Abstract
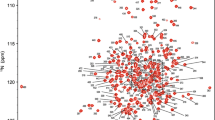
CGC-19, a 14 kDa proteic constituent of a non ribosomal peptide synthetase implicated in the biosynthesis of a secondary metabolite in Streptomyces ambofaciens, has been isotopically enriched and recombinantly expressed. Its nearly complete 1H, 13C and 15N resonance assignment is reported hereunder.

Similar content being viewed by others
References
Bahrami A, Assadi A, Markley JL, Eghbalnia H (2009) Probabilistic interaction network of evidence algorithm and its application to complete labeling of peak lists from protein NMR spectroscopy. PLoS Comput Biol 5(3):e1000307
Bialer M, El-On J, Yagen B, Mechoulam R (1981) Antiparasitic structure-activity relationships of congocidine derivatives. J Pharm Sci 70(7):822–824
Birlirakis N, Cerdan R, Guittet E (1996) A study of protein-water exchange through the off-resonance ROESY experiment: Application to the DNA-binding domain of AlcR. J Biomol NMR 8(4):487–491
Cai M, Huang Y, Sakaguchi K, Clore GM, Gronenborn AM, Craigie R (1998) An efficient and cost-effective isotope labeling protocol for proteins expressed in Escherichia coli. J Biomol NMR 11(1):97–102
Desvaux H, Berthault P, Birlirakis N, Goldman M (1994) Off-resonance ROESY: for the study of dynamics processes. J Magn Reson Series A 108:219–229
Juguet M, Lautru S, Francou FX, Nezbedová S, Leblond P, Gondry M, Pernodet JL (2009) An iterative nonribosomal peptide synthetase assembles the pyrrole-amide antibiotic congocidine in Streptomyces ambofaciens. Chem Biol 16(4):421–431
Kupce E, Schmidt P, Rance M, Wagner G (1998) Adiabatic mixing in the liquid state. J Magn Reson 135:361–367
McGuffin LJ, Bryson K, Jones DT (2000) The PSIPRED protein structure prediction server. Bioinformatics 16(4):404–405
Sattler M, Schleucher J, Griesinger C (1999) Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog Nucl Magn Reson Spectrosc 34:93–158
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44:213–223
Zimmerman DE, Kulikowski CA, Huang Y, Feng W, Tashiro M, Shimotakahara S, Chien C, Powers R, Montelione GT (1997) Automated analysis of protein NMR assignments using methods from artificial intelligence. J Mol Biol 269(4):592–610
Acknowledgments
We thank Dr. J.L. Pernodet and Dr. S. Lautru (Institut de Génétique et Microbiologie, UMR 8621, CNRS/Université Paris-sud 11) for supplying us with the CGC-19 plasmid and for fruitful discussions and Dr. J.P. Lecaer for recording the mass spectra. ICSN is acknowledged for a research fellowship to SN.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nogaret, S., Guittet, E. & Birlirakis, N. 1H, 13C and 15N NMR assignment of CGC-19, a single domain proteic constituent of a non ribosomal peptide synthetase. Biomol NMR Assign 7, 1–4 (2013). https://doi.org/10.1007/s12104-012-9364-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-012-9364-3