Abstract
Common wheat is one of the most important staple food crops worldwide. However, unlike other important staple crops such as maize or soybean, genetically modified (GM) wheat is not yet present in the global food market. Nonetheless, in the recent past, the adventitious presence of GM glyphosate-tolerant volunteers was reported in open wheat fields in the USA. The European Union Reference Laboratory for GM Food and Feed (EURL-GMFF) was therefore called to develop a strategy to detect such unauthorised GM wheat in wheat samples by using both taxon-specific and screening tests. Two candidate common wheat taxon-specific real-time PCR methods were suggested, one targeting ssII-D gene coding for starch synthase and the other targeting waxy-D1 gene, coding for granule-bound starch synthase. In the present study, the two above-mentioned real-time PCR taxon-specific methods were in-house verified and compared, proposing droplet digital PCR (ddPCR) as a new tool for supporting the application of the European Network of GMO Laboratories (ENGL) established method performance criteria. Preliminary performance data of waxy-D1 and ssII-D methods in ddPCR format are shown too to give a contribution to the bridging process from the consolidated to the emerging quantitative PCR methodology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Common wheat (Triticum aestivum) is one of the major food crops in the world. Belonging to the Triticum genus, it is one of the most widespread staple cereals in the food industry, and consists of several cold-resistant cultivars allowing its cultivation in frigid climate areas too. The genus Triticum includes very complex polyploid genomes, symbolised by the letters A, B, D and G. Common wheat (T. aestivum) is hexaploid AABBDD (1C value = 17.33 pg; Bennett and Smith 1976) and Triticum durum is tetraploid AABB.
Betting on its potential impact in the food sector, the public research biotech industry has been striving for decades to put genetically modified (GM) wheat on the market by trying to incorporate desired traits such as herbicide tolerance, disease resistance and improvements in the quality of this crop (Leckband and Lörz 1998; Anand et al. 2003; Hu et al. 2003; Abouseadaa et al. 2015; Shin et al. 2008). Within this line of research, in the 90s, Monsanto, one of the major agrobiotech companies, launched a programme for the development of genetically modified common wheat tolerant to glyphosate herbicide, and some promising GM “events” (such as MON71800) were authorised in the USA to be tested in field trials. However, after completion of the experimental phase, no GM wheat has ever been marketed. Nevertheless, in 2013, the presence in Oregon (USA) of unexpected glyphosate tolerant wheat was noticed and then confirmed and reported by the United States Department of Agriculture (USDA). This event was ascribed to MON71800 volunteers, most likely originating from the GM wheat field trials (United States Department of Agriculture (USDA) 2015). Moreover, in 2016, a similar situation was found in Washington state due to the adventitious presence of a MON71800 sister GM event named MON71700 (United States Department of Agriculture–USDA news 2016). As this GM crops are not authorised in the EU, in both cases, the US Animal and Plant Health Inspection Service (APHIS) informed the European Commission and the member states of the European Union of the event and the European Commission urged the member states to start testing consignments of common wheat at import from the USA for the presence of unauthorised GM wheat.
The authorisation process for introducing GM products in the EU market is ruled by Directive 2001/18/EC (European Parliament and Council of the European Union 2001), for the deliberate release into the environment of genetically modified organisms, and by Regulation (EC) 1829/2003 (European Parliament and Council of the European Union 2003) for GM food and feed. According to this legislation, in addition to the data concerning risk assessment, the applicant must provide a method for the detection and quantification of the transformation event under authorisation. The method has to be validated by the European Union Reference Laboratory for GM Food and Feed (EURL-GMFF), established at the Joint Research Centre and assisted by the European Network of GMO Laboratories (ENGL), according to the document “Definition of minimum performance requirements for analytical methods of GMO testing-MPR” (European Network of GMO Laboratories (ENGL) 2015).
Quantitative methods are required for enforcing EU labelling and traceability rules for food/feed containing authorised GM material >0.9% with respect to the specific ingredient. The quantitative analysis consists of a relative quantification of the GMO event to the plant species itself (taxon reference system). In this context, the selection of the species-specific reference system is crucial for the reliability of the quantitative analytical results obtained.
Real-time PCR represents the gold standard for GMO analysis, and the MPR guidelines refer exclusively to this technique. Nevertheless, digital PCR (dPCR) represents one of the more recent and promising quantitative PCR techniques. In dPCR, the sample is distributed across a large number of partitions containing zero, one, or more copies of target DNA. The absolute number of target DNA is then calculated by taking into consideration the ratio of positive (or negative) partitions to total partitions and using Poisson distribution model. According to this model, λ is the average number of target copies per partition, and it is determined by using λ = −ln(1 − k/n), where n is the total number of partitions detected and k is the number of positive partitions detected (Pinheiro et al. 2012).
Digital PCR platforms currently available are essentially classified by the way partitioning of reactions is obtained, either making use of microfluidic chambers or running individual reactions in droplets generated in a water-oil emulsion. The number of references to dPCR and its possible application in the field of GMO quantification have dramatically increased in recent years (Burns et al. 2010; Corbisier et al. 2010; Morisset et al. 2013; Demeke T. et al. 2014; Dobnik et al. 2015; Koppel and Bucher 2015; Dobnik et al. 2016; Félix-Urquídez et al. 2015; Gerdes et al. 2016; Zhu et al. 2016).
In the lack of an event-specific method for the identification of the above-mentioned unauthorised GM wheat events, the European Union Reference Laboratory for GM Food and Feed (EURL-GMFF) developed a strategy to detect GM wheat in wheat samples by using both taxon-specific and screening tests. This strategy was then described in a guidance document published on the EURL-GMFF website (European Union Reference Laboratory for GM Food and Feed 2013a), together with a comprehensive review and related bioinformatics analysis on all available relevant literature on taxon-specific real-time PCR systems for the identification of T. aestivum (European Union Reference Laboratory for GM Food and Feed 2013b). The results of the study suggested that the method described by Matsuoka et al. (2012), targeting ssII-D gene coding for starch synthase, and the one described by Iida et al. (2005), targeting waxy-D1 gene, coding for granule-bound starch synthase, may represent good candidates to specifically identify common wheat even in complex food samples. With regard to the latter method, what was reported in the above-mentioned EURL-GMFF guidelines had to be carefully considered, i.e. “according to in silico analyses conducted by the EURL GMFF, the primer sequences described in Iida et al. 2005 are not correct. The primers (Wx012F/Wx012R) and probe (Wx-Taq 1) sequences reported by the same authors in the following article: “Imai et al. (2012), Food Hyg. Saf. Sci. 204 Vol. 53 203–210 and the patent EP2180051” are recommended as reference.
In 2016, EURL-GMFF verified an endpoint real-time PCR method submitted by Monsanto Company, for detecting event MON71800 in wheat, but the wheat-specific system of choice was designed on T. aestivum acetyl-coenzyme A carboxylase, Acc-1, reference target (Genebank AF029897.1), although it is unable to discriminate between T. aestivum and T. durum (European Union Reference Laboratory for GM Food and Feed 2016). Given the importance of T. durum in food matrices, such an evident lack of specificity was considered to be a major drawback, and therefore, Acc-1 targeting method was not deemed appropriate for the aim of our study. The two above-mentioned methods (waxy-D1 and ssII-D), which had been previously recommended by EURL-GMFF, were then analysed and compared to select a unique taxon reference system for the identification and the quantification of wheat.
In order to evaluate possible within-species copy number variation of the two above-mentioned target sequences (waxy-D1 and ssII-D), a representative number of wheat cultivar samples were analysed in parallel using both real-time PCR and digital droplet PCR platform. The two taxon-specific systems, waxy-D1 and ssII-D, were compared for several validation parameters in accordance with “Definition of Minimum Performance Requirements for Analytical Methods of GMO Testing” document (European Network of GMO Laboratories (ENGL) 2015).
Materials and Methods
Plant Materials
Seeds from the following 16 wheat cultivars, provided by Consiglio per la Ricerca in agricoltura e l’analisi dell’Economia Agraria (CREA) seed testing laboratory, were used in the study: (A) Aquilante, (B) Palesio, (C) Bologna, (D) Blasco, (E) Aubusson, (F) Antille, (G) Bandiera, (H) Alatamira, (I) Pandas, (L) Ora, (M) Albachiara, (N) Laria, (O) Mieti, (P) Bisquit, (Q) Sagittario and (R) Bramante (Table S1 in supplementary material).
Sample Preparation and DNA Extraction
Seed samples were milled (GM200 RETSCH, Haan, Germany) and the ground material was used for DNA extraction. Total genomic DNA was extracted using the ION FORCE FAST DNA Extraction Kit for processed food and feed complex material (GENERON, Modena, Italy) from 2 g of each cultivar, according to the manufacturer’s instructions. DNA concentration was calculated from the absorption at 260 nm measured with a UV spectrophotometer (Biophotometer, Eppendorf, Hamburg, Germany). The DNA purity was checked by both calculating the ratio of the absorption values at 260/280 nm and carrying out a real-time PCR (inhibition run) targeting waxy-D1 encoding gene on two different amounts of each extracted DNA (300 and 75 ng, i.e. 1:4 dilution). As the theoretical ΔCq value is 2 for 1:4 dilution, the following acceptance criterion was considered: 1.5 < ΔCq < 2.5.
Oligonucleotide Primers and Probes
The oligonucleotide primers and TaqMan probes used in this study, both for real-time PCR and ddPCR, are listed in Table 1. The TaqMan probes were labelled on the 5′ end with (FAM) reporter dye and 3′ end (BHQ) quencher dye. All of the primers and probes were synthesised by Eurofins Genomics (Ebersberg, Germany).
Real-Time PCR
Real-time PCR reactions were performed in optical 96-well PCR plates and run on the ABI PRISM SEQUENCE DETECTION SYSTEM 7900HT (Thermo Fisher Scientific Waltham, MA USA), in a final volume of 25 μL. Real-time PCR mixture for both primers and probe set (waxy-D1 and ssIID) contained the following amounts: 200 ng DNA, 0.5 μM primers pairs, 0.2 μM probe and 12.5 μL Universal Master Mix (Thermo Fisher Scientific Waltham, MA USA). Real-time PCR runs had the following thermal profile: 2′ 50 °C, 10′ 95 °C; [15″ 95 °C, 60 °C 60″] × 50 cycles. Amplification plots were visualised in the logarithmic graph for manual setting of the fluorescent threshold value. This threshold was chosen in the middle of the linear phase of the PCR plots; the baseline was evaluated in the linear graph by manual setting too. After manual adjustment of these parameters, a regression curve for each DNA dilution set was obtained by plotting all of the Cq values, from each concentration level, against the log of the DNA copy number, obtained by ddPCR (see protocol below), at the corresponding concentration level. The regression curve Cq = a log(copy number) + b was used to calculate the experimental copy number corresponding to each specific Cq value.
Droplet Digital PCR
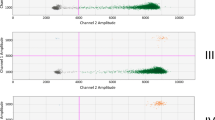
ddPCR reaction mixture contained 4 μL of DNA template (20 μL total reaction volume); final primers and probe concentrations in ddPCR mixes were identical to the real-time PCR conditions used in this study. Droplets were generated in 8-well cartridges, using the QX100 droplet generator (Bio-Rad, Pleasanton, CA). Water-in-oil emulsions were transferred to a 96-well plate and amplified in a PCR thermocycler GeneAmp PCR System 9700 (Thermo Fisher Scientific, Waltham, MA, USA) with the following thermal profile: 10 min at 95 °C, followed by 45 cycles of two steps of 30 s at 94 °C and 60 s annealing/extension at 60% ramp rate 60 °C; after amplification, products were denatured at 98 °C for 10 min, then cooled to 4 °C. Plates were then transferred to the QX100 droplet reader (Bio-Rad, Pleasanton, CA). The threshold was set manually, using the fluorescence amplitude versus event number (1D amplitude). Data generated by the QX100 droplet reader were analysed by Quantasoft 1.7.4.0917 software rejecting PCR reactions (single wells) with a too low number of droplets (<8000).
With regard to ddPCR-specific quality criteria (i.e. amount of rain and peak resolution R s ), the calculation procedures indicated in Lievens et al. (2016) were adopted.
Experimental Design for the Real-Time PCR Verification and Comparison Study
Evaluation of Within-Taxon Allelic Variation of the Target Sequences
For each cultivar, a DNA dilution set was prepared (D1 to D6) from 160 to 0.13 ng (see Table S2 in supplementary material). Each dilution set was tested on the same plate with waxy-D1 and ssII-D systems, both in real-time PCR and ddPCR (four replicates for D1 to D4 and eight replicates for D5 and D6). A total of 16 real-time PCR runs and 16 ddPCR runs were carried out and analysed. Cq values attained by real-time PCR were plotted against the log of the copy number estimated by ddPCR (for each corresponding concentration level) to create 16 regression curves for waxy-D1 system and 16 regression curves for ssII-D system.
Precision—Relative Repeatability Standard Deviation
In order to evaluate precision of waxy-D1 and ssII-D real-time PCR methods, five DNA dilutions (1:4) of two varieties, Aquilante (A) and Palesio (B), from S1 (300 ng) to S5 (1.17 ng) were performed and then tested in ddPCR for both endogenous systems.
Two different real-time PCR plates were run, one for waxy-D1 and one for ssII-D (Table S3 in supplementary material). In each plate, cultivar A was used as calibrator. Dilutions from S1 to S5 (three replicates each) were loaded three times in order to have three calibration curves in the same plate. Cq values, from each concentration level, were plotted against the corresponding log copy number, as estimated by ddPCR. An identical dilution set from cultivar B was loaded three times on the same plate as unknown samples. For each concentration level, 27 data points (9 by each dilution set) were generated to calculate percent relative standard deviation of repeatability (RSDr%).
Limit of Detection and Limit of Quantification
A subset of DNA dilution sets (D1 to D6) prepared as described above from 5 out of 16 cultivars (Aquilante, Palesio, Bologna, Blasco, Albachiara) were exploited in order to obtain 20 data points for D1–D4 and 40 data points for D5 and D6.
For improving the limit of quantification (LOQ) resolution, a second test was carried out on a new dilution set (i.e. 1400, 700, 350, 175, 90, 45, 23 and 11 theoretical copy numbers) from Aquilante cultivar DNA. For each level, 20 data points were obtained.
Robustness
In order to check the capacity of the methods to remain sufficiently unaffected by deliberate deviations, they were tested at different concentrations of master mix components (less/more 30% primers and probes and less/more master mix by addition or subtraction of 1 μL), as well as with a different real-time PCR instrument (FAST REAL TIME PCR SYSTEM 7900HT, Thermo Fisher Scientific, Waltham, MA, USA). Robustness was tested on 50 copies of the target (waxy-D1 and ssII-D) in six replicates at each condition.
Sequence Analysis of the waxy-D1 and ssII-D Target Sequences
Target sequences of both waxy-D1 and ssII-D PCR systems were amplified for all 16 wheat varieties, by using external primer pairs, conveniently designed (Primer-BLAST) on the sequences referred to on the respective original papers, to be specific for the corresponding flanking sequences from all wheat varieties. The amplicons obtained for both waxy-D1 and ssII-D from all 16 wheat varieties were sequenced by ABI Prism 3500 genetic analyser (Thermo Fisher Scientific, Waltham, MA, USA). All the sequences were then aligned by using Sequencing Analysis Software (Thermo Fisher Scientific, Waltham, MA, USA). Primers used in this study for sequence analysis (see Table S4 in supplementary material) have been selected on the corresponding reference sequences by using Primer-BLAST online resource (https://www.ncbi.nlm.nih.gov/tools/primer-blast/).
Results and Discussion
Selection of the Two Taxon-Specific Real-Time PCR Methods and Alignment of the Thermal Profiles
After having studied and considered all the relevant literature, with the fundamental contribution of a specific EURL-GMFF survey (European Union Reference Laboratory for GM Food and Feed 2013b), all the methods not strictly specific for T. aestivum were a priori left out of our study. Actually, according to the above-mentioned EURL-GMFF survey, the only two methods fulfilling this requirement were waxy-D1 (Imai et al. 2012) and ssII-D (Matsuoka et al. 2012), both designed on subgenome D. The only cross-reaction reported for these two methods is with the ancestor Aegilops tauschii, carrying the subgenome D, which is very unlikely to be found in food products. On the contrary, the other methods, designed on subgenomes A or B, show important cross-reactions with other related cereals, such as T. durum, which are easily found in food products. This was a significant reason for not selecting them. According to the experimental design adopted in this study, it was planned to carry out all the comparisons by always running the two reference real-time PCR methods in the same plate in order to exclude run-to-run variability. Therefore, the original thermal cycling conditions of the two real-time methods were harmonised as previously described. Once the annealing/extension temperature was increased to 60 °C, the next step was to look at the current version of the MPR guidelines for identifying the parameters already covered by the original studies and those to be included in our intralaboratory verification study. The following parameters were then not taken into consideration:
-
Specificity—“property of a method to respond exclusively to the characteristic or analyte of interest”. The annealing/extension temperature at 60 °C was in both cases increased by comparison with the original settings. The specificity of both PCR assays was then expected to be increased and, in any case, not decreased by this shift. We looked at the rather extensive specificity tests conducted in the two original papers as well as at the independent and exhaustive EURL-GMFF survey before deciding that such a parameter was already covered by the existing literature. So we just checked some GM CRMs (ERM and AOCS) available in our laboratory for routine tests (data not shown), but the specificity test was considered redundant;
-
Trueness—“the closeness of agreement between the average value obtained from a large series of test results and an accepted reference value”. Trueness was not assessed due to unavailability of wheat CRM in copy number and to the lack of clarity r in the scientific community regarding the use of dPCR data as “true value” for this purpose. In absence of an “accepted reference value”, this parameter was not included in the study;
All the other MPR guideline performance parameters were evaluated in this study.
Use of ddPCR Versions of waxy-D1 and ssII-D Real-Time PCR Methods in This Study and Some Performance Data
In this study, digital PCR was mainly used for supporting the evaluations and comparisons on the corresponding real-time PCR systems. Droplet digital PCR is a fairly new PCR format, and its performance criteria are still being defined and harmonised. With this regard, a recent paper (Lievens et al. 2016) has shed new light on ddPCR performance parameters, paying special attention to the GMO context. It recognises some performance parameters adopted for qPCR, such as applicability, practicability, specificity, dynamic range and trueness, to be directly transferable to the digital PCR. Nevertheless, some new ddPCR-specific criteria are suggested to ensure robust classification of positive and negative droplets, as this aspect shows to be of crucial importance for obtaining an accurate and reliable quantification of target copy number in ddPCR. In particular, it is recommended to check that (i) a single amplification product is present, (ii) the amount of rain (droplets having an intermediate fluorescence level between negative and positive populations) is lower than 2.5% and (iii) the peak resolution (R s ) is at least 2.5. According to the authors, the optimal conditions for evaluating such parameters could be achieved with control samples having a λ of approximately 0.7 as at this concentration, approximately half the partitions are positive and half are negative for the target. In the present study, which follows this suggestion as much as possible, the two droplet digital PCR methods (waxy-D1 and ssII-D) have been evaluated for such specific performance criteria. Indeed, the percentage of rain and the peak resolution were calculated at approximately 160 ng of wheat DNA, corresponding to the higher concentration level tested in this study, with a λ value ranging from 0.3 to 0.4. Both methods proved to be compliant with the proposed requirements, with a percentage of rain in the range of 0.403–1.154% for waxy-D1 and in the range of 0.651–1.427% for ssII-D and R s being between 9.21 and 10.10 for waxy-D1 and between 8.11 and 9.95 for ssII-D (see Fig. S1 and Table S5 in supplementary material).
Evaluation of Within-Taxon Allelic Variation of the Target Sequences
In the field of GMO analysis, the adoption of a reference gene target with a stable, known and low copy number, preferably 1 copy per haploid genome equivalent (HGE), and with no allelic variation amongst the cultivars of the same species, is recommended.
In order to evaluate the aforementioned requirements, 16 common wheat cultivars, acquired with the help of CREA (Italian seed testing laboratory and project partner) as varieties cultivated in Italy, were tested with both waxy-D1 and ssII-D endogenous reference PCR systems.
DNA extracted from each of the 16 cultivars was tested at 6 dilutions for both waxy-D1 and ssII-D methods by running real-time PCR and droplet digital PCR in parallel within the same working day. In Figs. 1 and 2, results obtained by ddPCR are displayed. In Fig. 1, copy number/microliter of one cultivar (Bologna cultivar) obtained on the dilution set D1–D6 (see the “Materials and Methods” section) is shown. Figure 2 shows log of waxy-D1 copy number plotted against log of ssII-D copy number from dilution D1 to D6, for all the cultivars tested. According to this plot, all data points referred to cultivars having the same copy number of waxy-D1 and ssII-D were expected to lay on the bisector of the Cartesian coordinate system, whilst cultivars having any copy number variation, for waxy-D1 or ssII-D, would have shown some deviation. The results confirm an absolute copy number ratio waxy-D1/ssII-D around 1 for all cultivars tested.
Furthermore, in order to verify the absence of single-nucleotide polymorphisms (SNPs) in the PCR target sequences, the amplicons generated for each cultivar by both waxy-D1 and ssII-D PCR systems were sequenced. The analysed DNA sequences were always identical to the corresponding reference sequences all across the cultivars tested (data not shown).
In the context of GMO quantification, the taxon-specific PCR system should target a single DNA copy per allele within the taxon of interest. The absence of copy number variation within the taxon is generally verified on a representative collection of cultivars, by using real-time PCR (Rønning et al. 2006; Vautrin and Zhang 2007; Chaouachi et al. 2008; Jacchia et al. 2015). The range of variability of Cq values in amplification should not exceed 1 Cq within the taxon (European Network of GMO Laboratories (ENGL) 2015). Moreover, in order to establish gene copy number, Southern blot is considered the most reliable technique, although the process is labour intensive and not automatable. A recent paper (Głowacka et al. 2016) compared Southern blot and PCR-based methods (qPCR and droplet digital PCR) for their ability to determine T-DNA copy number and homozygosity in transgenic plants. Setting Southern blotting as the gold standard, data obtained by ddPCR resulted more accurate than those generated by qPCR. On the basis of these findings, ddPCR technique seems to be, at present, the most convenient tool for routinely assess T-DNA copy number and homozygosity. The validity of ddPCR technique for the determination of zygosity in transgenic plants is also confirmed by Xu et al. (2016). Therefore, the experimental design of the present study is mainly based on ddPCR-generated data set.
Performance Data of waxy-D1 and ssII-D Real-Time PCR Methods in Comparison
Amplification Efficiency, R 2 and Dynamic Range
The efficiency of PCR can be expressed as the percentage of target molecules that are duplicated in one PCR cycle. PCR efficiency is mathematically correlated to the slope of a linear regression (plot of the Cq versus the logarithm of the target concentrations) according to the following equation: E = 10−1/slope − 1, where E is the efficiency; 100% efficiency corresponds to a −3.32 slope value. The R 2 (coefficient of determination) gives a measure of how well a linear regression fits the experimental data points used to obtain it. For both PCR methods, 16 regression curves (one per cultivar) were obtained by plotting Cq values, at each concentration level, against the log of target DNA copy number as calculated by ddPCR (see the “Materials and Methods” section). The slope and R 2 coefficient were determined for each regression curve (Table 2). The average slope is comparable for waxy-D1 and ssII-D methods, and in line with MPR guideline requirements. In terms of linearity, the two methods can be considered equivalent too. Dynamic range is defined as “the range of concentrations over which the module performs in a linear manner with an acceptable level of trueness and precision”. Considering that trueness was not measurable, the dynamic range was assessed relying upon precision data and LOQ evaluation. The two methods performed in a linear manner between 12,796 copies and 23 copies (waxy-D1), and between 15,348 copies and 23 copies (ssII-D) (see Tables S7 and S9 in supplementary material).
Precision—Relative Repeatability Standard Deviation
Precision is defined as the relative standard deviation of test results obtained under repeatability conditions. Five dilutions (1:4) of the two cultivars Aquilante (A) and Palesio (B) were set and analysed as described above, with the lower concentration level (S5) slightly above the limit of quantification of the two PCR methods. Cultivar A (Aquilante) was used as calibrator and cultivar B (Palesio) as unknown sample. According to the MPR guidelines, RSDr% should be ≤25% over the whole dynamic range of the PCR modules. In our results, RSDr% ranged between 3.75 and 20.98 for ssII-D, and between 5.68 and 15.56 for waxy-D1 (Tables S8 and S10 in supplementary material).
Figure 3 (box plot) displays log copy number per concentration level from S1 to S5 for waxy-D1 and ssII-D methods in parallel. In terms of repeatability, the two methods are comparable and in line with MPR guidelines, even including box plot outliers in the distributions.
Box plot analysis (R version 3.2.5) of repeatability (precision analysis) for waxy-D1 and ssII-D real-time PCR; copy number of each dilution (S1–S5) was obtained from three different calibration curves (see the “Materials and Methods” section and supplementary materials)
Moreover, waxy-D1 and ssII-D real-time PCR methods and the corresponding waxy-D1 and ssII-D ddPCR versions were compared. Table 3 shows, for both systems, the ability of the two techniques to quantify the analytical target in the S1–S5 range of dilution (see also Table S6 in supplementary material).
Limit of Detection and LOQ
Absolute limit of detection and absolute limit of quantification of the two real-time PCR methods, waxy-D1 and ssII-D, were estimated on DNA extracted from five cultivars (Aquilante, Palesio, Bologna, Blasco, Albachiara). In order to match each specific Cq value to the corresponding absolute target copy number estimated in ddPCR, data obtained by the evaluation of within-taxon allelic variation of the target sequences were exploited (Tables S11 and S12 in supplementary material). In both cases, limit of detection (LOD) is below 7 copies. Limit of quantification was set to be below the lowest tested amount showing a RSDr% ≤25%. In both cases, the LOQ is below 63 copies. This value is slightly outside the MPR guideline acceptance criteria (i.e. 50 copies), so it was necessary to refine our results by carrying out an additional test for improving the LOQ resolution below 62 copies. A new dilution set (i.e. 1400, 700, 350, 175, 90, 45, 23 and 11 theoretical copy numbers) from Aquilante cultivar DNA was tested in 20 replicates per level. The actual LOQ was then set at 23 theoretical copy number for both methods (see Tables S13, S14, S15 and S16 in supplementary material).
On the basis of our data obtained, both taxon-specific real-time PCR methods show to be promising for being adopted in quantitative GMO analysis. With regard to the choice, as already stated in a previous study (Paternò et al. 2009), from both a theoretical and technical point of view, there is no reason for changing the PCR reference system according to the GM event to be quantified (except for very peculiar situations related to food matrix and amplicon length). Therefore, the use of a single taxon-specific PCR reference system for enforcement purposes is not only advisable for practical reasons but also sensible from a scientific perspective. Nevertheless, the present study could not find any element to prefer ssII-D to waxy-D1 or vice versa, even though waxy-D1 is more supported by previous literature (Huang et al. 2013).
Robustness
The robustness of a method is defined as the “measure of its capacity to remain unaffected by small but deliberate deviations from the experimental conditions described in the procedure”. Currently, MPR guidelines provide some indications on how to evaluate robustness of qualitative and quantitative methods. Nevertheless, no specific suggestions are provided for assessing a taxon reference module to be used in a quantitative method. In our study, standard conditions were compared to deliberately altered conditions by testing ssII-D to waxy-D1 at 50 target copies (higher acceptance limit for LOQ according to the MPR guidelines) and reporting ΔCq values. ΔCq is the difference between the Cq mean obtained in standard condition and the Cq means obtained by each deviating condition. The acceptance criterion adopted was ΔCq ≤ 1. Deliberate changes to the initial protocol did not affect waxy-D1 and ssII-D targets; all the differences tested were ≤0.5 Cq (Tables S17 and S18 in supplementary material).
Conclusions
In the present study, two real-time PCR wheat taxon-specific methods were selected from the relevant literature available as potential candidates for playing the role of a taxon-specific reference system for GM wheat quantification. The first aim of our work was to provide in-house validation data according to MPR and the comparison of the two best-performing reference systems so far available in literature for GM wheat quantification. We hereby also propose the use of digital PCR (and its absolute quantification power) in support of the verification and the validation of taxon-specific reference systems, where intervarietal stability of the target sequence is a priority. Previous performance data were evaluated and, where necessary, integrated according to the MPR guideline requirements for single laboratory validation. The alignment of test conditions permitted the direct comparison of the two candidate methods in the same testing sessions. According to the data obtained, both methods have demonstrated to be in line with the MPR acceptance criteria. In order to support the evaluations and comparisons of these real-time PCR systems, the corresponding droplet digital PCR versions were used, therefore contributing to the bridging process from a consolidated to an emerging quantitative PCR methodology. The results suggest that the two real-time PCR methods are comparable and both eligible as wheat taxon-specific reference systems in GMO quantification assays.
With a view to having additional confirmation of this potential, it will be very interesting to test both methods in combination with a GM-specific ddPCR module (e.g. MON71800-specific or MON71700-specific) as soon as the corresponding CRMs are available.
References
Abouseadaa HH, Osman GH, Ramadan AM, Hassanein SE, Abdelsattar MT, Morsy YB, Alameldin HF, El-Ghareeb DK, Nour-Eldin HA, Salem R, Gad AA, Elkhodary SE, Shehata MM, Mahfouz HM, Eissa HF, Bahieldin A. (2015) Development of transgenic wheat (Triticum aestivum L.) expressing avidin gene conferring resistance to stored product insects. BMC Plant Biol. 22;15:183. doi: https://doi.org/10.1186/s12870-015-0570-x
Anand A, Zhou T, Trick HN, Gill BS, Bockus WW, Muthukrishnan S (2003) Greenhouse and field testing of transgenic wheat plants stably expressing genes for thaumatin-like protein, chitinase and glucanase against Fusarium graminearum. J Exp Bot 54(384):1101–1111. https://doi.org/10.1093/jxb/erg110
Bennett MD, Smith JB. (1976) Nuclear DNA amounts in angiosperms. Philosophical Transactions of the Royal Society of London Series B-Biological Sciences 274: 227–274. Kew Royal Botanic Garden. http://data.kew.org/cvalues/
Burns MJ, Burrell AM, Foy CA (2010) The applicability of digital PCR for the assessment of detection limits in GMO analysis. Eur Food Res Technol 231(3):353–362. https://doi.org/10.1007/s00217-010-1277-8
Chaouachi M, El Malki R, Berard A, Romaniuk M, Laval V, Brunel D, Bertheau Y (2008) Development of a real-time PCR method for the differential detection and quantification of four Solanaceae in GMO analysis: potato (Solanum tuberosum), tomato (Solanum lycopersicum), eggplant (Solanum melongena), and pepper (Capsicum annum). J Agric Food Chem 56(6):1818–1828. https://doi.org/10.1021/jf073313n
Corbisier P, Bhat S, Partis L, Xie VR, Emslie KR (2010) Absolute quantification of genetically modified MON810 maize (Zea mays L.) by digital polymerase chain reaction. Anal Bioanal Chem 396(6):2143–2150. https://doi.org/10.1007/s00216-009-3200-3
Demeke T, Gräfenhan T, Holigroski M, Fernando U, Bamforth J, Lee SJ (2014) Assessment of droplet digital PCR for absolute quantification of genetically engineered OXY235 canola and DP305423 soybean samples. Food Control 46:470–474. https://doi.org/10.1016/j.foodcont.2014.06.018
Dobnik D, Spilsberg B, Bogožalec Košir A, Holst-Jensen A, Žel J (2015) Multiplex quantification of 12 European Union authorised genetically modified maize lines with droplet digital polymerase chain reaction. Anal Chem 87(16):8218–8226. https://doi.org/10.1021/acs.analchem.5b01208
Dobnik D, Štebih D, Blejec A, Morisset D, Žel J (2016) Multiplex quantification of four DNA targets in one reaction with Bio-Rad droplet digital PCR system for GMO detection. Sci Rep 6(1):35451. https://doi.org/10.1038/srep35451
European Network of GMO Laboratories-ENGL (2015). Definition of minimum performance requirements for analytical methods of GMO testing. Guidance document http://gmo-crl.jrc.ec.europa.eu/guidancedocs.htm
European Parliament and Council of the European Union (2001) Directive 2001/18/EC of the European Parliament and of the Council of 12 March 2001 on the deliberate release into the environment of genetically modified organisms and repealing Council Directive 90/220/EEC. Official Journal of European Communities. 1742001 L106(/1)
European Parliament and Council of the European Union (2003). Regulation (EC) no 1829/2003 of the European Parliament and of the Council of 22 September 2003 on genetically modified food and feed
European Union Reference Laboratory for GM Food and Feed (2013a). EU-RL GMFF guidance on testing for GM glyphosate-resistant wheat (MON71800) in wheat grain or in food/feed products containing wheat flour originating or consigned from the US. http://gmo-crl.jrc.ec.europa.eu/doc/Wheat_EURL_testingstrategy_V2.pdf
European Union Reference Laboratory for GM Food and Feed (2013b). Literature and bioinformatics analysis of wheat-specific detection methods. http://publications.jrc.ec.europa.eu/repository /bitstream/JRC86656/wheat_bioinformatics%20technical%20report_with%20registration.pdf
European Union Reference Laboratory for GM Food and Feed (2016) - Report on the verification of the performance of a simplex endpoint event-specific method for the detection of event MON71800 in wheat using real-time PCR
Félix-Urquídez D, Pérez-Urquiza M, Valdez Torres JB, León-Félix J, García-Estrada R, Acatzi-Silva A (2015) Development, optimization, and evaluation of a duplex droplet digital PCR assay to quantify the T-nos/hmg copy number ratio in genetically modified maize. Anal Chem 88(1):812–819. https://doi.org/10.1021/acs.analchem.5b03238
Gerdes L, Iwobi A, Busch U, Pecoraro S (2016) Optimization of digital droplet polymerase chain reaction for quantification of genetically modified organisms. Biomol Detect Quantif 7:9–20. https://doi.org/10.1016/j.bdq.2015.12.003
Głowacka K, Kromdijk J, Leonelli L, Niyogi KK, Clemente TE, Long SP (2016) An evaluation of new and established methods to determine T-DNA copy number and homozygosity in transgenic plants. Plant Cell Environ 39(4):908–917. https://doi.org/10.1111/pce.12693
Hu T, Metz S, Chay C, Zhou HP, Biest N, Chen G, Cheng M, Feng X, Radionenko M, Lu F (2003) Fry J. Agrobacterium-mediated large-scale transformation of wheat (Triticum aestivum L.) using glyphosate selection. Plant Cell Rep 21(10):1010–1019. https://doi.org/10.1007/s00299-003-0617-6
Huang H, Cheng F, Wang R, Zhang D, Yang L (2013) Evaluation of four endogenous reference genes and their real-time PCR assays for common wheat quantification in GMOs detection. PLoS One 8(9):e75850. https://doi.org/10.1371/journal.pone.0075850
Iida M, Yamashiro S, Yamakawa H, Hayakawa K, Kuribara H, Kodama T, Furui S, Akiyama H, Maitani T, Hino A (2005) Development of taxon-specific sequences of common wheat for the detection of genetically modified wheat. J Agric Food Chem 53(16):6294–6300. https://doi.org/10.1021/jf0505731
Imai S, Tanaka K, Nishitsuji Y, Kikuchi Y, Matsuoka Y, Arami S, Sato M, Haraguchi H, Kurimoto Y, Mano J, Furui S, Kitta K. (2012) An endogenous reference gene of common and durum wheat for detection of genetically modified wheat. Food Hyg Saf Sci. 204 Vol. 53 203–210
Jacchia S, Nardini E, Savini C, Petrillo M, Angers-Loustau A, Shim JH, Trijatmiko K, Kreysa J, Mazzara M (2015) Development, optimization, and single laboratory validation of an event-specific real-time PCR method for the detection and quantification of Golden Rice 2 using a novel taxon-specific assay. J Agric Food Chem 63(6):1711–1721. https://doi.org/10.1021/jf505516y
Koppel R, Bucher T (2015) Rapid establishment of droplet digital PCR for quantitative GMO analysis. Eur Food Res Technol 241(3):427–439. https://doi.org/10.1007/s00217-015-2475-1
Leckband G, Lörz H (1998) Transformation and expression of a stilbene synthase gene of Vitis vinifera L. in barley and wheat for increased fungal resistance. Theor Appl Genet 96(8):1004–1012. https://doi.org/10.1007/s001220050832
Lievens A, Jacchia S, Kagkli D, Savini C, Querci M (2016) Measuring digital PCR quality: performance parameters and their optimization. PLoS One 11(5):e0153317. https://doi.org/10.1371/journal.pone.0153317
Matsuoka Y, Arami S, Sato M, Haraguchi H, Kurimoto Y, Imai S, Tanaka K, Mano J, Furui S, Kitta K (2012) Development of methods to distinguish between durum/common wheat and common wheat in blended flour using PCR. Food Hyg Saf Sci 53(5):195–202
Morisset D, Štebih D, Milavec M, Gruden K, Žel J (2013) Quantitative analysis of food and feed samples with droplet digital PCR. PLoS One 8(5):e62583. https://doi.org/10.1007/s00216-009-3200-3 2143, 2150
Paternò A, Marchesi U, Gatto F, Verginelli D, Quarchioni C, Fusco C, Zepparoni A, Amaddeo D, Ciabatti I (2009) Finding the joker among the maize endogenous reference genes for genetically modified organism (GMO) detection. J Agric Food Chem 57(23):11086–11091. https://doi.org/10.1021/jf902560x
Pinheiro LB, Coleman VA, Hindson CM, Herrmann J, Hindson BJ, Bhat S, Emslie KR (2012) Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Anal Chem 84(2):1003–1011. https://doi.org/10.1021/ac202578x
Rønning SB, Berdal KG, Andersen CB, Holst-Jensen A (2006) Novel reference gene, PKABA1, used in a duplex real-time polymerase chain reaction for detection and quantitation of wheat- and barley-derived DNA. J Agric Food Chem 54(3):682–687. https://doi.org/10.1021/jf052328n
Shin S, Mackintosh CA, Lewis J, Heinen SJ, Radmer L, Dill-Macky R, Baldridge GD, Zeyen RJ, Muehlbauer GJ (2008) Transgenic wheat expressing a barley class II chitinase gene has enhanced resistance against Fusarium graminearum. J Exp Bot 59(9):2371–2378. https://doi.org/10.1093/jxb/ern103
United States Department of Agriculture (USDA). (2015). https://www.aphis.usda.gov/brs/aphisdocs/mt140008_froi_red_20160801.pdf
United States Department of Agriculture–USDA news. (2016) https://www.aphis.usda.gov/aphis/ourfocus/biotechnology/brs-news-and-information/ge+wheat+washington+state
Vautrin S, Zhang D (2007) Real-time polymerase chain reaction assay for endogenous reference gene for specific detection and quantification of common wheat-derived DNA (Triticum aestivum L.) J AOAC Int 90(3):794–801
Xu X, Peng C, Wang X, Chen X, Wang Q, Xu J (2016) Comparison of droplet digital PCR with quantitative real-time PCR for determination of zygosity in transgenic maize. Transgenic Res 25(6):855–864. https://doi.org/10.1007/s11248-016-9982-0
Zhu P, Fu W, Wang C, Du Z, Huang K, Zhu S, Xu W (2016) Development and application of absolute quantitative detection by duplex chamber-based digital PCR of genetically modified maize events without pretreatment steps. Anal Chim Acta 916:60–66. https://doi.org/10.1016/j.aca.2016.02.020
Acknowledgments
We are very grateful to our project partners Dr. Elena Perri and Dr. Daniela Villa from Consiglio per la Ricerca in agricoltura e l’analisi dell’Economia Agraria–Centro di sperimentazione e certificazione delle sementi–Sede di Tavazzano (CREA-SCS) for providingthe wheat materials tested in this study.
Funding
Financial support was provided by the Italian Ministry of Health through the National Current Research Program (Ref. IZS LT 14/11 RC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Annalisa Paternò declares that she has no conflict of interest. Daniela Verginelli declares that she has no conflict of interest. Pamela Bonini declares that she has no conflict of interest. Marisa Misto declares that she has no conflict of interest. Cinzia Quarchioni declares that she has no conflict of interest. Enrico Dainese declares that he has no conflict of interest. Stefania Peddis declares that she has no conflict of interests. Cristiana Fusco declares that she has no conflict of interests. Daniela Vinciguerra declares that she has no conflict of interests. Ugo Marchesi declares that he has no conflict of interests.
Ethical Approval
This article does not contain any studies with human or animal subjects performed by any of the authors.
Informed Consent
Informed consent is not applicable for the nature of this study.
Electronic Supplementary Material
ESM 1
(PDF 97 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Paternò, A., Verginelli, D., Bonini, P. et al. In-House Validation and Comparison of Two Wheat (Triticum aestivum) Taxon-Specific Real-Time PCR Methods for GMO Quantification Supported by Droplet Digital PCR. Food Anal. Methods 11, 1281–1290 (2018). https://doi.org/10.1007/s12161-017-1097-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-017-1097-6