Abstract
The pneumococcal serine rich repeat protein (PsrP) is displayed on the surface of Streptococcus pneumoniae with a suggested role in colonization in the human upper respiratory tract. Full-length PsrP is a 4000 residue-long multi-domain protein comprising a positively charged functional binding region (BR) domain for interaction with keratin and extracellular DNA during pneumococcal adhesion and biofilm formation, respectively. The previously determined crystal structure of the BR domain revealed a flat compressed barrel comprising two sides with an extended β-sheet on one side, and another β-sheet that is distorted by loops and β-turns on the other side. Crystallographic B-factors indicated a relatively high mobility of loop regions that were hypothesized to be important for binding. Furthermore, the crystal structure revealed an inter-molecular β-sheet formed between edge strands of two symmetry-related molecules, which could promote bacterial aggregation during biofilm formation. Here we report the near complete 15N/13C/1H backbone resonance assignment of the BR domain of PsrP, revealing a secondary structure profile that is almost identical to the X-ray structure. Dynamic 15N-T1, T2 and NOE data suggest a monomeric and rigid structure of BR with disordered residues only at the N- and C-termini. The presented peak assignment will allow us to identify BR residues that are crucial for ligand binding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Biological context
The Gram-positive commensal and human-adapted bacterium Streptococcus pneumoniae colonizes the upper respiratory tract in about 10% of healthy adults and up to 60% of children, without necessarily causing any symptoms (van der Poll and Opal 2009). However, upon certain not yet well-defined triggers, pneumococcus is transformed from a silent colonizer to a virulent pathogen causing diseases such as otitis media, sinusitis, pneumonia, septicemia and meningitis (Weiser et al. 2018). For efficient colonization within the nasopharynx, pneumococcus displays a multitude of surface-exposed modular adhesive and catalytic proteins (Perez-Dorado et al. 2012).
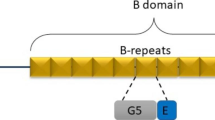
One such adhesin is the giant pneumococcal serine rich repeat protein (PsrP), that was discovered as a key colonization factor present in 60% of pneumococcal strains capable of causing pneumonia in children (Blanchette-Cain et al. 2013; Sanchez et al. 2010). PsrP belongs to a family of serine rich repeat proteins (SRRP) displayed on the surface of Gram-positive bacteria for bacterial attachment to host cells (Lizcano et al. 2012). The C-terminal LPXTG motif of PsrP is recognized by Sortases and covalently linked to the bacterial peptidoglycan cell wall. SRRPs share a long, highly repetitive and glycosylated C-terminal serine rich-repeat (SRR) region that varies in length between 400 and 4000 residues. Their functional binding region (BR) domains, which bind to a broad range of targets including extracellular DNA (eDNA), glyco-conjugates and keratins, are variable in sequence and organized into modular domains (Lizcano et al. 2012).
The positively charged BR domain of PsrP binds to negatively charged helical structures such as keratin-10 (KRT-10) and eDNA, possibly to promote efficient bacterial attachment to the upper respiratory tract and during biofilm formation, respectively (Sanchez et al. 2010; Shivshankar et al. 2009; Blanchette-Cain et al. 2013; Schulte et al. 2014, 2016). The crystal structure of the KRT10- and DNA-binding domain of PsrP (BR187–385) revealed a fold topology that is distantly related to the adhesin CnaA of S. aureus, a microbial surface component recognizing adhesive molecule (MSCRAMM) (Deivanayagam et al. 2002; Schulte et al. 2014) (Fig. 1a). MSCRAMMs were defined by a common ligand binding mechanism that is mediated by two adjacent subdomains comprising Ig-like folds (Foster et al. 2014). In the structurally and mechanistically well-described ‘dock, lock and latch’ (DLL) binding mode, extracellular matrix-derived peptide ligands dock to the open apo form of MSCRAMMs and conformational changes create a closed form, in which the ligands are locked into place (Foster et al. 2014). Most of the described DLL and associated binding mechanisms were derived from X-ray structures of apo- and ligand-bound forms of MSCRAMMs, and in-depth biophysical investigations revealed strong interactions even withstanding forces in the covalent bond range (Deivanayagam et al. 2002; Xiang et al. 2012; Ross et al. 2012; Milles et al. 2018; Herman et al. 2014).
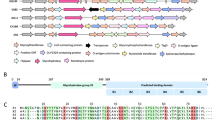
Near complete 15N/13C/1H backbone resonance assignment of BR187–385 revealed a rigid monomer with secondary structure almost identical to the X-ray structure (PDB:3ZGH). a The crystal structure of BR187–385 is visualized as in Fig. 3 of our previous publication (Schulte et al. 2014). b Assigned non-proline backbone 15N and 1HN as well as 13Cα are shown in green. Non-assigned residues are shown in black. Proline residues are highlighted (P). c The secondary structure of BR187–385 calculated from the NMR chemical shifts was compared to the crystal structure at the residue-level, and color-coded according to the cartoon representation in panel a. Prior to the secondary structure calculations, random coil chemical shifts of BR187–385, were calculated using POTENCI (results shown in Figure S1) (Nielsen and Mulder 2018; Schwarzinger et al. 2000). d T2 relaxation times, 15N-NOEs data as well as the general order parameter S2 reveal a rigid barrel with flexible N- and C-termini. Panels b–d are visualized on a common axis corresponding to the residue numbers of BR187–385. T1 relaxation times are shown in Figure S2
While we have presented structural docking models for binding of KRT-10 or eDNA to the BR domain of PsrP, neither their complex structures nor defined molecular binding mechanisms have been demonstrated. It is also yet unclear whether the previously identified inter-molecular β-sheet dimer of BR is required for ligand binding. In this report we present the complete 1H, 13C and 15N backbone assignment of the PsrP-BR domain that will allow us to identify BR residues that are crucial for ligand binding and involved in binding-associated conformational dynamics. Our on-going ligand titration experiments will reveal if BR adopts a binding mode similar to or different from the DLL mode of MSCRAMMs.
Methods and experiments
Protein production
A TEV-cleavage site (ENLYFQG) was inserted between the poly-His tag and the BR domain of the previously described BR187−385 construct using sequence and ligation-independent cloning to yield His6-TEV-BR187−385 (Li and Elledge 2007; Schulte et al. 2014). The protein was expressed heterologously in E. coli and purified as previously described (Schulte et al. 2014). Single 15N and double 15N/13C labeled proteins were expressed in minimal M9 medium according to published protocols (15NH4Cl: 1 g/L and 13C d-glucose: 10 g/L) (Sivashanmugam et al. 2009; Fox and Blommel 2009). Bacteria were grown to an OD of about 0.4–0.7 in 4L TB medium, pelleted and re-suspended in 400 mL of minimal NMR-growth medium for o/n expression at 25 °C.
All purification buffers were based on 20 mM HEPES, 300 mM NaCl, 10% glycerol pH 7.5. His6-TEV-BR187−385 was purified using immobilized metal affinity (IMAC, HisTrap FF GE-Healthcare) and size exclusion chromatography (SEC, Superdex 75, GE Healthcare). The poly-His tag was removed by TEV cleavage in HEPES buffer comprising 5 mM EDTA and 1 mM DTT. Cleaved protein was passed through the Ni-NTA column as flow-through. The final sample was purified over Superdex 75 column in 50 mM sodium-phosphate buffer pH 5.0, 100 mM NaCl, and concentrated using centrifugation ultrafiltration.
NMR samples preparation
All NMR experiments were performed in the buffer containing 50 mM Na–phosphate pH 5.0, 100 mM NaCl, with added 1 mM NaN3 and 10 (v/v) % D2O. The buffer-exchanged protein was concentrated to at least 0.7 mM using centrifugation ultrafiltration and 280 μL was placed in a 5 mm shigemi tube.
NMR spectroscopy
NMR experiments were acquired on Bruker Avance III spectrometers operating at 14.1 T, equipped with a cryo-enhanced QCI-P probe at a temperature of 298 K. The backbone residues were assigned, based on standard 3D TROSY triple resonance experiments. The iterative non-uniform sampling protocol (NUS) comprised HNCO, HNCA and HN(CO)CA, HN(CA)CO, HN(CO)CACB and HNCACB experiments (Jaravine et al. 2008; Orekhov and Jaravine 2011; Salzmann et al. 1998, 1999). A 25% sampling schedule was used for all other 3D spectra, yielding a total acquisition time of 153 h (about 1 week). Targeted acquisition (TA) was used for automatic processing and analysis of spectra as described previously (Jaravine and Orekhov 2006; Jaravine et al. 2008; Orekhov and Jaravine 2011). This novel procedure reduces significantly data acquisition and analysis time to assign backbone resonance peaks of proteins (Unnerstale et al. 2016; Agback et al. 2019). The automatic assignment was validated manually using CcpNmr Analysis 2.4.2 (Vranken et al. 2005).
Hα protons were assigned using a 3D HCACO sampling schedule comprising 25% NUS and 15N-hsqc-NOESY, 13C-hsqc-NOESY (Kay et al. 1990, 1992; Schleucher et al. 1994). Data were processed and assigned by Topspin 4.0.6 (Bruker) and CcpNmr Analysis 2.4.2, respectively (Vranken et al. 2005). The 1H, 13C and 15N backbone chemical shifts were referred to DSS-d6 directly, while 13C and 15N chemical shifts were referred to indirectly.
Random coil chemical shifts of BR187–385, were calculated using POTENCI with neighbour correction and subtracted from the experimental 1HN, 15N, 13Cα, 13Cβ, 13C′ and Hα chemical shifts (Nielsen and Mulder 2018; Schwarzinger et al. 2000) (Figure S1). The chemical shift index (CSI) was calculated according to the original method (Wishart et al. 1992). Residues with consecutive Δδ13C′ or Δδ13Cα values above 0.7 ppm and below − 0.7 ppm indicate alpha helix, and beta strands, respectively. The opposite is valid for Δδ13Cβ. The CSI for the three nuclei were averaged and reported as “consensus” CSI.
T1, T2 and NOEs were determined using sensitivity enhanced TROSY-type pulse-sequences with temperature compensation train of pulses after acquisition time (Zhu et al. 2000). T1 relaxation was determined from the following series of relaxation delays: 10, 90, 192, 320, 480, 690, 980, 1220 and 1444 ms. T2 relaxation was measured using CPMG delays of 8.5, 17.0, 25.4, 33.9, 42.4, 50.9, 59.4, 76.3 and 93.3 ms. Both T1 and T2 experiments were repeated to estimate the experimental fitting to about 2%. The same error was set for the NOE experiment. All spectra were processed by Topspin 4.0.6 (Bruker) and evaluated using Dynamics Center 2.1 (Bruker), in which T1 and T2 data were fit using mono exponential decay functions. NOEs were obtained by dividing the 1H–15N peak intensities in a NOE-enhanced spectrum by the corresponding intensities in an unsaturated spectrum. The order parameters, S2, and the fast internal correlation time, τE were obtained by fitting the relaxation parameters at one field using the Lipari–Szabo model-free approach with a NH bond length of 1.02 Å and a CSA of − 160 ppm (Hiyama et al. 1988; Lipari and Szabo 1982a, b).
In the figures and in the text, the standard nomenclature for amino acids of the carbon atoms was used, where 13Cα is the carbon next to the carbonyl group 13C′ and 13Cβ is the carbon next to 13Cα (Markley et al. 1998). The secondary structures obtained from NMR and X-ray crystallography were compared using CSI 3.0 (Hafsa et al. 2015). The crystal structure and NMR data presented in Fig. 1 were visualized using PyMol and the R tidyverse, respectively (Schrödinger 2010; Wickham 2016, 2017).
Extent of assignments and data deposition
Targeted Acquisition (TA) and conventional approaches were combined to assign 94% of non-proline backbone 15N and 1HN, 98% of 13Cα, 96% of 13Cβ, 96% of 13C′ and 91% of non-glycine Hα (Fig. 1b). All assigned chemical shifts are labelled in Fig. 2 and the associated peak table has been deposited to BMRB with accession code 50157. The secondary structure profile derived from the NMR data was almost identical to the previously determined crystal structure, thus validating our resonance assignment (Fig. 1a, c). In more detail, the previously determined crystal structure of BR187–385 revealed a fold that is distantly related to the DEv-IgG fold topology of MSCRAMMs (Schulte et al. 2014). The DEv-IgG fold topology can be described as a compressed barrel composed of two opposing β-sheets that are formed by β-strands ABED (sheet I) and CFG (sheet II), and is distinguished from the IgG-fold by the insertion of two extra strands between strands D and E (Deivanayagam et al. 2002; Vengadesan and Narayana 2011). In BR187–385, the CFG sheet is heavily distorted by loops and β-turns, and four strands (D1 to D4) are inserted between strands D and E (Fig. 1a). The NMR assignment revealed that residues T353-A356 between strands F1 and F2 adopted a β-strand in solution, thus combining the two short strands into a single F strand comprising nine residues (Fig. 1a, c). Subtle differences between the crystal and NMR secondary structures were noticed for turn and disordered regions following strand F. The two short strands C1 and C2 were each extended by two residues in solution, but a short sequence comprising residues Y267-G270 in between the two strands was not assigned. Furthermore, the short G-strand was identified as unstructured in solution. The N- and C-termini comprising G186-E208 and T359-Q385 respectively, were identified as unstructured.
Dynamic backbone motions of BR187–385 on a pico- to nanosecond timescale were evaluated by determining the longitudinal (15N T1) and transverse (15N T2) relaxation times as well as steady-state heteronuclear nuclear Overhauser enhancement (15N NOE) of each 1H–15N amide bond (Figs. 1 and S2). High T2 relaxation times, large negative 15N-NOE values and low general order parameter S2 values indicated highly dynamic N- and C-termini of BR187–385 (Fig. 1d). Indeed, these regions were not observed in the previously determined crystal structure. However, low T2 relaxation times, positive 15N-NOE values as well as S2-values between 0.8 and 1 revealed a rigid structure of the compressed BR187–385 barrel (Fig. 1). Furthermore, the correlation time for molecular reorientation (τc) was estimated to 13 ns as expected for a 20 kDa protein, indicating that in solution BR187–385 is a monomer.
In conclusion, the near complete 15N/13C/1H backbone resonance assignment of BR187–385 revealed a secondary structure profile almost identical to the X-ray structure. BR187–385 was monomeric and rigid in solution exhibiting disordered flexible N- and C-termini. Studies of the structure and dynamics of the BR187–385 in complex with ligands in solution are on-going and will provide important insights in the molecular bases underlying these interactions.
References
Agback P et al (2019) Structural characterization and biological function of bivalent binding of CD2AP to intrinsically disordered domain of chikungunya virus nsP3 protein. Virology 537:130–142. https://doi.org/10.1016/j.virol.2019.08.022
Blanchette-Cain K et al (2013) Streptococcus pneumoniae biofilm formation is strain dependent, multifactorial, and associated with reduced invasiveness and immunoreactivity during colonization. mBio. https://doi.org/10.1128/mBio.00745-13
Deivanayagam CCS, Wann ER, Chen W, Carson M, Rajashankar KR, Hook M, Narayana SVL (2002) A novel variant of the immunoglobulin fold in surface adhesins of Staphylococcus aureus: crystal structure of the fibrinogen-binding MSCRAMM, clumping factor A. EMBO J 21:6660–6672. https://doi.org/10.1093/emboj/cdf619
Foster TJ, Geoghegan JA, Ganesh VK, Hoeoek M (2014) Adhesion, invasion and evasion: the many functions of the surface proteins of Staphylococcus aureus. Nat Rev Microbiol 12:49–62. https://doi.org/10.1038/nrmicro3161
Fox BG, Blommel PG (2009) Autoinduction of protein expression current protocols in protein science CHAPTER 5:Unit-5.23. https://doi.org/10.1002/0471140864.ps0523s56
Hafsa NE, Arndt D, Wishart DS (2015) CSI 3.0: a web server for identifying secondary and super-secondary structure in proteins using NMR chemical shifts. Nucleic Acids Res 43:W370–W377. https://doi.org/10.1093/nar/gkv494
Herman P, El-Kirat-Chatel S, Beaussart A, Geoghegan JA, Foster TJ, Dufrêne YF (2014) The binding force of the staphylococcal adhesin SdrG is remarkably strong. Mol Microbiol 93:356–368. https://doi.org/10.1111/mmi.12663
Hiyama Y, Niu CH, Silverton JV, Bavoso A, Torchia DA (1988) Determination of N-15 chemical-shift tensor via N-15-H-2 dipolar coupling in boc-glycylglycyl[N-15]glycine benzyl ester. J Am Chem Soc 110:2378–2383. https://doi.org/10.1021/ja00216a006
Jaravine VA, Orekhov VY (2006) Targeted acquisition for real-time NMR spectroscopy. J Am Chem Soc 128:13421–13426. https://doi.org/10.1021/ja062146p
Jaravine VA, Zhuravleva AV, Permi P, Ibraghimov I, Orekhov VY (2008) Hyperdimensional NMR spectroscopy with nonlinear sampling. J Am Chem Soc 130:3927–3936. https://doi.org/10.1021/ja077282o
Kay LE, Ikura M, Tschudin R, Bax A (1990) 3-Dimensional triple-resonance nmr-spectroscopy of isotopically enriched proteins. J Magn Reson 89:496–514. https://doi.org/10.1016/0022-2364(90)90333-5
Kay LE, Keifer P, Saarinen T (1992) Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J Am Chem Soc 114:10663–10665. https://doi.org/10.1021/ja00052a088
Li MZ, Elledge SJ (2007) Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC. Nat Methods 4:251–256. https://doi.org/10.1038/nmeth1010
Lipari G, Szabo A (1982a) Model-free approach to the interpretation of nuclear magnetic-resonance relaxation in macromolecules. 1. Theory and range of validity. J Am Chem Soc 104:4546–4559. https://doi.org/10.1021/ja00381a009
Lipari G, Szabo A (1982b) Model-free approach to the interpretation of nuclear magnetic-resonance relaxation in macromolecules. 2. Analysis of experimental results. J Am Chem Soc 104:4559–4570. https://doi.org/10.1021/ja00381a010
Lizcano A, Sanchez CJ, Orihuela CJ (2012) A role for glycosylated serine-rich repeat proteins in Gram-positive bacterial pathogenesis. Mol Oral Microbiol 27:257–269. https://doi.org/10.1111/j.2041-1014.2012.00653.x
Markley JL et al (1998) Recommendations for the presentation of NMR structures of proteins and nucleic acids—IUPAC-IUBMB-IUPAB Inter-Union Task Group on the standardization of data bases of protein and nucleic acid structures determined by NMR spectroscopy. Eur J Biochem 256:1–15. https://doi.org/10.1046/j.1432-1327.1998.2560001.x
Milles LF, Schulten K, Gaub HE, Bernardi RC (2018) Molecular mechanism of extreme mechanostability in a pathogen adhesin. Science 359:1527–1533. https://doi.org/10.1126/science.aar2094
Nielsen JT, Mulder FAA (2018) POTENCI: prediction of temperature, neighbor and pH-corrected chemical shifts for intrinsically disordered proteins. J Biomol NMR 70:141–165. https://doi.org/10.1007/s10858-018-0166-5
Orekhov V, Jaravine VA (2011) Analysis of non-uniformly sampled spectra with multi-dimensional decomposition. Prog Nucl Magn Reson Spectrosc 59:271–292. https://doi.org/10.1021/ja062146p
Perez-Dorado I, Galan-Bartual S, Hermoso JA (2012) Pneumococcal surface proteins: when the whole is greater than the sum of its parts. Mol Oral Microbiol 27:221–245. https://doi.org/10.1111/j.2041-1014.2012.00655.x
Ross CL, Liang X, Liu Q, Murray BE, Höök M, Ganesh VK (2012) Targeted protein engineering provides insights into binding mechanism and affinities of bacterial collagen adhesins. J Biol Chem 287:34856–34865. https://doi.org/10.1074/jbc.M112.371054
Salzmann M, Pervushin K, Wider G, Senn H, Wuthrich K (1998) TROSY in triple-resonance experiments: new perspectives for sequential NMR assignment of large proteins. Proc Natl Acad Sci USA 95:13585–13590. https://doi.org/10.1073/pnas.95.23.13585
Salzmann M, Wider G, Pervushin K, Senn H, Wuthrich K (1999) TROSY-type triple-resonance experiments for sequential NMR assignments of large proteins. J Am Chem Soc 121:844–848
Sanchez CJ et al (2010) The pneumococcal serine-rich repeat protein is an intra-species bacterial adhesin that promotes bacterial aggregation in vivo and in biofilms. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1001044
Schleucher J et al (1994) A general enhancement scheme in heteronuclear multidimensional NMR employing pulsed field gradients. J Biomol NMR 4:301–306. https://doi.org/10.1007/bf00175254
Schrödinger L (2010) PyMOL molecular graphics system, version 2.0 Schrödinger, LLC
Schulte T et al (2014) The basic keratin 10-binding domain of the virulence-associated pneumococcal serine-rich protein PsrP adopts a novel MSCRAMM fold. Open Biol 4:130090. https://doi.org/10.1098/rsob.130090
Schulte T et al (2016) The BR domain of PsrP interacts with extracellular DNA to promote bacterial aggregation; structural insights into pneumococcal biofilm formation. Sci Rep 6:32371. https://doi.org/10.1038/srep32371
Schwarzinger S, Kroon GJ, Foss TR, Wright PE, Dyson HJ (2000) Random coil chemical shifts in acidic 8 M urea: implementation of random coil shift data in NMRView. J Biomol NMR 18:43–48. https://doi.org/10.1023/a:1008386816521
Shivshankar P, Sanchez C, Rose LF, Orihuela CJ (2009) The Streptococcus pneumoniae adhesin PsrP binds to Keratin 10 on lung cells. Mol Microbiol 73:663–679. https://doi.org/10.1111/j.1365-2958.2009.06796.x
Sivashanmugam A, Murray V, Cui C, Zhang Y, Wang J, Li Q (2009) Practical protocols for production of very high yields of recombinant proteins using Escherichia coli. Protein Sci 18:936–948. https://doi.org/10.1002/pro.102
Unnerstale S et al (2016) Backbone assignment of the MALT1 paracaspase by solution NMR. PLoS ONE. https://doi.org/10.1371/journal.pone.0146496
van der Poll T, Opal SM (2009) Pathogenesis, treatment, and prevention of pneumococcal pneumonia. Lancet 374:1543–1556. https://doi.org/10.1016/S0140-6736(09)61114-4
Weiser JN, Ferreira DM, Paton JC (2018) Streptococcus pneumoniae: transmission, colonization and invasion. Nat Rev Microbiol. https://doi.org/10.1038/s41579-018-0001-8
Vengadesan K, Narayana SVL (2011) Structural biology of Gram-positive bacterial adhesins. Protein Sci 20:759–772. https://doi.org/10.1002/pro.613
Wickham H (2016) ggplot2: elegant graphics for data analysis, vol 2nd. Springer International Publishing, New York
Wickham H (2017) Tidyverse: R-packages for data science. tidyverse.org
Wishart DS, Sykes BD, Richards FM (1992) The chemical shift index: a fast and simple method for the assignment of protein secondary structure through. NMR Spectrosc Biochem 31:1647–1651. https://doi.org/10.1021/bi00121a010
Vranken WF et al (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696. https://doi.org/10.1002/prot.20449
Xiang H et al (2012) Crystal structures reveal the multi-ligand binding mechanism of Staphylococcus aureus ClfB. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1002751
Zhu G, Xia Y, Nicholson LK, Sze KH (2000) Protein dynamics measurements by TROSY-based NMR experiments. J Magn Reson 143:423–426. https://doi.org/10.1006/jmre.2000.2022
Acknowledgements
Open access funding provided by Swedish University of Agricultural Sciences. This work was supported by Swedish Foundation for Strategic Research grant ITM17-0218 to P.A.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schulte, T., Sala, B.M., Nilvebrant, J. et al. Assigned NMR backbone resonances of the ligand-binding region domain of the pneumococcal serine-rich repeat protein (PsrP-BR) reveal a rigid monomer in solution. Biomol NMR Assign 14, 195–200 (2020). https://doi.org/10.1007/s12104-020-09944-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-020-09944-9