Abstract
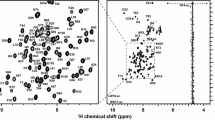
Nucleosome core particle (NCP), the basic unit of chromatin in eukaryotic cells, consists of ~ 147 bp DNA wrapped around a histone octamer (HO) formed by two H2A–H2B dimers and one (H3–H4)2 tetramer. Histones undergo various post-translational modifications (PTMs), which regulates genomic activities in different cellular phases. High-resolution structures have been solved for many nucleosomes primarily including NCPs. However, the atomic-resolution structures of nucleosome arrays and chromatin fiber, as well as the dynamics of nucleosomes remain poorly understood. Solid-state NMR (SSNMR) is one of the premier techniques to answer these questions. In this study, we present the 13C and 15N chemical shifts assignments for the globular domain of human histone H3 (hH3) using multidimensional SSNMR experiments. The obtained spectra are of outstanding resolution and the assignments are nearly 100% complete for the backbone 13C and 15N spins of R42–G132 and ~ 80% when taking into account the side chains. The secondary structure derived from the chemical shifts agrees with the previously reported X-ray crystal structure. The reported chemical shifts can be carried over to future SSNMR studies of structure and dynamics of hH3 in NCPs, nucleosome array, chromatin fibers and nucleosome-protein complexes.




Similar content being viewed by others
References
Baxevanis AD, Landsman D (1996) Histone sequence database: a compilation of highly-conserved nucleoprotein sequences. Nucleic Acids Res 24:245–247. https://doi.org/10.1093/nar/24.1.245
Berezhnoy NV, Liu Y, Allahverdi A, Yang R, Su CJ, Liu CF, Korolev N, Nordenskiöld L (2016) The influence of ionic environment and histone tails on columnar order of nucleosome core particles. Biophys J 110:1720–1731. https://doi.org/10.1016/j.bpj.2016.03.016
Bertin A, Mangenot S, Renouard M, Durand D, Livolant F (2007) Structure and phase diagram of nucleosome core particles aggregated by multivalent cations. Biophys J 93:3562–3663. https://doi.org/10.1529/biophysj.107.108365
Bohrmann B, Haider M, Kellenberger E (1993) Concentration evaluation of chromatin in unstained resin-embedded sections by means of low-dose ratio-contrast imaging in STEM. Ultramicroscopy 49:235–251. https://doi.org/10.1016/j.bpj.2016.03.016
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293. https://doi.org/10.1007/BF00197809
Felsenfeld G, Groudine M (2003) Controlling the double helix. Nature 424:448–453. https://doi.org/10.1038/nature01410
Gao M, Nadaud PS, Bernier MW, North JA, Hammel PC, Poirier MG, Jaroniec CP (2013) Histone H3 and H4 N-terminal tails in nucleosome arrays at cellular concentrations probed by magic angle spinning NMR spectroscopy. J Am Chem Soc 135:15278–15281. https://doi.org/10.1021/ja407526s
Jones B, Su H, Bhat A, Lei H, Bajko J, Hevi S, Baltus GA, Kadam S, Zhai H, Valdez R, Gonzalo S, Zhang Y, Li E, Chen T (2008) The histone H3K79 methyltransferase Dot1L is essential for mammalian development and heterochromatin structure. PLoS Genet 4:e1000190. https://doi.org/10.1371/journal.pgen.1000190
Kato H, van Ingen H, Zhou B, Feng H, Bustin M, Kay LE, Bai Y (2011) Architecture of the high mobility group nucleosomal protein 2-nucleosome complex as revealed by methyl-based NMR. Proc Natl Acad Sci USA 108:12283–12288. https://doi.org/10.1073/pnas.1105848108
Lawrence M, Daujat S, Schneider R (2016) Lateral thinking: how histone modifications regulate gene expression. Trends Genet 32:42–56. https://doi.org/10.1016/j.tig.2015.10.007
Li G, Levitus M, Bustamante C, Widom J (2005) Rapid spontaneous accessibility of nucleosomal DNA. Nat Struct Mol Biol 12:46–53. https://doi.org/10.1038/nsmb869
Lowary PT, Widom J (1998) New DNA sequence rules for high affinity bindingto histone octamer and sequence-directed nucleosome positioning. J Mol Biol 276:19–42. https://doi.org/10.1006/jmbi.1997.1494
Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ (1997) Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389:251–260. https://doi.org/10.1038/38444
Luger K, Rechsteiner TJ, Richmond TJ (1999) Expression and purification of recombinant histones and nucleosome reconstitution. Methods Mol Biol 119:1–16. https://doi.org/10.1385/1-59259-681-9:1
Masumoto H, Hawke D, Kobayashi R, Verreault A (2005) A role for cell-cycle-regulated histone H3 lysine 56 acetylation in the DNA damage response. Nature 436:294–298. https://doi.org/10.1038/nature03714
Ng HH, Feng Q, Wang H, Erdjument-Bromage H, Tempst P, Zhang Y, Struhl K (2002) Lysine methylation within the globular domain of histone H3 by Dot1 is important for telomeric silencing and Sir protein association. Genes Dev 16:1518–1527. https://doi.org/10.1101/gad.1001502
Schanda P, Ernst M (2016) Studying dynamics by magic-angle spinning solid-state NMR spectroscopy: principles and applications to biomolecules. Prog Nucl Magn Reson Spectrosc 96:1–46. https://doi.org/10.1016/j.pnmrs.2016.02.001
Shen Y, Bax A (2013) Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J Biomol NMR 56:227–241. https://doi.org/10.1007/s10858-013-9741-y
Shi X, Rienstra CM (2016) Site-specific internal motions in GB1 protein microcrystals revealed by 3D 2H–13C–13C solid-state NMR spectroscopy. J Am Chem Soc 138:4105–4119. https://doi.org/10.1021/jacs.5b12974
Shi X, Prasanna C, Nagashima T, Yamazaki T, Pervushin K, Nordenskiöld L (2018) Structure and dynamics in the nucleosome revealed by solid-state NMR. Angew Chem Int Ed 57:9734–9738. https://doi.org/10.1002/anie.201804707
Takegoshi K, Nakamura S, Terao T (2001) 13C–1H dipolar-assisted rotational resonance in magic-angle spinning NMR. Chem Phys Lett 344:631–637. https://doi.org/10.1016/s0009-2614(01)00791-6
Tessarz P, Kouzarides T (2014) Histone core modifications regulating nucleosome structure and dynamics. Nat Rev Mol Cell Biol 15:703–708. https://doi.org/10.1038/nrm3890
Vasudevan D, Chua EYD, Davey CA (2010) Crystal structures of nucleosome core particles containing the ‘601’ strong positioning sequence. J Mol Biol 403:1–10. https://doi.org/10.1016/j.jmb.2010.08.039
Verel R, Baldus M, Ernst M, Meier BH (1998) homonuclear spin-pair filter for solid-state NMR based on adiabatic-passage techniques. Chem Phys Lett 287:421–428. https://doi.org/10.1016/S0009-2614(98)00172-9
Wei S, Falk SJ, Black BE, Lee T (2015) A novel hybrid single molecule approach reveals spontaneous DNA motion in the nucleosome. Nucleic Acids Res 43:e111. https://doi.org/10.1093/nar/gkv549
Xiang SQ, le Paige UB, Horn V, Houben K, Baldus M, van Ingen H (2018) Site specific studies of nucleosome interactions by solid-state NMR spectroscopy. Angew Chem Int Ed 57:4571–4575. https://doi.org/10.1002/anie.201713158
Xu F, Zhang K, Grunstein M (2005) Acetylation in histone H3 globular domain regulates gene expression in yeast. Cell 121:375–385. https://doi.org/10.1016/j.cell.2005.03.011
Zhou K, Gaullier G, Luger K (2019) Nucleosome structure and dynamics are coming of age. Nat Struct Mol Biol 26:3–13. https://doi.org/10.1038/s41594-018-0166-x
Acknowledgements
This work was supported by the Singapore Ministry of Education (MOE) Academic Research Fund (AcRF) Tier 2 (MOE2018-T2-1-112) and Tier 3 Grants (MOE2012-T3-1-001). SSNMR experiments were conducted at the high-field NMR facility at SPMS, Nanyang Technological University. We acknowledge the NTU Institute of Structural Biology facilities for supporting this research.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shi, X., Prasanna, C., Pervushin, K. et al. Solid-state NMR 13C, 15N assignments of human histone H3 in the nucleosome core particle. Biomol NMR Assign 14, 99–104 (2020). https://doi.org/10.1007/s12104-020-09927-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-020-09927-w