Abstract
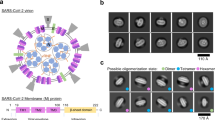
Vaccinia virus (VACV) belonging to the poxvirus family enters the host cell via two different entry pathways; either endocytosis or virus/host cell membrane fusion. With respect to the virus/host cell membrane fusion, there are eleven viral membrane proteins forming a complicated entry-fusion complex (EFC), including A28, A21, A16, F9, G9, G3, H2, J5, L5, L1 and O3, to conduct the fusion function. These EFC components are highly conserved in all poxviruses and each of them is essential and necessary for the fusion activity. So far, with the exceptions of L1 and F9 whose crystal structures were reported, the structural information about other EFC components remains largely unclear. We aim to conduct a structural and functional investigation of VACV virus-entry membrane protein A28. In this work, we expressed and purified a truncated form of A28 (14 kDa; residues 38–146, abbreviated as tA28 hereinafter), with deletion of its transmembrane domain (residues 1–22) and a hydrophobic segment (residues 23–37). And the assignments of its backbone and side chain 1H, 13C and 15N chemical shifts of tA28 are reported. The secondary structure propensity from TALOS+ indicates that tA28 does contain three α-helices, six β-strands and connecting loops. Aside from this, we demonstrated that tA28 does interact with fusion suppressor viral protein A26 (residues 351–500) by the 1H–15N HSQC spectrum. We interpret that A28 binding to A26 deactivates EFC fusion activity. The current study provides a valuable framework towards further structural analyses of this protein and for better understanding virus/host cell membrane fusion mechanism in association with virus entry.
Similar content being viewed by others
Biological context
Vaccinia virus (VAVC) is the prototype of the Orthopoxvirus genus of the Poxviridae family. VAVC has a large double-stranded DNA genome of 190 kb and it encodes more than 200 proteins (Goebel et al. 1990; Moss 1996). It infects a wide range of hosts and enters host cells via different entry pathways; either endocytosis or plasma membrane fusion (Carter et al. 2005; Armstrong et al. 1973; Chang and Metz 1976). During virus entry, vaccinia mature virus utilizes a highly conserved entry fusion protein complex (EFC) to mediate the virus/host cell membrane fusion. The EFC consists of eleven viral membrane proteins forming a complicated protein complex, including A28, A21, A16, F9, G9, G3, H2, J5, L1, L5 and O3, and each is essential for the virus/host membrane fusion activity. It was demonstrated that A28-deficient virions are noninfectious due to the inability to initiate virus entry (Senkevich et al. 2004). So far, the underlying mechanism triggers the virus/host cell membrane fusion remains largely unknown.
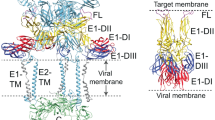
VAVC entry-fusion protein A28 contains a transmembrane anchor (residues 1–22) and a highly hydrophobic region (residues 23–37) at the N-terminus. The secondary structure prediction suggested that it has primarily a β-sheet structure, with six putative β-strands and two flanking α-helices (Senkevich et al. 2004). Based on the pull-down experiments, it was shown that A28 and H2 form a sub-complex, so does A16 and G9 (Wagenaar et al. 2008; Chang et al. 2012), and G3 and L5 (Wolfe and Moss 2011). Noteworthy the EFC sub-complexes are able to anchor to mature virus membrane even in the absence of other EFC components, with the exception of A28 (Senkevich et al. 2005). Aside from this, it was reported that EFC proteins A16 and G9 from mature virion do bind to its envelope protein A26 at neutral pH. When the mature virion was treated with acidic buffer, the A26-A27 protein complex dissociated from the virion at low pH (Chang et al. 2012, 2019). Most importantly, crystal structure of A26 protein was recently resolved and further mutagenesis revealed that His48 and His53 located in the N-terminal alpha2 helix are responsible for acid-induced conformational changes of A26 protein, leading to EFC activation and viral and vesicular membrane fusion (Chang et al. 2012, 2019). Thus, based on these data, it was concluded that VAVC envelope protein A26 acts as an acid-sensitive fusion suppressor regulating the virus entry pathways.
We aim to launch a structural and functional investigation of soluble A28 fusion protein in order to better understand the virus entry mechanism. To this end, we expressed and purified a truncated form of A28 (14 kDa; residues 38–146, abbreviated as tA28 hereinafter), with deletion of its transmembrane domain (residues 1–22) and a hydrophobic segment (residues 23–37). In this communication, we report the 1H, 13C and 15N NMR assignments of tA28 by 3D heteronuclear NMR spectroscopy and the tA28-A26 viral protein–protein interaction by 2D HSQC spectroscopy. The structural and functional study of tA28 is currently undergoing in our lab might help to further advance our knowledge about the viral entry mechanism at the molecular level.
Methods and experiments
Expression and purification of A28 soluble domain
The cDNA corresponding to soluble tA28 (residues 38–146) tagged with His6-yeast SUMO (Smt3) at the N-terminus was constructed. The tA28 construct was cloned into pETDuet-1 vector and expressed in E. coli BL21 (DE3) cells via a T7 promoter. The U-15N and 13C labeled protein was expressed in M9 minimal media, containing 1 g/L 15NH4Cl (Sigma-Aldrich) and 1 g/L U-13C6 glucose (Cambridge Isotope laboratories) as the sole nitrogen and carbon source, respectively, supplemented with 2 mM [0.24 g/L] MgSO4, 0.1 mM [11.1 mg/L] CaCl2, 1 mg/L thiamin, 1 mg/L biotin. The cells were first grown overnight in LB medium and transferred into M9 medium after removal of excess LB medium. The protein expression was induced by addition of 1 mM isopropyl-β-d-thiogalactoside (IPTG) when the OD600 of the cells reached 0.6. Cells were incubating for 18 h at 17 °C and then harvested by centrifugation at 8000 rpm, 25 min, 4 °C. The cell pellets were suspended in lysis buffer A (20 mM Tris, 300 mM NaCl, 8 M Urea, 1 mM Phenylmethylsulfonyl fluoride (PMSF), 1 mM 1,4-Dithiothreitol (DTT), lysozyme, pH 8.0) and were lysed by sonication on ice bath. The lysate was cleared from cell debris by centrifugation (1.5 h, 4500 rpm at 4 °C). The supernatant was incubated with Ni2+-NTA affinity resin (BioRad) using a 5 ml HisTrap FF column (GE Healthcare Life Sciences) with a gradient from 100% buffer A to 100% buffer B (20 mM Tris, 300 mM NaCl, 20 mM Imidazole, 1 mM PMSF, 1 mM DTT, pH 8.0). The protein was eluted with buffer C containing 500 mM imidazole, 20 mM Tris, 300 mM NaCl, 1 mM DTT, pH 8.0. The His6-SUMO-tag was cleaved by Ulp1 protease (Reverter and Lima 2009) during incubation with buffer D (20 mM Tris, 100 mM NaCl, 2 mM CaCl2, pH 8.0) and then purified using HiLoad 16/60 Superdex 75 size-exclusion chromatography column (GE Healthcare) in buffer E (20 mM Tris, 150 mM NaCl, 2 mM DTT, 0.02% NaN3, pH 8.0) and then concentrated to 1.0 mM using Amicon Ultra Centrifugal Filter unit with a 3 kDa molecular weight cut-off (Millipore). The concentrated protein was exchanged into NMR buffer containing 20 mM MES (pH 6.5), 50 mM NaCl. The protein concentration was determined by absorbance at 280 nm (ε280 = 29,160 M−1 cm−1) and purity of the protein was confirmed by 15% SDS–PAGE.
NMR spectroscopy
NMR samples containing 10% v/v D2O and 4,4-dimethyl-4-silapentane-1-sulfonic acid (DSS) as an internal reference were loaded into 5 mm Shigemi NMR tubes. All NMR experiments were performed at 298 K and pH 6.5 on Bruker AVANCE 600 or 800 MHz spectrometers equipped with 5 mm triple resonance TXI cryogenic probes including shielded Z-gradient. Sequence-specific backbone resonance assignments were achieved by the independent connectivity analysis of HNCACB, CBCA(CO)NH, HNCO and HN(CA)CO. The aliphatic side chain assignments were determined using aliphatic HCCH-TOCSY, C(CO)NH, H(CCO)NH, HBHA(CO)NH and 1H–15N NOESY-HSQC (mixing time 150 ms). Aromatic side chain resonances were assigned using HBCBCGCDHD, HBCBCGCDCEHE, 13C-HSQC, and 1H–13C NOESY-HSQC (mixing time 150 ms). All NMR spectra were processed using Bruker Topspin 3.6 and analyzed using NMRView. DSS was used for the 1H, 13C and 15N chemical shift references, using appropriate conversion equations (Wishart et al. 1995). Secondary structure analysis was performed using backbone chemical shifts (13Cα, 13C’, 13Cβ, 15N, 1Hα and 1HN) by Talos+ webserver (Shen et al. 2009).
Assignments and data deposition
The 2D 1H–15N HSQC spectrum of tA28 (residues 38–146) showed well resolved resonances for non-proline residues, indicative of a well-folded structure (Fig. 1). We have completed 98.5% of backbone assignment among which 101 out of 104 non-proline residues with exceptions of Ala38, Thr39 and Asn127. All NƐ1 and HƐ1 resonances of 17 amide side-chain of Asn and Gln were assigned and indicated in Fig. 1. Overall, 93.5% of all backbone and side chain resonances were assigned. For aromatic residues, Phe43, Trp73, Phe89, Phe106, Phe115 and Phe141 were partially assigned. Secondary structure analysis using 13Cα, 13C’, 13Cβ, 15N, 1Hα and 1HN chemical shifts reveal two α-helices, α1: residues 109–116, α2: 137–144 and six β-strands, β1: residues 74–77, β2: 82–87, β3: 93–95, β4: 105–106, β5: 129–130 and β6: 134–135 (Fig. 2).
2D 1H–15N HSQC spectrum of 13C/15N-labeled tA28 in 20 mM MES (pH 6.5), 50 mM NaCl and 10% D2O, in the presence (blue) and absence of A26-C (residues 351–500) (black), recorded at 298 K on a Bruker 600 MHz AVANCE spectrometer equipped with a TXI cryogenic probe. In the presence of the A26-C, for those residues that were involved in the tA28/A26 interaction, their cross-peaks were missing from the tA28 HSQC spectrum, including 40–52, 56–65, 70–73, 89–93, 118–119, 121–122 and 124–126 residues. Thus, it was suggested that these residues were directly involved in the viral protein–protein interaction. The assigned residues are indicated using single letter codes. An expended section of the central, overlapped region is shown by red frame. Assignments of the side-chain NH2 groups from the Asn and Gln residues are indicated with an asterisk, and the pairs of protons are connected by horizontal gray lines. Resonance assignments are available online at the BMRB repository (Accession Number 50469)
Secondary structures prediction of tA28 from TALOS+ derived from 13Cα, 13C’, 13Cβ, 15N, 1Hα and 1HN chemical shifts. The secondary structure of α-helix, β-strand and random coil are represented by gray cylinders, black arrows and black lines, respectively. The residue with no NMR assignment was marked with X
To verify whether A28 interacts with fusion suppressor A26 (full length; residues 1–500) mediating virus entry, we then mixed 15N-labeled tA28 with natural abundant N- and C-terminal A26 fragments (residues 1–397; A26-N & residues 351–500; A26-C), at a molar ratio 1:0.7, and examined by 2D 1H–15N HSQC spectroscopy, respectively. As shown, for those residues that were involved in the tA28/A26 interaction, their cross-peaks were missing from the tA28 HSQC spectrum when A26-C was added (see Fig. 1), including 40–52, 56–65, 70–73, 89–93, 118–119, 121–122 and 124–126 residues. No change in the tA28 HSQC spectrum was observed when A26-N was added (data not shown). Our 2D HSQC data indicated that tA28 does interact specifically with A26-C (at pH 6.5), instead of A26-N. Thus, it is interpreted that the tA28/A26 protein–protein interaction might regulate the EFC membrane fusion activity.
The determination of the molecular structure of tA28 using 13C- and 15N-edited NOESY is underway. The 1H, 13C and 15N chemical shifts have been deposited into the Biological Magnetic Resonance Databank (http://www.bmrb.wisc.edu) under the ascension code BMRB-50469.
References
Armstrong JA, Metz DH, Young MR (1973) The mode of entry of vaccinia virus into L cells. J Gen Virol 21(3):533–537. https://doi.org/10.1099/0022-1317-21-3-533
Carter GC, Law M, Hollinshead M, Smith GL (2005) Entry of the vaccinia virus intracellular mature virion and its interactions with glycosaminoglycans. J Gen Virol 86(Pt 5):1279–1290. https://doi.org/10.1099/vir.0.80831-0
Chang A, Metz DH (1976) Further investigations on the mode of entry of vaccinia virus into cells. J Gen Virol 32(2):275–282. https://doi.org/10.1099/0022-1317-32-2-275
Chang SJ, Shih AC, Tang YL, Chang W (2012) Vaccinia mature virus fusion regulator A26 protein binds to A16 and G9 proteins of the viral entry fusion complex and dissociates from mature virions at low pH. J Virol 86(7):3809–3818. https://doi.org/10.1128/JVI.06081-11
Chang H-W, Yang C-H, Luo Y-C, Su B-G, Cheng H-Y, Tung S-Y, Carillo KJD, Liao Y-T, Tzou D-LM, Wang H-C, Chang W (2019) Vaccinia viral A26 protein is a fusion suppressor of mature virus and triggers membrane fusion through conformational change at low pH. PLOS Pathog 15(6):e1007826. https://doi.org/10.1371/journal.ppat.1007826
Goebel SJ, Johnson GP, Perkus ME, Davis SW, Winslow JP, Paoletti E (1990) The complete DNA sequence of vaccinia virus. Virology 179(1):247–266. https://doi.org/10.1016/0042-6822(90)90294-2
Moss B (1996) Poxviridae: the viruses and thier replication. In: Fields BN, Knipe DM, Howley PM (eds) Fields virology. Lippincott - Raven Publishers, pp 2637–2671. https://ci.nii.ac.jp/naid/10004889159/en/
Reverter D, Lima CD (2009) Preparation of SUMO proteases and kinetic analysis using endogenous substrates. Methods Mol Biol 497:225–239. https://doi.org/10.1007/978-1-59745-566-4_15
Senkevich TG, Ward BM, Moss B (2004) Vaccinia virus A28L gene encodes an essential protein component of the virion membrane with intramolecular disulfide bonds formed by the viral cytoplasmic redox pathway. J Virol 78(5):2348–2356. https://doi.org/10.1128/JVI.78.5.2348-2356.2004
Senkevich TG, Ojeda S, Townsley A, Nelson GE, Moss B (2005) Poxvirus multiprotein entry–fusion complex. PNAS 102(51):18572. https://doi.org/10.1073/pnas.0509239102
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44(4):213–223. https://doi.org/10.1007/s10858-009-9333-z
Wagenaar TR, Ojeda S, Moss B (2008) Vaccinia virus A56/K2 fusion regulatory protein interacts with the A16 and G9 subunits of the entry fusion complex. J Virol 82(11):5153–5160. https://doi.org/10.1128/JVI.00162-08
Wishart DS, Bigam CG, Yao J, Abildgaard F, Dyson HJ, Oldfield E, Markley JL, Sykes BD (1995) 1H, 13C and 15N chemical shift referencing in biomolecular NMR. J Biomol NMR 6(2):135–140. https://doi.org/10.1007/BF00211777
Wolfe CL, Moss B (2011) Interaction between the G3 and L5 proteins of the vaccinia virus entry-fusion complex. Virology 412(2):278–283. https://doi.org/10.1016/j.virol.2011.01.014
Acknowledgements
We gratefully thank Claire Yang (Institute of Chemistry, Academia Sinica) for help in preparing the manuscript. NMR spectra were collected at the High-field NMR Center (HFNMRC), Academia Sinica, supported by Academia Sinica Core Facility and Innovative Instrument Project (AS-CFII-108-112). This research is funded by Grant MOST 106-2113-M-001-025 and 108-2113-M-001-016 (DLT) from the Ministry of Science and Technology of Taiwan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wu, D., Lou, YC., Chang, W. et al. NMR assignments of vaccinia virus protein A28: an entry-fusion complex component. Biomol NMR Assign 15, 117–120 (2021). https://doi.org/10.1007/s12104-020-09993-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-020-09993-0