Abstract
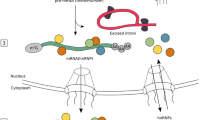
Adenosine-to-inosine (A-to-I) editing of a subset of RNAs in a eukaryotic cell is required in order to avoid triggering the innate immune system. Editing is carried out by ADAR1, which exists as short (p110) and long (p150) isoforms. ADAR1p150 is mostly cytoplasmic, possesses a Z-RNA binding domain (Zα), and is only expressed during the innate immune response. A structurally homologous domain to Zα, the Zβ domain, is separated by a long linker from Zα on the N-terminus of ADAR1 but its function remains unknown. Zβ does not bind to RNA in isolation, yet the binding kinetics of the segment encompassing Zα, Zβ and the 95-residue linker between the two domains (Zα–Zβ) are markedly different compared to Zα alone. Here we present the solution NMR backbone assignment of Zα–Zβ from H. Sapiens ADAR1. The predicted secondary structure of Zα–Zβ based on chemical shifts is in agreement with previously determined structures of Zα and Zβ in isolation, and indicates that the linker is intrinsically disordered. Comparison of the chemical shifts between the individual Zα and Zβ domains to the full Zα–Zβ construct suggests that Zβ may interact with the linker, the function of which is currently unknown.




Similar content being viewed by others
Data availability
The chemical shift assignments for Zα (BMRB 50714), Zβ (BMRB 50713) and Zα–Zβ (BMRB 50715) have been deposited in the Biological Magnetic Resonance Data Bank.
References
Athanasiadis A (2012) Zalpha-domains: At the intersection between RNA editing and innate immunity. Semin Cell Dev Biol 23:275–280
Athanasiadis A, Placido D, Maas S, Brown BA, Lowenhaupt K, Rich A (2005) The crystal structure of the Zβ domain of the RNA-editing enzyme ADAR1 reveals distinct conserved surfaces among Z-domains. J Mol Biol 351(3):496–507
Bass BL, Weintraub H (1988) An unwinding activity that covalently modifies its double-stranded RNA substrate. Cell. https://doi.org/10.1016/0092-8674(88)90253-X
Brown BA, Lowenhaupt K, Wilbert CM et al (2000) The Zα domain of the editing enzyme dsRNA adenosine deaminase binds left-handed Z-RNA as well as Z-DNA. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.240464097
Cai M, Huang Y, Yang R et al (2016) A simple and robust protocol for high-yield expression of perdeuterated proteins in Escherichia coli grown in shaker flasks. J Biomol NMR. https://doi.org/10.1007/s10858-016-0052-y
Cavanagh J (2007) Protein NMR spectroscopy: principles and practice. Academic Press, London
Chung H, Calis JJA, Wu X et al (2018) Human ADAR1 prevents endogenous RNA from triggering translational shutdown. Cell. https://doi.org/10.1016/j.cell.2017.12.038
Delaglio F, Grzesiek S, Vuister GW et al (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293. https://doi.org/10.1007/BF00197809
George CX, Samuel CE (1999) Human RNA-specific adenosine deaminase ADAR1 transcripts possess alternative exon 1 structures that initiate from different promoters, one constitutively active and the other interferon inducible. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.96.8.4621
Herbert A, Schade M, Lowenhaupt K et al (1998) The Zα domain from human ADAR1 binds to the Z-DNA conformer of many different sequences. Nucleic Acids Res. https://doi.org/10.1093/nar/26.15.3486
Hyberts SG, Milbradt AG, Wagner AB et al (2012) Application of iterative soft thresholding for fast reconstruction of NMR data non-uniformly sampled with multidimensional Poisson Gap scheduling. J Biomol NMR. https://doi.org/10.1007/s10858-012-9611-z
Kruse H, Mrazikova K, D’Ascenzo L et al (2020) Short but weak: the Z-DNA Lone-Pair⋅⋅⋅π conundrum challenges standard carbon Van der Waals radii. Angew Chemie - Int Ed. https://doi.org/10.1002/anie.202004201
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327. https://doi.org/10.1093/bioinformatics/btu830
Marsh JA, Singh VK, Jia Z, Forman-Kay JD (2006) Sensitivity of secondary structure propensities to sequence differences between α- and γ-synuclein: implications for fibrillation. Protein Sci. https://doi.org/10.1110/ps.062465306
Montaville P, Coudevylle N, Radhakrishnan A et al (2008) The PIP2 binding mode of the C2 domains of rabphilin-3A. Protein Sci. https://doi.org/10.1110/ps.073326608
Nishikura K (2016) A-to-I editing of coding and non-coding RNAs by ADARs. Nat Rev Mol Cell Biol 17:83–96
O’Connell MA, Keller W (1994) Purification and properties of double-stranded RNA-specific adenosine deaminase from calf thymus. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.91.22.10596
O’Connell MA, Krause S, Higuchi M et al (1995) Cloning of cDNAs encoding mammalian double-stranded RNA-specific adenosine deaminase. Mol Cell Biol. https://doi.org/10.1128/mcb.15.3.1389
Patterson JB, Samual SE (1995) Expression and regulation by interferon of a double-stranded-RNA-specific adenosine deaminase from human cells: evidence for two forms of the deaminase. Mol Cell Biol 15:5376–5388
Placido D, Brown BA, Lowenhaupt K et al (2007) A left-handed RNA double helix bound by the Zα domain of the RNA-editing enzyme ADAR1. Structure. https://doi.org/10.1016/j.str.2007.03.001
Schade M, Turner CJ, Kühne R et al (1999a) The solution structure of the Za domain of the human RNA editing enzyme ADAR1 reveals a prepositioned binding surface for Z-DNA. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.96.22.12465
Schade M, Turner CJ, Lowenhaupt K et al (1999b) Structure-function analysis of the Z-DNA-binding domain Zα of dsRNA adenosine deaminase type I reveals similarity to the (α + β) family of helix-turn-helix proteins. EMBO J. https://doi.org/10.1093/emboj/18.2.470
Schwartz T, Lowenhaupt K, Kim YG et al (1999a) Proteolytic dissection of Zab, the Z-DNA-binding domain of human ADAR1. J Biol Chem. https://doi.org/10.1074/jbc.274.5.2899
Schwartz T, Rould MA, Lowenhaupt K et al (1999b) Crystal structure of the Zalpha domain of the human editing enzyme ADAR1 bound to left-handed Z-DNA. Science 11:1841–1845
Vranken WF, Boucher W, Stevens TJ et al (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins Struct Funct Genet 59:687–696. https://doi.org/10.1002/prot.20449
Wagner RW, Smith JE, Cooperman BS, Nishikura K (1989) A double-stranded RNA unwinding activity introduces structural alterations by means of adenosine to inosine conversions in mammalian cells and Xenopus eggs. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.86.8.2647
Williamson MP (2013) Using chemical shift perturbation to characterise ligand binding. Progr Nuclear Magn Reson Spectrosc 73:1–6
Acknowledgements
The authors thank David Jones (University of Colorado, Aurora) for help with NMR spectroscopy, and Jeffrey Kieft for support. This project was supported by NIH Grant R01GM130694-01A1, NSF Grant 1917254 for Infrastructure Innovation for Biological Research, and a start-up package by the University of Colorado to B.V., and University of Colorado Cancer Center Support Grant P30 CA046934 and NIH Biomedical Research Support Shared Grant S10 OD025020-01.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nichols, P.J., Henen, M.A., Vicens, Q. et al. Solution NMR backbone assignments of the N-terminal Zα-linker-Zβ segment from Homo sapiens ADAR1p150. Biomol NMR Assign 15, 273–279 (2021). https://doi.org/10.1007/s12104-021-10017-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-021-10017-8