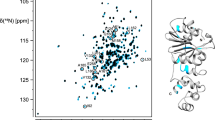
Abstract
Almost complete sequence specific 1H, 13C and 15N resonance assignments of a mutant of UV inducible transcript (S55A-UVI31+) from Chlamydomonas reinhardtii are reported, as a prelude to its structural and functional characterization. Site-directed mutagenesis of uvi31+ was carried out using complementary mutation harbouring oligonucleotides for S55A mutant. The resulting mutant was sequenced and then S55A mutant of uvi31+ cDNA were expressed in E. coli, and purification were carried out from where the protein was purified to high homogeneity. The point mutation S55A in UVI31+ results in a significant enhancement in its DNA endonuclease activity as compared to its wild type protein.


Similar content being viewed by others
References
Aldea M, Hernandez-Chico C, de la Campa AG, Kushner SR, Vicente M (1988) Identification, cloning, and expression of bolA, an ftsZ-dependent morphogene of Escherichia coli. J Bacteriol 170:5169–5176
Atreya HS, Sahu SC, Chary KV, Govil G (2000) A tracked approach for automated NMR assignments in proteins (TATAPRO). J Biomol NMR 17:125–136
Atreya HS, Chary KVR, Govil G (2002) Automated NMR assignments of proteins for high throughput structure determination: TATAPRO II. Curr Sci Bangalore 83(11):1372–1375
Bax A, Grzesiek S (1993) Methodological advances in protein NMR. Acc Chem Res 22:131–138
Bax A, Ikura M, Kay LE, Barbato G, Spera S (1991) Multidimensional triple resonance NMR spectroscopy of isotopically uniformly enriched proteins: a powerful new strategy for structure determination. Ciba Found Symp 161:108–119
Chary KVR, Govil G (2008) NMR in biological systems: from molecules to humans. Springer, The Netherlands
Edison AS, Abildgaard F, Westler WM, Mooberry ES, Markley JL (1994) Practical introduction to theory and implementation of multinuclear, multidimensional nuclear magnetic resonance experiments. Methods Enzymol 239:3–79
Huynen MA, Spronk CA, Gabaldon T, Snel B (2005) Combining data from genomes, Y2H and 3D structure indicates that BolA is a reductase interacting with a glutaredoxin. FEBS Lett 579:591–596
Kasai T, Inoue M, Koshiba S, Yabuki T, Aoki M, Nunokawa E, Seki E, Matsuda T, Matsuda N, Tomo Y (2004) Solution structure of a BolA-like protein from Mus musculus. Protein Sci 13:545
Keller R (2002) The computer aided NMR resonance assignment tutorial. CANTINA, Goldau
Kim SH, Kim M, Lee JK, Kim MJ, Jin YH, Seong RH, Hong SH, Joe CO, Park SD (1997) Identification and expression of uvi31+, a UV-inducible gene from Schizosaccharomyces pombe. Environ Mol Mutagen 30:72–81
Kim MJ, Kim HS, Lee JK, Lee CB, Park SD (2002) Regulation of septation and cytokinesis during resumption of cell division requires uvi31+, a UV-inducible gene of fission yeast. Mol Cells 14:425–430
Lee JK, Park EJ, Chung HK, Hong SH, Joe CO, Park SD (1994) Isolation of UV-inducible transcripts from Schizosaccharomyces pombe. Biochem Biophys Res Commun 202:1113–1119
Moharikar S, D’Souza JS, Kulkarni AB, Rao BJ (2006) Apoptotic-like cell death pathway is induced in unicellular chlorophyte Chlamydomonas reinhardtii cells following UV irradiation: detection and functional analyses. J Phycol 42:423–433
Moharikar S, D’Souza JS, Rao BJ (2007) A homologue of the defender against the apoptotic death gene (dad1) in UV-exposed Chlamydomonas cells is downregulated with the onset of programmed cell death. J Biosci 32:261–270
Montelione GT, Lyons BA, Emerson SD, Tashiro M (1992) An efficient triple resonance experiment using carbon-13 isotopic mixing for determining sequence-specific resonance assignment of isotopically enriched proteins. J Am Chem Soc 113:5490–5492
Muhandiram DR, Farrow NA, Xu GY, Smallcombe SH, Kay LE (1993) A gradient C NOESY-HSQC experiment for recording NOESY spectra of “C-labeled proteins dissolved in H20”. J Magn Reson B 102:317–321
Rout AK, Minda R, Peri D, Ramakrishnan V, Bhattacharjee SK, Rao BJ, Chary KVR (2010) Sequence specific 1H, 13C and 15N backbone resonance assignments of UVI31+ from Chlamydomonas reinhardtii. Biomol NMR Assign 4:171–174
Santos JM, Freire P, Vicente M, Arraiano CM (1999) The stationary-phase morphogene bolA from Escherichia coli is induced by stress during early stages of growth. Mol Microbiol 32:789–798
Shukla M, Minda R, Singh H, Tirumani S, Chary KVR, Rao BJ (2012) UVI31+ is a DNA endonuclease that dynamically localizes to chloroplast pyrenoids in C. reinhardtii. PLoS One 7(12):e51913
Spera S, Bax A (1991) Empirical correlation between protein backbone conformation and Ca and Cb 13C nuclear magnetic resonance chemical shifts. J Am Chem Soc 113:5490–5492
Vieira HL, Freire P, Arraiano CM (2004) Effect of Escherichia coli morphogene bolA on biofilms. Appl Environ Microbiol 70:5682–5684
Wishart DS, Bigam CG, Holm A, Hodges RS, Sykes BD (1995) 1H, 13C, and 15N random coil NMR chemical shifts of the common amino acids. I. Investigations of nearest-neighbour effects. J Biomol NMR 5:67–81
Wuthrich K, Thrich KW, Wuthrich K (1986) NMR of proteins and nucleic acids. Wiley, London
Zhou YB, Cao JB, Wan BB, Wang XR, Ding GH, Zhu H, Yang HM, Wang KS, Zhang X, Han ZG (2008) hBolA, novel non-classical secreted proteins, belonging to different BolA family with functional divergence. Mol Cell Biochem 317:61–68
Acknowledgments
The facilities provided by National Facility for High Field NMR at Tata Institute of Fundamental Research, Mumbai, supported by Department of Science and Technology, New Delhi, Department of Biotechnology, New Delhi, Council of Scientific and Industrial Research, New Delhi and Tata Institute of Fundamental Research, Mumbai are acknowledged. KVRC and BJ thank DST for their JC Bose Fellowships.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Singh, H., Rao, B.J. & Chary, K.V.R. 1H, 13C and 15N NMR assignments of a mutant of UV inducible transcript (S55A-UVI31+) from Chlamydomonas reinhardtii . Biomol NMR Assign 8, 371–374 (2014). https://doi.org/10.1007/s12104-013-9520-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-013-9520-4