methodology, and significant differences in free amino acid content were observed depending on the plant origin (19–192 mg/g of total content), PRO always being detected at the highest concentrations (> 3 mg/g). It can be concluded that determining free amino acid content in bee pollen could be useful not only for assessing the quality and nutritional value of this product, but also for pinpointing the origin of the samples.
Data Availability
The datasets generated during the current study are included in this published article and the Supplementary Information, or they are available from the corresponding author on reasonable request.
References
Ares AM, Valverde S, Bernal JL, Nozal MJ, Bernal J (2018) Extraction and determination of bioactive compounds from bee pollen. J Pharm Biomed Anal 147:110–124. https://doi.org/10.1016/j.jpba.2017.08.009
Biluca FC, Bernal J, Valverde S, Ares AM, Gonzaga LV, Costa ACO, Fett R (2019) Determination of free amino acids in stingless bee (Meliponinae) honey. Food Anal Methods 12:902–907. https://doi.org/10.1007/s12161-018-01427-x
Campos MG, Anjos O, Chica M, Campoy P, Nozkova J, Almaraz-Abarca N, Barreto LMRC, Nordi JC, Estevinho LM, Pascoal A, Paula VB, Chopina A, Dias LG, Tešić ZLJ, Mosić MD, Kostić AŽ, Pešić MB, Milojković-Opsenica DM, Sickel W, Ankenbrand MJ, Grimmer G, Steffan-Dewenter I, Keller A, Förster F, Tananaki CH, Liolios V, Kanelis D, Rodopoulou MA, Thrasyvoulou A, Paulo L, Kast C, Lucchetti MA, Glauser G, Lokutova O, de Almeida-Muradian LB, Szczęsna T, Carreck NL (2021) Standard methods for pollen research. J Apic Res 60:1–109. https://doi.org/10.1080/00218839.2021.1948240
Canale A, Benelli G, Castagna A, Sgherri C, Poli P, Serra A, Mele M, Ranieri A, Signorini F, Bientinesi M, Nicolella C (2016) Microwave-assisted drying for the conservation of honeybee pollen. Materials 9:363. https://doi.org/10.3390/ma9050363
Chantarudee A, Phuwapraisirisan P, Kimura K, Okuyama M, Mori H, Kimura A, Chanchao C (2012) Chemical constituents and free radical scavenging activity of corn pollen collected from Apismellifera hives compared to floral corn pollen at Nan. Thailand BMC Complement Altern Med 12:45. https://doi.org/10.1186/1472-6882-12-45
Gałuszka A, Migaszewski Z, Namieśnik J (2013) The 12 principles of green analytical chemistry and the SIGNIFICANCE mnemonic of green analytical practices. Trends Anal Chem 50:78–84. https://doi.org/10.1016/j.trac.2013.04.010
González-Paramás AM, Gómes-Bárez JA, Cordón-Marcos C, García-Villanova RJ, Sánchez-Sánchez J (2006) HPLC-fluorimetric method for analysis of amino acids in products of the hive (honey and bee-pollen). Food Chem 95:148–146. https://doi.org/10.1016/j.foodchem.2005.02.008
Kafantaris I, Amoutzias GD, Mossialos D (2021) Foodomics in bee product research: a systematic literature review. Eur Food Res Technol 247:309–331. https://doi.org/10.1007/s00217-020-03634-5
Li F, Guo S, Zhang S, Peng S, Cao W, Ho CT, Bai N (2019) Bioactive constituents of F esculentum bee pollen and quantitative analysis of samples collected from seven areas by HPLC. Molecules 24:2705. https://doi.org/10.3390/molecules24152705
Lilek N, Gonzales AP, Božič J, Borovšak AK, Bertoncelj J (2015) Chemical composition and content of free tryptophan in Slovenian bee pollen. J Food Nut Res 54:323–333. https://www.vup.sk/index.php?mainID=2&navID=36&version=2&volume=54&article=1988
Nikkeshi A, Kuramitsu K, Yokoi T, Yamaji K (2021) Simple methods of analyzing proteins and amino acids in small pollen samples. J Apic Res 61:107–113. https://doi.org/10.1080/00218839.2021.1915633
Perucho J, Gonzalo-Gobernado R, Bazan E, Casarejos MJ, Jiménez-Escrif A, Asensio MJ, Herranz AS (2015) Optimal excitation and emission wavelengths to analyze amino acids and optimize neurotransmitters quantification using precolumn OPA-derivatization by HPLC. Amino Acids 47:963–973. https://doi.org/10.1007/s00726-015-1925-1
Prđun S, Svečnjak L, Valentić M, Marijanović Z, Jerković I (2021) Characterization of bee pollen: physico-chemical properties, headspace composition and FTIR spectral profiles. Foods 10:2103. https://doi.org/10.3390/foods10092103
Stabler D, Power EF, Borland AM, Barnes JDM, Wright GA (2018) A method for analysing small samples of floral pollen for free and protein-bound amino acids. Methods Ecol Evol 9:430–438. https://doi.org/10.1111/2041-210X.12867
Taha EKA, Al-Kahtani S, Taha R (2019) Protein content and amino acids composition of bee-pollens from major floral sources in Al-Ahsa, eastern Saudi Arabia. Saudi J Biol Sci 26:232–237. https://doi.org/10.1016/j.sjbs.2017.06.003
Themelis T, Gotti R, Orlandini S, Gatti R (2019) Quantitative amino acids profile of monofloral bee pollens by microwave hydrolysis and fluorimetric high performance liquid chromatography. J Pharm Biomed Anal 173:144–153. https://doi.org/10.1016/j.jpba.2019.05.031
Omar WAW, Yahaya N, Ghaffar ZA, Fadzilah NH (2018) GC-MS analysis of chemical constituents in ethanolic bee pollen extracts from three species of Malaysian stingless bee. J Apic Sci 62:275–284. https://doi.org/10.2478/jas-2018-0022
Walker V, Mills GA (1995) Quantitative methods for amino acid analysis in biological fluids. Ann Clin Biochem 32:28–57. https://doi.org/10.1177/000456329503200103
Wu G, Wu Z, Dai Z, Yang Y, Wang W, Liu C, Wang B, Wang J, Yin Y (2013) Dietary requirements of ‘“nutritionally non-essential amino acids”’ by animals and humans. Amino Acids 44:1107–1113. https://doi.org/10.1007/s00726-012-1444-2
Yang K, Wu D, Ye X, Liu D, Chen J, Sun J (2013) Characterization of chemical composition of bee pollen in China. J Agric Food Chem 61:708–718. https://doi.org/10.1021/jf304056b
You J, Liu L, Zhao W, Zhao X, Suo Y, Wang H, Li Y (2007) Study of a new derivatizing reagent that improves the analysis of amino acids by HPLC with fluorescence detection: application to hydrolyzed rape bee pollen. Anal Bioanal Chem 387:2705–2718. https://doi.org/10.1007/s00216-007-1155-9
Zhang JZ, Xue XF, Zhou JH, Chen F, Wu LM, Li Y, Zhao J (2009) Determination of tryptophan in bee pollen and royal jelly by high-performance liquid chromatography with fluorescence detection. Biomed Chromatogr 23:994–998
Ziegler J, Abel S (2014) Analysis of amino acids by HPLC-electrospray negative ion tandem mass spectrometry using 9-fluorenylmethoxycarbonyl chloride (FMOC-Cl) derivatization. Amino Acids 46:2799–2808. https://doi.org/10.1007/s00726-014-1837-5
Acknowledgements
The authors wish also to thank David Rixham (White Rose English School, Valladolid, Spain) for performing the English revision, N. Agüero, I. Cívicos and A. Redondo for their technical support.
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature. This work was supported by the Spanish “Ministerio de Economía y Competitividad” and the “Instituto Nacional de Investigación y Tecnología Agraria y Alimentaria” (project number RTA 2015–00013-C03-03).
Author information
Authors and Affiliations
Contributions
José Bernal: Conceptualization, funding acquisition, investigation, project administration, resources; visualization, writing—original draft preparation, Writing—review & editing; María T. Martín: conceptualization, investigation, methodology, validation, visualization writing—original draft preparation; Laura Toribio: conceptualization, investigation, methodology, validation, visualization writing—original draft preparation; Ana M. Ares: conceptualization, investigation, methodology, project administration, supervision, validation, writing—original draft preparation, writing—review and editing.
Corresponding authors
Ethics declarations
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed Consent
Not applicable.
Conflict of Interest
José Bernal declares that he has no conflict of interest. María T. Martín declares that she has no conflict of interest. Laura Toribio declares that she has no conflict of interest. Ana M. Ares declares that she has no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ares, A.M., Martín, M.T., Toribio, L. et al. Determination of Free Amino Acids in Bee Pollen by Liquid Chromatography with Fluorescence Detection. Food Anal. Methods 15, 2172–2180 (2022). https://doi.org/10.1007/s12161-022-02281-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-022-02281-8