Abstract
Neuroplasticity and synaptic transmission in the brain are regulated by N-methyl-D-aspartate receptors (NMDARs) that consist of hetero-tetrameric combinations of the glycine-binding GluN1 and glutamate-binding GluN2 subunits. Calmodulin (CaM) binds to the cytosolic C0 domain of GluN1 (residues 841–865) that may play a role in the Ca2+-dependent inactivation (CDI) of NMDAR channel activity. Dysregulation of NMDARs are linked to various neurological disorders, including Alzheimer’s disease, depression, stroke, epilepsy, and schizophrenia. Here, we report complete NMR chemical shift assignments of Ca2+-saturated CaM bound to the GluN1 C0 domain of the human NMDAR (BMRB no. 51715).
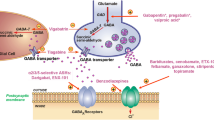
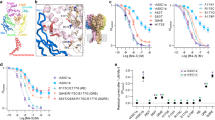
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Biological context
N-methyl-D-aspartate receptors (NMDARs) in the brain are localized at the post-synaptic membrane where they regulate neuronal excitability and confer synaptic plasticity (Traynelis et al. 2010). NMDARs contain two copies each of GluN1 and GluN2 subunits, which activate upon binding to the co-agonist glycine and neurotransmitter agonist glutamate, respectively (Benveniste and Mayer 1991; Clements and Westbrook 1991). Under resting basal conditions, the intracellular Ca2+ concentration is kept below 100 nM due to the powerful action of Ca2+ pumps and exchangers (Berridge et al. 2003; Clapham 2007), and Ca2+ sequestration into stores (Berridge 2002; Clapham 2007). Ligand-gated opening of NMDAR channels at the postsynaptic membrane causes intracellular Ca2+ levels to increase into the micromolar range (Wadel et al. 2007), causing a wide range of Ca2+-dependent processes (Luscher and Malenka 2012; Kunz et al. 2013; Puri 2020). Prolonged elevation of intracellular Ca2+ levels is cytotoxic (Stanika et al. 2012), and NMDAR channels are negatively regulated by a process known as Ca2+-dependent inactivation (CDI) (Iacobucci and Popescu 2017, 2019, 2020). The Ca2+-dependent inactivation of NMDA receptors requires CaM binding to the cytosolic C0 domain in GluN1 (Zhang and Majerus 1998; Iacobucci and Popescu 2017, 2019, 2020). Ca2+-free CaM (apoCaM) is believed to be pre-associated with the C0 domain that may cause channel activation at low Ca2+ levels (Wang et al. 2008; Iacobucci and Popescu 2017, 2019, 2020). Neurotransmitter binding to NMDAR causes channel opening, which triggers a rise in intracellular Ca2+ that promotes a conformational change in the NMDAR/CaM complex, leading to CDI (Krupp et al. 1999; Wang et al. 2008; Iacobucci and Popescu 2020).
Atomic-level structures of NMDARs have been solved by x-ray crystallography (Karakas and Furukawa 2014; Lee et al. 2014) and cryo-EM (Jalali-Yazdi et al. 2018; Regan et al. 2018; Chou et al. 2020) that show detailed inter-subunit interactions between the extracellular ligand-binding domain and transmembrane channel domain. However, the C-terminal cytosolic domain (that mediates CDI) is not visible in any of the published structures and its structure has remained elusive. The cytosolic region of GluN1 (residues 830–938) is comprised of helical domains called C0 (residues 841–865) and C1 (residues 875–898) that interact with CaM (Ehlers et al. 1996; Krupp et al. 1999; Ataman et al. 2007). A crystal structure of CaM bound to the C1 domain has been reported (Ataman et al. 2007), but a structure of CaM bound to C0 is currently not known. We report here NMR resonance assignments of Ca2+-saturated CaM bound to the C0 domain of GluN1 (hereafter called CaM/GluN1 C0). These assignments are an important step toward elucidating the complete structure of CaM bound to GluN1, which may be important for understanding the mechanism of CDI.
Methods and experiments
Expression and purification of CaM
Human CaM was overexpressed in E. coli strain BL21(DE3) using pET11b (Novagen) and the expressed protein was purified as described previously (Bej and Ames 2022a). The CaM protein samples typically have greater than 99% purity as confirmed by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE). A peptide fragment of the GluN1 C0 domain of the NMDA receptor (residues 841–865) was purchased from GenScript, dissolved in DMSO-d6, and quantified using UV–Vis absorption spectroscopy (ε280 = 5500 M−1 cm−1). A 2.5-fold excess of the peptide was added to Ca2+-bound CaM, incubated at room temperature for 30 min, and concentrated to 0.4 mM in a final volume of 0.3 ml.
NMR spectroscopy
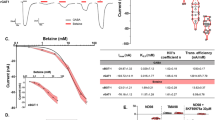
NMR samples of isotopically labeled CaM bound to the unlabeled GluN1 C0 peptide (complex is called CaM/GluN1 C0) were dissolved in 20 mM Tris-d11 (pH 7.0) and 1 mM CaCl2 containing 8% (v/v) D2O and placed in a Shigemi NMR tube (Shigemi Inc.). All NMR experiments on CaM/GluN1 C0 were performed at 308 K using a Bruker Avance III 800 MHz spectrometer equipped with a triple resonance cryogenic (TCI) probe. The 15N-1H HSQC spectrum of CaM/GluN1 C0 (Fig. 1A, B) was recorded with 182 × 2048 complex points for 15N(F1) and 1H(F2). Backbone resonances were assigned by analyzing triple resonance spectra: HNCACB, HN(CO)CACB, HNCO, HBHA(CO)NH, and HBHANH. Side chain resonances (aliphatic (Fig. 1C) and aromatic) were assigned by analyzing HCCCONH-TOCSY, HBCBCGCDHD and HBCBCGCDCEHE as described by (Ikura et al. 1991). The NMR data were processed by NMRPipe (Delaglio et al. 1995) and assignments were obtained using Sparky (Lee et al. 2015).
NMR spectra of Ca2+-saturated CaM bound to unlabeled GluN1 C0 peptide. A 15N-1H HSQC spectrum recorded at 800 MHz 1H frequency. The black-dashed box highlights spectrally crowded region. B Expanded view of the resonance assignment from the spectrally crowded region highlighted with black-dashed box. C Constant-time 13C-1H HSQC spectrum of Ca2+-saturated CaM bound to the peptide
Extent of assignments and data deposition
Backbone resonance assignments of CaM/GluN1 C0 are illustrated in the 15N-1H HSQC spectrum of 15N-labeled CaM bound to unlabeled C0 peptide (Fig. 1A, B). Side chain aliphatic resonance assignments are shown by labeled peaks in the constant-time 13C-1H HSQC spectrum (Fig. 1C). NMR assignments were derived from the analysis of 3D heteronuclear NMR experiments performed on 13C/15N-labeled CaM bound to unlabeled C0 peptide. The highly dispersed spectral peaks and uniform peak intensities suggest that CaM/GluN1 C0 complex is stable in solution and properly folded. The downfield amide peaks assigned to G26, G62, G99 and G135 indicate that Ca2+ is bound to each of the four EF-hands (Fig. 1A). Upfield-shifted side chain methyl peaks assigned to I28, L33, V36, L40, M72, I101, V109 and V143 (Fig. 1C) suggest these residues may interact with aromatic side chain atoms located in the hydrophobic core. At least 97% of the non-proline backbone resonances (1HN, 15N, 13Cα, 13Cβ, and 13CO) and 97% of side-chain resonances were assigned. Only K78 in the second EF-hand of CaM remains unassigned, because its HSQC peak could not be detected. The chemical shift assignments (1H, 15N, 13C) for CaM/GluN1 C0 have been deposited in the BioMagResBank (http://www.bmrb.wisc.edu) under accession number 51715.
The CaM/GluN1 C0 secondary structure was calculated based on chemical shift index (Wishart et al. 1992) and ANN-Secondary structure prediction using TALOS + (Shen et al. 2009) (Fig. 2). The CaM/GluN1 C0 secondary structure is similar to that reported previously for free CaM (Bej and Ames 2022b), and is depicted by cylinders and triangles in Fig. 2A. The analysis of sequential NOE patterns along with long-range NOE-derived distances identify a total of eight α-helices and four β-strands that make up four EF-hands (EF1: residues 7–39, EF2: residues 45–76, EF3: residues 83–112 and EF4: residues 119–144) as seen in the crystal structure of Ca2+-bound CaM (Babu et al. 1988). Two N-terminal EF-hands (EF1 and EF2) form the CaM N-lobe, and C-terminal EF-hands (EF3 and EF4) form the C-lobe. The GluN1 C0 peptide binds to CaM and causes chemical shift perturbations (CSPs) for hydrophobic CaM residues in the N-lobe (F20, I28, L33, L40, L49, M52, A58, M72) and C-lobe (A89, V92, L106, V109, M145, M146) (Fig. 3A, B), suggesting that the GluN1 C0 peptide is making contact with both lobes of CaM as seen in previous collapsed structures of CaM bound to other peptides (Hoeflich and Ikura 2002).
Secondary structure and RCI order parameters (S2) of Ca2+-saturated CaM bound to GluN1 C0 peptide predicted from the assigned backbone chemical shifts. A Probability of secondary structural elements (cyan for helix and magenta for strand) and B RCI S2 of Ca2+-saturated CaM bound to the GluN1 C0 peptide were predicted using TALOS + server (Shen et al. 2009). The wire diagram depicting the secondary structural elements (cyan cylinder for helix and magenta triangle for strand) was obtained from the CaM structure (PDB ID: 2VAY (Halling et al. 2009))
Residue-specific chemical shift perturbation (CSP) for Ca2+-saturated CaM in GluN1 C0 peptide-bound and unbound conditions. A Backbone amide CSP was calculated as: \(CSP= \sqrt{{\left(\Delta {H}^{N}\right)}^{2}+{\left(0.14\times \Delta N\right)}^{2}}\). ΔHN and ΔN are the observed difference in the 1HN and 15N chemical shifts, respectively between peptide-bound and unbound Ca2+-saturated CaM (Bej and Ames 2022b). B Side-chain methyl CSP was calculated as: \(CSP= \sqrt{{\left(\Delta H\right)}^{2}+{\left(0.14\times \Delta C\right)}^{2}}\). ΔH and ΔC are the observed difference in the 1H and 13C methyl chemical shifts, respectively between peptide-bound and unbound Ca2+-saturated CaM. CSP values are mapped on to the CaM structure (PDB ID: 2VAY (Halling et al. 2009)). Residues with significant CSP are showed in spheres and labeled accordingly. Residues, without CSP values including proline, amino acids without methyl group, or unassigned resonances, are colored as gray
The amino acid sequence of GluN1 C0 when aligned with the IQ-motif of L-type Ca2+ channels (CaV1.1 and CaV1.2) reveal critical conserved hydrophobic residues that likely contact CaM (Fig. 4A). The GluN1 C0 residues L850, A854 and W858 align with conserved hydrophobic residues of the IQ-motif that contact the CaM C-lobe (highlighted red in Fig. 4A) as seen in the known structures of CaM bound to the IQ-motif of CaV1.2 (Fig. 4B) (Fallon et al. 2005) and CaV1.1 (Halling et al. 2009). Therefore, we predict that these residues in GluN1 C0 likely contact the CaM C-lobe. In addition, the GluN1 C0 residues M848, A851 and F852 align with conserved hydrophobic residues (highlighted blue in Fig. 4A) that contact the CaM N-lobe in the crystal structure of CaM bound to the CaV1.2 IQ peptide (Fig. 4B), suggesting that these residues from GluN1 C0 may contact the CaM N-lobe. Future NMR studies are needed to determine the NMR structure of CaM/GluN1 C0 and test whether it is similar to the collapsed structure of CaM bound to CaV1.2 IQ (Fig. 4B). The NMR assignments of CaM/GluN1 C0 presented here are an important first step toward determining the full three-dimensional structure of CaM bound to GluN1 C0.
Conserved residues in GluN1 C0 that may contact CaM. A Sequence alignment of GluN1 C0 with the IQ motif of CaV1.1 and CaV1.2. Residues contacting the N-lobe and C-lobe of CaM are highlighted in blue and red, respectively. B Crystal structure of CaM bound to CaV1.2 IQ peptide (PDB: 2F3Y (Fallon et al. 2005)). Key residues involved in interaction with CaM are labeled
Data availability
The assignments have been deposited to the BMRB under the accession code: 51715.
References
Ataman ZA, Gakhar L, Sorensen BR, Hell JW, Shea MA (2007) The NMDA receptor NR1 C1 region bound to calmodulin: structural insights into functional differences between homologous domains. Structure 15:1603–1617
Babu YS, Bugg CE, Cook WJ (1988) Structure of calmodulin refined at 2.2 A resolution. J Mol Biol 204:191–204
Bej A, Ames JB (2022a) Chemical shift assignments of calmodulin bound to the beta-subunit of a retinal cyclic nucleotide-gated channel (CNGB1). Biomol NMR Assign 16:147–151
Bej A, Ames JB (2022b) Chemical shift assignments of calmodulin under standard conditions at neutral pH. Biomol NMR Assign 16:213
Benveniste M, Mayer ML (1991) Kinetic analysis of antagonist action at N-methyl-D-aspartic acid receptors. Two binding sites each for glutamate and glycine. Biophys J 59:560–573
Berridge MJ (2002) The endoplasmic reticulum: a multifunctional signaling organelle. Cell Calcium 32:235–249
Berridge MJ, Bootman MD, Roderick HL (2003) Calcium signalling: dynamics, homeostasis and remodelling. Nat Rev Mol Cell Biol 4:517–529
Chou TH, Tajima N, Romero-Hernandez A, Furukawa H (2020) Structural basis of functional transitions in mammalian NMDA receptors. Cell 182(357–371):e13
Clapham DE (2007) Calcium signaling. Cell 131:1047–1058
Clements JD, Westbrook GL (1991) Activation kinetics reveal the number of glutamate and glycine binding sites on the N-methyl-D-aspartate receptor. Neuron 7:605–613
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeiffer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Ehlers MD, Zhang S, Bernhadt JP, Huganir RL (1996) Inactivation of NMDA receptors by direct interaction of calmodulin with the NR1 subunit. Cell 84:745–755
Fallon JL, Halling DB, Hamilton SL, Quiocho FA (2005) Structure of calmodulin bound to the hydrophobic IQ domain of the cardiac Ca(v)1.2 calcium channel. Structure 13:1881–1886
Halling DB, Georgiou DK, Black DJ, Yang G, Fallon JL, Quiocho FA, Pedersen SE, Hamilton SL (2009) Determinants in CaV1 channels that regulate the Ca2+ sensitivity of bound calmodulin. J Biol Chem 284:20041–20051
Hoeflich KP, Ikura M (2002) Calmodulin in action: diversity in target recognition and activation mechanisms. Cell 108:739–742
Iacobucci GJ, Popescu GK (2017) Resident calmodulin primes NMDA receptors for Ca(2+)-dependent inactivation. Biophys J 113:2236–2248
Iacobucci GJ, Popescu GK (2019) Spatial coupling tunes NMDA receptor responses via Ca(2+) diffusion. J Neurosci 39:8831–8844
Iacobucci GJ, Popescu GK (2020) Ca(2+)-Dependent Inactivation of GluN2A and GluN2B NMDA receptors occurs by a common kinetic mechanism. Biophys J 118:798–812
Ikura M, Spera S, Barbato G, Kay LE, Krinks M, Bax A (1991) Secondary structure and side-chain 1H and 13C resonance assignments of calmodulin in solution by heteronuclear multidimensional NMR spectroscopy. Biochemistry 30:9216–9228
Jalali-Yazdi F, Chowdhury S, Yoshioka C, Gouaux E (2018) Mechanisms for zinc and proton inhibition of the GluN1/GluN2A NMDA receptor. Cell 175(1520–1532):e15
Karakas E, Furukawa H (2014) Crystal structure of a heterotetrameric NMDA receptor ion channel. Science 344:992–997
Krupp JJ, Vissel B, Thomas CG, Heinemann SF, Westbrook GL (1999) Interactions of calmodulin and alpha-actinin with the NR1 subunit modulate Ca2+-dependent inactivation of NMDA receptors. J Neurosci 19:1165–1178
Kunz PA, Roberts AC, Philpot BD (2013) Presynaptic NMDA receptor mechanisms for enhancing spontaneous neurotransmitter release. J Neurosci 33:7762–7769
Lee CH, Lu W, Michel JC, Goehring A, Du J, Song X, Gouaux E (2014) NMDA receptor structures reveal subunit arrangement and pore architecture. Nature 511:191–197
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327
Luscher C, Malenka RC (2012) NMDA receptor-dependent long-term potentiation and long-term depression (LTP/LTD). Cold Spring Harb Perspect Biol 4:a005710
Puri BK (2020) Calcium signaling and gene expression. Adv Exp Med Biol 1131:537–545
Regan MC, Grant T, McDaniel MJ, Karakas E, Zhang J, Traynelis SF, Grigorieff N, Furukawa H (2018) Structural mechanism of functional modulation by gene splicing in NMDA receptors. Neuron 98(521–529):e3
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44:213–223
Stanika RI, Villanueva I, Kazanina G, Andrews SB, Pivovarova NB (2012) Comparative impact of voltage-gated calcium channels and NMDA receptors on mitochondria-mediated neuronal injury. J Neurosci 32:6642–6650
Traynelis SF, Wollmuth LP, McBain CJ, Menniti FS, Vance KM, Ogden KK, Hansen KB, Yuan H, Myers SJ, Dingledine R (2010) Glutamate receptor ion channels: structure, regulation, and function. Pharmacol Rev 62:405–496
Wadel K, Neher E, Sakaba T (2007) The coupling between synaptic vesicles and Ca2+ channels determines fast neurotransmitter release. Neuron 53:563–575
Wang C, Wang HG, Xie H, Pitt GS (2008) Ca2+/CaM controls Ca2+-dependent inactivation of NMDA receptors by dimerizing the NR1 C termini. J Neurosci 28:1865–1870
Wishart DS, Sykes BD, Richards FM (1992) The chemical shift index: a fast and simple method for the assignment of protein secondary structure through NMR spectroscopy. Biochemistry 31:1647–1651
Zhang X, Majerus PW (1998) Phosphatidylinositol signalling reactions. Semin Cell Dev Biol 9:153–160
Acknowledgements
We thank Derrick Kaseman and Ping Yu for help with NMR experiments performed at the UC Davis NMR Facility.
Funding
Work supported by NIH grants to J.B.A (R01 EY012347) and to the UC Davis NMR Facility (RR11973).
Author information
Authors and Affiliations
Contributions
A.B. performed all experiments, analyzed data and helped write the manuscript. J.B.A directed the overall project and wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no competing conflict of interest.
Ethical approval
The experiments comply with the current laws of the United States.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bej, A., Ames, J.B. Chemical shift assignments of calmodulin bound to the GluN1 C0 domain (residues 841–865) of the NMDA receptor. Biomol NMR Assign 17, 61–65 (2023). https://doi.org/10.1007/s12104-023-10121-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-023-10121-x