Abstract
Protein p53 is mostly known for playing a key role in tumour suppression, and mutations in the p53 gene are amongst the most frequent genomic events accompanying oncogenic transformation. Continuous research is conducted to target disordered proteins/protein regions for cancer therapy, for which atomic level information is also necessary. The disordered N-terminal part of p53 contains the transactivation and the proline-rich domains—which besides being abundant in proline residues—contains repetitive Pro-Ala motifs. NMR assignment of such repetitive, proline-rich regions is challenging due to the lack of amide protons in the 1HN-detected approaches, as well as due to the small chemical shift dispersion. In the present study we perform the full assignment of the p531–100 region by applying a combination of 1HN- and 1Hα-detected NMR experiments. We also show the increased information content when using real-time homo- and heteronuclear decoupled acquisition schemes. On the other hand, we highlight the presence of minor proline species, and using Pro-selective experiments we determine the corresponding cis or trans conformation. Secondary chemical shifts for (Cα–Cβ) atoms indicate the disordered nature of this region, with expected helical tendency for the TAD1 region. As the role of the proline-rich domain is yet not well understood our results can contribute to further successful investigations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Biological context
The tumour suppressor p53 protein (human p53 UniProt ID: P04637) is a well-known transcriptional factor regulating key cellular processes upon genotoxic stress: controls DNA damage repair mechanisms, triggers cell cycle arrest and induces apoptosis in the nucleus or in the cytosol (Green and Kroemer 2009; Follis et al. 2015). The loss of p53 function results in uncontrolled cell growth. In normal cells, p53 level is low, and it is regulated by murine double minute 2 (MDM2) ubiquitin ligase. This interaction is a major cancer therapy target (Espadinha et al. 2022; Koo et al. 2022; Vassilev et al. 2004).
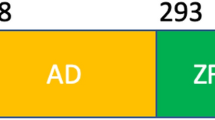
In its functional state, the 393 residue-long p53 is a homotetramer. Each monomer contains all main functional domains (Fig. 1). The N-terminal transactivation domain (TAD) is highly mobile and disordered with nascent helical elements (Wells et al. 2008). This region is followed by the proline-rich domain (PRD). Previous studies suggest that PRD plays mainly a structural role (Wells et al. 2008; Toledo et al. 2007), however several hot spot mutations are localized between residues 55–100 (72, 73, 82, 84, 89, 98) (Hoyos et al. 2022). Residues involved in the mutations can be found in PXXP sequence repeats (X: any amino acids) which is the consensusSH3 domain interaction site. The mutations can alter several protein-protein interactions (Toledo et al. 2006; Berger et al. 2001). On the other hand, it has been revealed that cis-trans isomerization of Pro82 by Pin1 is essential for consequent phosphorylation on Ser20 by checkpoint kinase 2 (CHK-2). Thus, p53 sensitivity to inhibition by MDM-2 increases (Berger et al. 2005).
The N-terminal TAD and PRD region of p53 protein has been investigated by NMR with various sequence length and sample conditions, however, full-assignment of the proline-rich region has not been achieved (BMRB ID: 17760, 50960, 51124) (Wong et al. 2009; Usluer et al. 2021; Mandal et al. 2022). All studies showed successful assignment for more than 95% of the TAD domain (1–60), but proline resonance assignment and the full assignment of the repetitive motifs in PRD region is missing.
Here, we aim to further explore the N-terminal region of p531–100 towards better understanding the interactions of the proline-rich region. We report the first, full, solution state backbone assignment of the wild type p531–100 and minor conformers arising from proline cis-trans isomerization using recently published 1HN and 1Hα-detected NMR approaches combined with real-time homo- and heteronuclear decoupling schemes.
1HN-detection does not to provide full assignment of disordered proteins, especially if the sequence is abundant in proline residues—which lack the amide proton. A solution to this is 1Hα -detection, where all residues are detectable. Another favourable feature of this approach is that Hα is a not an exchangeable proton, therefore it is suitable for samples at physiological pH and temperature (Mantylahti et al. 2010; Wong et al. 2020; Bodor et al. 2020). In the present study, we used the previously introduced sensitive, high resolution 2D SHACA-HSQC including the BASEREX hetero- and homonuclear decoupling scheme during acquisition (Haller et al. 2019; Bodor et al. 2020). BASEREX was also applied in various 3D experiments to improve resolution (Sebák et al. 2022; Szabó et al. 2022). Determination of the exact proline conformation was done by the selective Pro-(H)CBCGCAHA experiment. On the other hand, in case of 1HN-detection we applied the BEST-TROSY measurement incorporating real-time pure shift acquisition containing the 13C-BIRDX inversion element to increase spectral resolution in peak picking (Haller et al. 2022).
Domain structure of p53 protein (Krois et al. 2016) with the sequence of p531–100. The studied mutations, P82S and A84G are highlighted with green and red colours. Note, that the P72R natural polymorph was used for all p53 variants in this study. TAD transactivation domain, PRD proline-rich domain, DBD DNA-binding domain, NLS nuclear localization signal, TET tetramerization domain, CTD C-terminal regulatory domain
Methods and experiments
Protein expression and purification
Human p531–100 (UniProt code: P04637) was cloned into a modified pGEX vector (pETARA) containing an N-terminal TEV-cleavable glutathione S-transferase (GST) tag. This construct was modified using the Quickchange mutagenesis technique to generate the p531–100(P82S) and p531–100(A84G) point mutants.
The different 13C/15 N-labelled p53 peptides were expressed and purified as previously (Dudás et al. 2020; Sebák et al. 2022). The constructs were transformed into Escherichia coli BL21(DE) cells and the cultures were grown in Luria-Bertani (LB) broth complemented with 100 µg/mL ampicillin. Before induction the cells were transferred into a minimal broth containing 50 mM Na2HPO4, 20 mM KH2PO4, 8.5 mM NaCl, 1 mM CaCl2, 2 mM MgSO4, 18.7 mM NH4Cl and 22.2 mM glucose (in 15N/13C-labelled form). The expression was induced with 0.25 mM isopropyl β-d-1-thiogalactopyranoside (IPTG) and the cells were incubated at 28 °C for 3 h. Pelleted cells were disintegrated by ultrasonication in a buffer containing 20 mM Tris, pH 8, 300 mM NaCl, 0.1 mM tris(2-carboxyethyl)phosphine (TCEP). Cell lysates were clarified by centrifugation at 48,000×g. The supernatants were loaded onto Protino Glutathione Agarose 4B resin (Macherey-Nagel) and the p53 constructs were eluted using 10 mM reduced glutathione in the buffer. The GST tag was eliminated using TEV protease at room temperature, overnight. After complete cleavage, GST was removed from solution by heat denaturation followed by centrifugation. P53 fragments were further purified by reversed-phase HPLC on a Jupiter 300 C5 column (Phenomenex). The peptide containing fractions were lyophilized and stored at − 80 °C. All peptides contain additional 2 residues on the N-terminus (Gly-Ser) from cloning, the assignment of these residues is not included in the study.
NMR spectroscopy
NMR samples of p531–100 variants (wt, A84G, P82S) contained 1 mM 13C, 15N-labelled protein, 20 mM MES, 20 mM NaCl, 10 mM TCEP, 3 mM NaN3, 10% D2O, 1% DSS at pH 6.0. All NMR spectra were recorded on a Bruker Avance III 700 spectrometer operating at 700.05 MHz using a Prodigy TCI H&F-C/N-D, z-gradient probe-head. 1H chemical shifts were referenced to the internal DSS standard, whereas 15N and 13C chemical shifts were referenced indirectly via the gyromagnetic ratios. Temperature was calibrated against the methanol standard sample (Findeisen et al. 2007). All measurements were performed at 298 K.
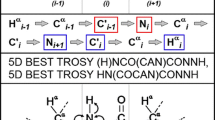
Peak assignment and sequential connectivities were determined from 1HN- and 1Hα-detected experiments: 2D 1H,15 N-BEST-HSQC and 1H,15 N-BEST-TROSY (Haller et al. 2022) and 3D BEST-type HNCACB, HN(CO)CACB and HNCO (Lescop et al. 2007). High resolution, 1Hα -detected 2D SHACA-HSQC spectra (Bodor et al. 2020) were recorded for proline peak detection, 3D HCAN and HCACON spectra were used for unambiguous peak assignment for 1Hα-13Cα correlation-based measurements (Kanelis et al. 2000; Szabó et al. 2022). Proline conformation was determined from 3D Pro-(H)CBCGCAHA measurements (Sebák et al. 2022). Experimental details and acquisition parameters are provided in Table 1. All spectra were processed with TopSpin 3.6.2 and analysed with CARA 1.8.4.2.(Keller 2004).
Extent of assignments and data deposition
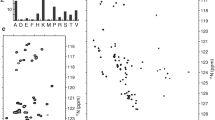
p531–100 protein is intrinsically disordered with repetitive Ala and Pro-rich motifs; thus, the signal dispersion is narrow in the 1H,15N-HSQC spectrum causing severe signal overlap (Fig. 2). The real-time homo- and heteronuclear decoupling acquisition schemes help in obtaining an increased spectral resolution (see experiments in Table 1). This allowed us to separate peaks even in the Ala-rich region. Still, this was still not sufficient for unambiguous resonance assignment. In this respect two mutations—both bearing biological relevance—were chosen: A84G and P82S. The introduction of these mutations cause perturbation in the chemical environment of the neighbouring peaks on the 1H,15N-HSQC spectra, and this helps to perform a 97% assignment of backbone 1HN and 15N resonances of the non-proline residues (76 of 78) (Fig. 2A). Only the peaks of Ser 95 and Ser 96 could not be unambiguously distinguished.
Proline residues were characterized using the 1Hα-detected approach. The high resolution 2D SHACA-HSQC spectrum allows to distinguish the 22 proline residues (Fig. 2B). Sequential connectivities were done using 3D HCAN and HCACON spectra. 1Hα-detected 2 and 3D measurements allowed 90% assignment of all residues (90 of 100). The 10 unassigned residues involve Gly59 which is not detected on the 2D SHACA-HSQC, and several distorted serine residues (94–96) still overlap. Resonance overlap hinders unambiguous assignment of 2 prolines and 1 alanine in the PRD region, as well as 2 Asp and 1 Glu residues. The type of proline isomer was determined from the Cβ and Cγ chemical shift difference, being ~ 5 ppm for trans, and ~ 10 ppm for the cis-Pro isomer (Schubert et al. 2002). For this purpose, the 3D Pro- (H)CBCGCAHA spectrum was recorded which correlates the proline sidechain Cβ and Cγ signals with the Hα–Cα crosspeaks. Results indicate that the major proline peaks are exclusively trans isomers.
2D 1H,15N-BEST-HSQC (A) and SHACA-HSQC (B) spectra of the p531–100 variants: wild type (blue), A84G (red) and P82S (green). Zoomed insets show the crowded Ala and Pro-rich parts of the BEST-HSQC and SHACA-HSQC spectra, as well as the Trp sidechain Hε1-Nε1 signals. The minor peaks are labelled with lowercase letters
As the disordered p531–100 is highly enriched in prolines (22 out of 100 residues), several small intensity minor peaks arise from the Pro cis-trans isomerization (Fig. 3). The high resolution 2D BEST-TROSY spectrum allowed the detection of more than 40 minor peaks, however due to the signal overlap in the 3D spectra and the repetitive sequence, the assignment of only 24 peaks was successful. In our previous work, the assignment of the minor peaks in p531–60 was published (Sebák et al. 2022), these peaks also appear for the p531–100 variant. As the amount of these minor conformers is between 3 and 15%, we can conclude that ratio of these minor peaks is not affected by the longer protein sequence.
In the PRD region the assignment was successful for several minor peaks. Proline isomerisation results in the appearance of minor peaks also for the neighbouring residues. For example, in the case of Pro80-Thr81-Pro82 segment - besides the major peaks - two minor peaks were detected for Thr81. The major peaks are assigned, as expected as the trans-Pro80-Thr81-trans-Pro82 segment. For the minor peaks the following connectivites are revealed: cis-pro80_1-thr81_1 fragment, as δ(13Cβ) ~ 34.2 ppm for pro80_1 is detected, and the succeeding proline is most probably trans-pro82, though the detection of this peak was not possible. Similarly, the other minor fragment is trans-pro80_2-thr81_2, as δ(13Cβ) ~ 31.7 ppm for pro80_2 is observed. The existence of thr81_2 is a consequence of the proline 82 cis-trans isomerization. This observation is strengthened by the disappearance of this minor peak for the P82S mutant. Regarding the amount of the two minor peaks: these are 3.2% and 7.0% respectively. The polar Thr81 residue in the Pro preceding position slightly increases the cis-Pro82 amount (Sebák et al. 2022).
The most intensive minor peak belongs to Trp91, as in this case the interaction between the aromatic sidechain and the Pro92 is energetically favourable. Consequently, the amount of the minor peak is more than 30%, which is in agreement with previous literature findings (Sebák et al. 2023). Most minor peak intensities in the repetitive Pro- and Ala-rich region are ~ 3–10% of the corresponding major conformer, however, unambiguous assignment is not possible. In the A84G mutant, again two minor peaks arise for Gly84 with intensities 11.7% and 7.7%, respectively.
In conclusion, using the combination of 1H-detected approaches, 100% assignment of Cα resonances was possible, including both Pro and non-Pro residues. Moreover, characterization of minor species is also given.
Further on, the obtained Cα and Cβ chemical shifts were used for secondary chemical shift calculations (SCS) using the following equation: SCS = δmeasured − δrandom coil. Random coil chemical shifts and neighbour corrections were derived from Kjaergaard et al. (Kjaergaard et al. 2011; Kjaergaard and Poulsen 2011). The calculated chemical shifts predicts the p531–100 protein to be highly disordered with a nascent helicity in TAD1 region (Ser20- Pro27)(Fig. 4.), in accordance with earlier findings.
Data Availability
The 1H, 13C and 15N backbone and sidechain chemical shifts of p531–100 have been deposited in the Biological Magnetic Resonance Data Bank (BMRB) under the accession number 51984.
Abbreviations
- BASEREX:
-
Band-selective refocusing on the X-nuclei
- CHK-2:
-
Checkpoint kinase 2
- PRD:
-
Proline rich domain
- TAD:
-
Transactivation domain
References
Berger M, Vogt Sionov R, Levine AJ, Haupt Y (2001) A role for the polyproline domain of p53 in its regulation by Mdm2. J Biol Chem 276:3785–3790. https://doi.org/10.1074/jbc.m008879200
Berger M, Stahl N, Del Sal G, Haupt Y (2005) Mutations in proline 82 of p53 impair its activation by Pin1 and Chk2 in response to DNA damage. Mol Cell Biol 25:5380–5388. https://doi.org/10.1128/mcb.25.13.5380-5388.2005
Bodor A, Haller JD, Bouguechtouli C, Theillet FX, Nyitray L, Luy B (2020) Power of pure shift HαCα correlations: a way to characterize biomolecules under physiological conditions. Anal Chem 92:12423–12428. https://doi.org/10.1021/acs.analchem.0c02182
Dudás EF, Pálfy G, Menyhárd DK, Sebák F, Ecsédi P, Nyitray L, Bodor A (2020) Tumor-suppressor p53TAD(1–60) forms a fuzzy complex with metastasis-associated S100A4: structural insights and dynamics by an NMR/MD approach. ChemBioChem 21:3087–3095. https://doi.org/10.1002/cbic.202000348
Espadinha M, Lopes EA, Marques V, Amaral JD, Djva Dos Santos, Mori M, Daniele S, Piccarducci R, Zappelli E, Martini C, Rodrigues CMP, Santos MMM (2022) Discovery of MDM2-p53 and MDM4-p53 protein-protein interactions small molecule dual inhibitors. Eur J Med Chem 241:114637. https://doi.org/10.1016/j.ejmech.2022.114637
Findeisen M, Brand T, Berger S (2007) A 1H-NMR thermometer suitable for cryoprobes. Magn Reson Chem 45:175–178. https://doi.org/10.1002/mrc.1941
Follis AV, Llambi F, Merritt P, Chipuk JE, Green DR, Kriwacki RW (2015) Pin1-induced proline isomerization in cytosolic p53 mediates BAX activation and apoptosis. Mol Cell 59:677–684. https://doi.org/10.1016/j.molcel.2015.06.029
Green DR, Kroemer G (2009) Cytoplasmic functions of the tumour suppressor p53. Nature 458:1127–1130. https://doi.org/10.1038/nature07986
Haller JD, Bodor A, Luy B (2019) Real-time pure shift measurements for uniformly isotope-labeled molecules using X-selective BIRD homonuclear decoupling. J Magn Reson 302:64–71. https://doi.org/10.1016/j.jmr.2019.03.011
Haller JD, Bodor A, Luy B (2022) Pure shift amide detection in conventional and TROSY-type experiments of (13)C,(15)N-labeled proteins. J Biomol NMR 76:213–221. https://doi.org/10.1007/s10858-022-00406-z
Hoyos D, Greenbaum B, Levine AJ (2022) The genotypes and phenotypes of missense mutations in the proline domain of the p53 protein. Cell Death Differ 29:938–945. https://doi.org/10.1038/s41418-022-00980-7
Kanelis V, Donaldson L, Muhandiram DR, Rotin D, Forman-Kay JD, Kay LE (2000) Sequential assignment of proline-rich regions in proteins: application to modular binding domain complexes. J Biomol NMR 16:253–259. https://doi.org/10.1023/a:1008355012528
Keller RLJ (2004) The computer aided resonance assignment tutorial. CANTINA Verlag, Goldau, Switzerland
Kjaergaard M, Poulsen FM (2011) Sequence correction of random coil chemical shifts: correlation between neighbor correction factors and changes in the Ramachandran distribution. J Biomol NMR 50:157–165. https://doi.org/10.1007/s10858-011-9508-2
Kjaergaard M, Brander S, Poulsen FM (2011) Random coil chemical shift for intrinsically disordered proteins: effects of temperature and pH. J Biomol NMR 49:139–149. https://doi.org/10.1007/s10858-011-9472-x
Koo N, Sharma AK, Narayan S (2022) Therapeutics targeting p53-MDM2 interaction to induce cancer cell death. Int J Mol Sci 23:5005. https://doi.org/10.3390/ijms23095005
Krois AS, Ferreon JC, Martinez-Yamout MA, Dyson HJ, Wright PE (2016) Recognition of the disordered p53 transactivation domain by the transcriptional adapter zinc finger domains of CREB-binding protein. Proc Natl Acad Sci USA 113:E1853–E1862. https://doi.org/10.1073/pnas.1602487113
Lescop E, Schanda P, Brutscher B (2007) A set of BEST triple-resonance experiments for time-optimized protein resonance assignment. J Magn Reson 187:163–169. https://doi.org/10.1016/j.jmr.2007.04.002
Mandal R, Kohoutova K, Petrvalska O, Horvath M, Srb P, Veverka V, Obsilova V, Obsil T (2022) FOXO4 interacts with p53 TAD and CRD and inhibits its binding to DNA. Protein Sci 31:e4287. https://doi.org/10.1002/pro.4287
Mantylahti S, Aitio O, Hellman M, Permi P (2010) HA-detected experiments for the backbone assignment of intrinsically disordered proteins. J Biomol NMR 47:171–181. https://doi.org/10.1007/s10858-010-9421-0
Schubert M, Labudde D, Oschkinat H, Schmieder P (2002) A software tool for the prediction of xaa-pro peptide bond conformations in proteins based on C-13 chemical shift statistics. J Biomol NMR 24:149–154. https://doi.org/10.1023/a:1020997118364
Sebák F, Ecsédi P, Bermel W, Luy B, Nyitray L, Bodor A (2022) Selective 1Hα NMR methods reveal functionally relevant proline cis/trans isomers in intrinsically disordered proteins: characterization of minor forms, effects of phosphorylation, and occurrence in proteome. Angew Chem Int Ed Engl 61:e202108361. https://doi.org/10.1002/anie.202108361
Sebák F, Szolomájer J, Papp N, Tóth GK, Bodor A (2023) Proline cis/trans isomerization in intrinsically disordered proteins and peptides. Front Biosci (Landmark Ed) 28:127. https://doi.org/10.31083/j.fbl2806127
Szabó CL, Szabó B, Sebák F, Bermel W, Tantos A, Bodor A (2022) The disordered EZH2 loop: atomic level characterization by 1HN- and 1Hα-detected NMR approaches, interaction with the long noncoding HOTAIR RNA. Int J Mol Sci 23:6150. https://doi.org/10.3390/ijms23116150
Toledo F, Krummel KA, Lee CJ, Liu CW, Rodewald LW, Tang M, Wahl GM (2006) A mouse p53 mutant lacking the proline-rich domain rescues Mdm4 deficiency and provides insight into the Mdm2-Mdm4-p53 regulatory network. Cancer Cell 9:273–285. https://doi.org/10.1016/j.ccr.2006.03.014
Toledo F, Lee CJ, Krummel KA, Rodewald LW, Liu CW, Wahl GM (2007) Mouse mutants reveal that putative protein interaction sites in the p53 proline-rich domain are dispensable for tumor suppression. Mol Cell Biol 27:1425–1432. https://doi.org/10.1128/mcb.00999-06
Usluer S, Spreitzer W, Bourgeois B, Madl T (2021) p53 transactivation domain mediates binding and phase separation with Poly-PR/GR. Int J Mol Sci 22:11431. https://doi.org/10.3390/ijms222111431
Vassilev LT, Vu BT, Graves B, Carvajal D, Podlaski F, Filipovic Z, Kong N, Kammlott U, Lukacs C, Klein C, Fotouhi N, Liu EA (2004) In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 303:844–848. https://doi.org/10.1126/science.1092472
Wells M, Tidow H, Rutherford TJ, Markwick P, Jensen MR, Mylonas E, Svergun DI, Blackledge M, Fersht AR (2008) Structure of tumor suppressor p53 and its intrinsically disordered N-terminal transactivation domain. Proc Natl Acad Sci USA 105:5762–5767. https://doi.org/10.1073/pnas.0801353105
Wong TS, Rajagopalan S, Freund SM, Rutherford TJ, Andreeva A, Townsley FM, Petrovich M, Fersht AR (2009) Biophysical characterizations of human mitochondrial transcription factor A and its binding to tumor suppressor p53. Nucleic Acids Res 37:6765–6783. https://doi.org/10.1093/nar/gkp750
Wong LE, Kim TH, Muhandiram DR, Forman-Kay JD, Kay LE (2020) NMR experiments for studies of dilute and condensed protein phases: application to the phase-separating protein CAPRIN1. J Am Chem Soc 142:2471–2489. https://doi.org/10.1021/jacs.9b12208
Acknowledgements
Financial support of the National Research, Development and Innovation Office, Hungary, grant numbers NKFI-K124900, K137940 is greatly acknowledged.
Funding
Open access funding provided by Eötvös Loránd University. This research was supported by the National Research, Development and Innovation Office, Hungary, grant numbers NKFI-K124900, K137940.
Author information
Authors and Affiliations
Contributions
FS and AB performed experiments and did the resonance assignment. PE and LN contributed to protein expression. All authors wrote the manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
All authors have agreed to the publication of the manuscript.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sebák, F., Ecsédi, P., Nyitray, L. et al. Assignment of the disordered, proline-rich N-terminal domain of the tumour suppressor p53 protein using 1HN and 1Hα-detected NMR measurements. Biomol NMR Assign 17, 309–314 (2023). https://doi.org/10.1007/s12104-023-10160-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-023-10160-4