Abstract
Introduction
Lung cancer is one of the most prevalent cancers and the leading cause of cancer death. Advanced non-small cell lung cancer (aNSCLC) patients frequently harbor mutations that impact their survival outcomes. There are limited data regarding the prognostic and predictive significance of these mutations on survival outcomes in the real-world setting.
Methods
This observational retrospective study analyzed de-identified electronic medical records from the Flatiron Health Clinico-Genomic and FoundationCore® databases to identify patients with aNSCLC who initiated first-line immune checkpoint inhibitors (ICI; alone or in combination) or chemotherapy under routine care between 2016 and 2021. The primary objectives were to assess the prevalence of non-actionable mutations and to determine their association with overall survival (OS). Real-world progression-free survival (rwPFS) and real-world response (rwR) were investigated as secondary exploratory outcomes.
Results
Based on an assessment of 185 non-actionable mutations in 2999 patients, the most prevalent mutations were TP53 (70%), KRAS (42%), CDKN2A/B (31%), and STK11 (21%). STK11, KEAP1, and CDKN2A/B mutations were significantly associated with lower rwR, shorter rwPFS and OS. KRAS mutations were clinically associated with shorter rwPFS in CIT-treated patients. Subgroup analysis revealed that fast progressors were significantly more likely to harbor STK11, KEAP1, and CDKN2A/B mutations. Accordingly, long-term survivors (LTS) showed a significantly lower prevalence of these mutations.
Conclusion
Our results provide evidence on the prognostic value of STK11, KEAP1, and CDKN2A/B mutations in patients with aNSCLC. Further research is required to better understand the implications of these findings on patient management and future trial design and treatment selection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is the second most diagnosed cancer worldwide and the leading cause of cancer death to date [1]. Based on histology, it can be classified into small cell lung cancer and non-small cell lung cancer (NSCLC), which approximately represent 25% and 85% of lung cancer diagnoses, respectively. NSCLC can be further classified into lung adenocarcinoma (45–60%), squamous cell carcinoma (20–25%), and neuroendocrine carcinoma (10–15%), which are treated using diverse therapeutic strategies. Recently, the development of new targeted therapies and the use of immunotherapy have increased 5-year overall survival (OS) rates in patients with NSCLC [2, 3]. However, it remains unclear why certain subgroups of patients either do not respond to treatment or present significantly different survival rates than others.
In NSCLC, multiple driver mutations responsible for the initiation and maintenance of the cancer have been described (i.e., EGFR, ALK, ROS1, BRAF). This, in turn, has prompted the development of targeted therapies against them [3, 4]. However, most NSCLC patients do not harbor known driver mutations, or they have mutations that are not actionable [5]. In these cases, their care relies on immunotherapy ± chemotherapy [6], but they frequently present with non-driver or non-actionable mutations that affect disease progression, response to treatment and survival [3, 4]. Non-actionable mutations in certain tumor suppressor genes have been described to predict survival or response to treatment [6,7,8,9]. For instance, mutations in STK11, KEAP1, and CDKN2A/B genes have been linked to shorter survival and resistance to immunotherapy [10]. Nevertheless, the real-world evidence on non-driver and non-actionable mutations in advanced NSCLC (aNSCLC) patients remains limited, largely due to the small sizes of patient populations involved in the studies [11, 12]. Further research on the non-driver and non-actionable mutations associated with efficacy outcomes could help identify aNSCLC populations with high unmet needs, thus, guiding the choice of the best treatments based on their mutational pattern.
Our primary objective was to identify the predictive and prognostic value of non-driver and non-actionable mutations in a large real-world cohort of aNSCLC patients undergoing first-line (1L) chemotherapy and/or immune checkpoint inhibitors (ICI). Moreover, this study aimed to identify specific mutational profiles that could predict a patient’s response to treatment. Secondary objectives included: determining the overall prevalence of non-driver and non-actionable mutations in the selected patient cohort and characterizing the mutation profiles of specific patient subgroups known to respond differently to treatment, such as fast progressors (real-world time to progression [rwTP] < 3 months), long-term survivors (LTS, rwTP > 12 months), women, and never-smokers.
Methods
Study design
This observational retrospective cohort study was conducted employing the Flatiron Health–Foundation Medicine Clinico-Genomic database (FH-FMI CGDB), which includes patients from a subset of the Flatiron Health network of ~ 280 US cancer clinics (approximately 800 care sites). Retrospective longitudinal clinical data were derived from electronic health records and comprise patient-level structured and unstructured data including: patient demographics, precise diagnosis details such as staging, histopathology and biomarkers, along with selected treatment and outcomes. The clinical data are further enriched by linkage to genomic data that are procured from Foundation Medicine’s Core® database, enabling a deeper understanding of the patients’ genomic profiles.
Participants
All patients with aNSCLC who had initiated 1L ICI (alone or in combination with chemotherapy) or chemotherapy under routine clinical practice between January 1, 2016, and June 30, 2021, were selected. 2016 was set as the start of the study period to include the following immunotherapy approvals in the United States: pembrolizumab, nivolumab, ipilimumab, durvalumab and atezolizumab. Eligible patients were only those who received 1L ICI and/or chemotherapy, had next-generation sequencing (NGS) reports prior to the 1L treatment end date, had tissue sample only, non-driver or non-actionable mutations (EGFR, ALK, ROS1, BRAF V600E, RET, METex14, NTRK3 were considered as driver mutations), recorded activity within 90 days of advanced diagnosis, and absence of multiple primary cancers. There was no requirement for informed consent or ethical review and approval.
Study outcomes
The study aimed to estimate the prevalence of non-actionable mutations deemed clinically relevant by expert opinion.
Real-world overall survival (rwOS) for each patient was measured, defined as the duration from the index date (the start of the 1L treatment to the date of death).
Real-world progression-free survival (rwPFS) was defined as the period from the initiation of 1L therapy until the earliest recorded occurrence of any form of disease progression or death.
Real-world response rates (rwR) were determined by analyzing the numbers and percentages of patients who responded to treatment compared to the total population. RwR was considered as complete response upon clearance of all lesions and pathological nodes, while partial response was recorded for a decrease ≥ 30% of the sum of the maximum diameters.
The study also evaluated the association between non-actionable mutational patterns and the following patient subgroups of special interest: fast progressors, LTS, women, [13], and never-smokers [14]. The last two were selected based on previous results from meta-analyses on immunotherapy.
Statistical analysis
A descriptive analysis of the sociodemographic and clinical characteristics of this population was performed. Cox regression models were used to calculate hazard ratios (HR) and 95% confidence intervals (CI) for rwPFS and rwOS. Logistic regression models were used to obtain odds ratios (OR) and 95% CI for rwR.
To balance the differences in baseline characteristics between the mutated and wild-type groups, inverse probability weights were utilized. Propensity score models were employed to determine the associations between mutational status and rwPFS, rwOS, and rwR. This involved the inclusion of various prognostic variables: age, Eastern Cooperative Oncology Group performance status (ECOG PS) at 1L therapy, race, sex, type of diagnosis, smoking status, time from diagnosis to 1L therapy start date, histology, and brain/CNS metastasis at baseline. For each covariate, the balance between mutated and wild-type groups was evaluated using standard mean difference (SMD), where ideal balance was defined as SMD < 0.1. Multivariable modelling was restricted to mutations where the exposure group had at least 10 patients. The statistical analysis was performed using R package version 4.1.0. The significance level was set to alpha 0.05.
Results
Characteristics of the study population
A total of 10,795 patients with aNSCLC, who had initiated 1L ICI (alone or in combination with chemotherapy) or chemotherapy at data cut-off were selected. Of these, the 2999 patients (27.8%) who met the inclusion/exclusion criteria for this in-depth analysis were finally included in the study cohort (Fig. 1).
In the overall population, 65.3% received a ICI-containing treatment while the remaining 34.7% were treated with chemotherapy (Table 1). Mean age was 67.9 years. A higher proportion of patients aged ≥ 65 years was observed in the ICI-containing group compared to the chemotherapy one. Approximately half (53.4%) of the patients were men and nearly all had a history of smoking (95.1%). Histologically, non-squamous cell carcinoma was the most frequent type (67.7%), with a higher prevalence in the ICI-treated group. Overall, 58.4% of the patients had de novo diagnosis (63.2% in patients receiving ICI-containing treatment). Generally, ECOG PS was good (61.7% scoring 0). Regarding programmed cell death-ligand 1 (PD-L1) status, 32.1% patients in the ICI-containing group showed a PD-L1 expression of at least 50% (Table 1).
Prevalence of identified non-actionable mutations
A total of 185 different non-actionable mutations were identified. These mutations were grouped into 58 mutation families. The most prevalent mutations in the overall population were TP53, observed in 70% of patients; KRAS in 42%; CDKN2A/B in 31%; and STK11 in 21% (Fig. 2A). Interestingly, when the prevalence of these mutations was stratified by advanced diagnosis type (de novo or recurrent), no statistically significant differences were observed (Fig. 2B).
Response and survival outcomes
We assessed the association between the mutational status in patients with aNSCLC and the overall rwR, rwPFS, and rwOS. Of all identified mutations, STK11, KEAP1, and CDKN2A/B were significantly associated with all three effectiveness outcomes (Fig. 3). Patients harboring mutations in STK11 showed a statistically significant lower rwR (OR: 0.49 [0.39–0.62], p < 0.0001), shorter rwPFS (OR: 1.38 [1.19–1.59], p < 0.0001) and reduced rwOS (OR: 1.6 [1.39–1.84], p < 0.0001) than the wild-type population. APC and KRAS mutations were only significantly associated with lower rwR (Fig. 3A), while FGFR and HRAS mutations were related to worse rwPFS and rwOS (Fig. 3B, C) respectively. In contrast, a significantly higher likelihood of response or rwPFS was associated with ATM/R/RX and GATA3 mutations (Fig. 3B).
Volcano plots and real-world response values (A), real-world progression-free survival (B), and overall survival (C) in the overall population of patients with aNSCLC according to their mutational status. CI confidence interval, HR hazard ratio, OR odds ratio, OS overall survival, PFS progression-free survival, rw real-world
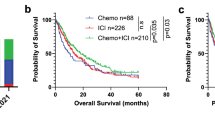
Furthermore, we evaluated effectiveness outcomes depending on the treatment regimen (Fig. 4). Low rwR and short rwPFS were found in patients harboring KRAS mutations treated with chemotherapy. In the ICI-containing group, patients with KEAP1 mutations showed low rwR and short rwPFS and rwOS, patients with CDKN2A/B mutations showed low rwR and short rwOS, while patients with STK11 mutations showed short rwOS. Despite these results, we did not find statistically significant efficacy differences between treatment regimens in patients harboring STK11, KEAP1 or CDKN2A/B mutations (Supplementary Fig. S1).
Analysis of subgroups of especial interest
Fast progressors were characterized by a significantly higher prevalence of STK11 (OR 1.68 [95% CI 1.41–2.01]), KEAP1 (OR 1.60 [95% CI 1.29–1.99]), and CDKN2A/B (OR 1.28 [95% CI 1.09–1.50]) mutations compared with non-fast progressors. Consistent with these results, LTS showed a significantly lower prevalence of these mutations (Table 2). In the LTS subgroup, we also observed significantly lower prevalence of FGFR1/2/3 mutations (OR 0.57 [95% CI 0.35–0.94]) and higher of KRAS mutations (OR 1.43 [95% CI 1.17–1.74]). In women, APC (OR 0.53 [95% CI 0.34–0.82]) and FGFR1/2/3 (OR 0.58 [95% CI 0.43–0.79]) mutations were less frequent, while KRAS mutations were more frequent (OR 1.98 [95% CI 1.71–2.30]). In never-smokers, STK11 (OR 2.30 [95% CI 1.36–3.91]) and KEAP1 (OR 4.48 [95% CI 1.82–11.00]) mutations were significantly less prevalent. We also observed a trend towards a lower prevalence of FGFR1/2/3 (OR 2.45 [95% CI 0.90–6.71]).
In addition, in order to determine whether the treatment regimen was associated with fast progression or LTS in patients with a certain mutation, we performed a subgroup analysis. The logistic regression analysis did not show statistically significant differences between treatment groups. However, numerical differences were observed. Regarding KEAP1-mutated patients, there was a larger proportion of fast progressors in the ICI-containing-treated group (55.9%) compared to the chemotherapy-treated group (48.9%). We also observed a higher proportion of fast progressors among patients with FGFR1/2/3 (53.5% vs 39.3%) and APC mutations (49.1% vs 41.5%) in the ICI-containing group. Lastly, KRAS (19.8% vs 14.2%) and APC-mutant patients (18.9 vs 7.3%) showed a larger proportion of LTS in the ICI-containing group compared with those who received chemotherapy.
Discussion
To our knowledge, this is the largest real-world study to date describing the non-actionable mutational profile and its association with prognosis and predictive value in 1L patients with aNSCLC. In this dataset, we identified over 180 mutations, the most prevalent being TP53, KRAS, CDKN2A/B, and STK11 mutations. These results are in line with those reported in the literature, except for a higher prevalence of TP53 mutation, which is commonly associated with squamous histology [10, 15,16,17]. In the literature, there is very little evidence regarding CDKN2A/B in aNSCLC, with contradictory findings [10]. In this context, the large dataset used in our study and the significant outcomes confirms the importance of evaluating CDKN2A/B mutation as part of the mutational pattern in aNSCLC. We report that STK11, KEAP1, and CDKN2A/B mutations were significantly associated with poor prognosis in all effectiveness outcomes. In addition, KRAS mutations led to a lower rwR and clinically significant differences in rwPFS associated with different treatment regimens.
STK11 is a tumor suppressor kinase, which negatively regulates the AMPK/mTOR pathway and is somatically inactivated in up to 30% of patients with NSCLC [18, 19]. A large real-world observational genome study found that STK11 mutations had a negative prognostic value in patients with metastatic NSCLC treated with chemotherapy or immunotherapy [17]. Similarly, another observational study determined that STK11 and KEAP1 mutations were associated with shorter rwPFS and rwOS in all treatment groups, suggesting a prognostic but not predictive value for these biomarkers [16]. A more recent study showed that treatment with atezolizumab in patients harboring STK11 or KEAP1 mutations resulted in longer OS [20]. In our overall population, patients with mutated STK11 showed lower rwR, rwPFS, and rwOS, and, in line with other reports, we did not observe statistically significant differences between chemotherapy and ICI-containing groups [16, 17].
KEAP1 is a negative regulator of nuclear factor erythroid 2-related factor 2 involved in cell defense, and cytoprotective response to endogenous and exogenous stress [21]. Somatic mutations in KEAP1 are found in about 20% of patients with NSCLC [19]. Goeman and colleagues showed that KEAP1 mutations were associated with shorter survival outcomes in patients with aNSCLC [22]. Additional studies have also reported that patients harboring KEAP1 mutations showed shorter survival regardless of treatment type [23, 24]. In this context, there are several ongoing clinical trials evaluating the efficacy of targeted therapy [25]. In our study, patients in the ICI-containing group exhibited worse outcomes compared to those treated with chemotherapy. This suggests a potential predictive role of KEAP1 for ICI-containing treatment, although further studies are required to validate these results. Overall, harboring a KEAP1 mutation led to lower rwR, rwPFS, and rwOS and our results are consistent with those published elsewhere [16, 20].
Mutations in STK11 and KEAP1 are associated with poor outcomes in patients with NSCLC, despite high TMB, including outcomes with PD-1 inhibitors [16, 26]. Inactivation of STK11 in lung cancer appears to result in an immunologically cold tumor microenvironment, with reduced T cell infiltration [26,27,28]. KEAP1 appears to interact functionally with STK11 [29] and these two proteins are significantly co-mutated in NSCLC, and result in a poor OS prognosis [30, 31]. OS and PFS outcomes in mSTK11 and mKEAP1 patients were improved by ICI treatment in several studies [32].
CDKN2A/B genes encode potent tumor suppressor proteins. In agreement, loss of function mutations in these genes negatively impact patient outcomes [33]. In this regard, our study showed that mutations in CDKN2A/B genes were associated with reduced rwR and shorter rwPFS and rwOS. Similarly, Gutiontov and colleagues showed that CDKN2A loss of function worsened clinical outcomes in aNSCLC patients treated with ICI [10]. In the same line, our ICI-containing group patients showed a trend towards lower rwR and shorter rwOS compared with those treated with chemotherapy. Only a few, small-scale studies have assessed the prevalence of CDKN2A/B mutations in aNSCLC and its impact on treatment outcomes [10, 34]; ours is one of the largest describing and confirming the role of CDKN2A/B in this setting.
KRAS is one of the most frequently mutated genes in cancer, being observed in up to 30% of patients with NSCLC [9, 35, 36]. Some studies suggest that KRAS mutations are associated with poor prognosis, while others found no correlation [35, 36]. In this context, a recent systematic review and meta-analysis analyzed 43 clinical studies to assess KRAS impact. Authors concluded that KRAS mutations may be associated with poor prognosis and response outcomes, but more evidence of its predictive value is needed [37]. We observed similar results, especially in patients treated with chemotherapy: KRAS mutation in these patients resulted in a numerically lower rwR and shorter rwPFS compared with patients in the ICI-containing group. Similarly, a recently published pooled analysis described that patients with KRAS mutations treated with ICI-containing therapy displayed a greater response and survival compared with those treated only with chemotherapy [38]. Overall, our data suggest that KRAS mutation may have predictive value for PFS in ICI-containing treated patients. However, additional research is needed to validate this clinical significance.
In the subgroup analyses, we observed that STK11, KEAP1, and CDKN2A/B mutations were significantly associated with fast disease progression, and together with FGFR1/2/3 mutations with shorter survival. These results are consistent with our data on rwR, rwPFS, and rwOS; fast progressors seem to be more likely to harbor mutations in STK11, KEAP1, and CDKN2A/B and, therefore, have a poor prognosis. In a previous study, it was reported that KEAP1 mutations are overrepresented in fast progressors, and it was suggested that they could define a molecular subset of patients characterized by resistance to chemotherapy [22]. Our results support this conclusion and also suggest that KEAP1 mutations could be a potential predictive biomarker for poor survival in patients treated with ICI-containing therapy. We observed a larger proportion of LTS in KRAS-mutated patients when treated with ICI-containing therapy, compared to chemotherapy. Given that this result is in line with the lower rwR and shorter rwPFS observed in patients treated with chemotherapy, it supports the idea of KRAS being a potential predictive biomarker.
The present study makes a significant contribution to the current literature on the mutational profile of patients with aNSCLC, although its retrospective nature could be considered a limitation. However, the real-world data and the large size of our dataset ensure representativity of the aNSCLC population. An additional limitation is the fact that certain mutations were observed in a limited number of patients, leading to CIs too large to draw any conclusions when carrying out comparisons. Furthermore, co-occurrence of mutations was not considered and the effect on tumor mutation burden was not investigated, making it impossible to exclude a potential selection bias. Finally, since sotorasib was approved while the study was ongoing (June 2021), we could not indicate whether patients with KRAS mutations were treated with this therapy, and we did not investigate different KRAS variants separately.
In conclusion, our study describes the prevalence and mutational pattern of 1L aNSCLC and shows that mutations in genes such as STK11, KEAP1 and CDKN2A/B are significantly associated with poor efficacy outcomes. Thus, they could be considered prognostic factors. The same mutational profile was observed in de novo and recurrent patients, but other subgroups of patients, such as fast progressors and LTS, were characterized by distinct patterns of STK11, KEAP1, and CDKN2A/B mutations that could guide clinical decision making and help predict treatment response in patients with aNSCLC. Overall, our results contribute to the identification of novel biomarkers that could help clinicians determine the degree of treatment response expected from certain subgroups of patients. Further studies are needed to support these results and to evaluate their impact in clinical trial design or clinical decision making.
Data availability
Qualified researchers may request access to individual patient-level data through the clinical study data request platform (https://vivli.org/). Further details on Roche’s criteria for eligible studies are available here: https://vivli.org/members/ourmembers/. For further details on Roche’s Global Policy on the Sharing of Clinical Information and how to request access to related clinical study documents, see here: https://www.roche.com/research_and_development/who_we_are_how_we_work/clinical_trials/our_commitment_to_data_sharing.htm.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49.
Reck M, Rodriguez-Abreu D, Robinson AG, Hui R, Csoszi T, Fulop A, et al. Five-year outcomes with pembrolizumab versus chemotherapy for metastatic non-small-cell lung cancer with PD-L1 tumor proportion score >/= 50. J Clin Oncol. 2021;39(21):2339–49.
Rodak O, Peris-Diaz MD, Olbromski M, Podhorska-Okolow M, Dziegiel P. Current landscape of non-small cell lung cancer: epidemiology, histological classification, targeted therapies, and immunotherapy. Cancers (Basel). 2021;13(18):4705.
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr, Kinzler KW. Cancer genome landscapes. Science. 2013;339(6127):1546–58.
Le X, Elamin YY, Zhang J. New actions on actionable mutations in lung cancers. Cancers (Basel). 2023;15(11):2917.
Spira AI, Tu H, Aggarwal S, Hsu H, Carrigan G, Wang X, et al. A retrospective observational study of the natural history of advanced non-small-cell lung cancer in patients with KRAS p.G12C mutated or wild-type disease. Lung Cancer. 2021;159:1–9.
Bazhenova L, Lokker A, Snider J, Castellanos E, Fisher V, Fellous M, et al. TRK fusion cancer: patient characteristics and survival analysis in the real-world setting. Target Oncol. 2021;16(3):389–99.
Hellmann MD, Nathanson T, Rizvi H, Creelan BC, Sanchez-Vega F, Ahuja A, et al. Genomic features of response to combination immunotherapy in patients with advanced non-small-cell lung cancer. Cancer Cell. 2018;33(5):843-52.e4.
Skoulidis F, Heymach JV. Co-occurring genomic alterations in non-small-cell lung cancer biology and therapy. Nat Rev Cancer. 2019;19(9):495–509.
Gutiontov SI, Turchan WT, Spurr LF, Rouhani SJ, Chervin CS, Steinhardt G, et al. CDKN2A loss-of-function predicts immunotherapy resistance in non-small cell lung cancer. Sci Rep. 2021;11(1):20059.
Arbour KC, Shen R, Plodkowski A, Rizvi H, Ni A, Long N, et al. Concurrent mutations in STK11 and KEAP1 is associated with resistance to PD-(L)1 blockade in patients with NSCLC despite high TMB. J Thorac Oncol. 2018;13(10):S424.
Cho BC, Lopes G, Kowalski DM, Kasahara K, Wu Y-L, Castro G et al. Relationship between STK11 and KEAP1 mutational status and efficacy in KEYNOTE-042: pembrolizumab monotherapy versus platinum-based chemotherapy as first-line therapy for PD-L1-positive advanced NSCLC. Presented at: AACR Virtual Annual Meeting I 2020.
Pinto JA, Vallejos CS, Raez LE, Mas LA, Ruiz R, Torres-Roman JS, et al. Gender and outcomes in non-small cell lung cancer: an old prognostic variable comes back for targeted therapy and immunotherapy? ESMO Open. 2018;3(3): e000344.
Zhao W, Jiang W, Wang H, He J, Su C, Yu Q. Impact of smoking history on response to immunotherapy in non-small-cell lung cancer: a systematic review and meta-analysis. Front Oncol. 2021;11: 703143.
Arbour KC, Jordan E, Kim HR, Dienstag J, Yu HA, Sanchez-Vega F, et al. Effects of co-occurring genomic alterations on outcomes in patients with KRAS-mutant non-small cell lung cancer. Clin Cancer Res. 2018;24(2):334–40.
Papillon-Cavanagh S, Doshi P, Dobrin R, Szustakowski J, Walsh AM. STK11 and KEAP1 mutations as prognostic biomarkers in an observational real-world lung adenocarcinoma cohort. ESMO Open. 2020;5(2): e000706.
Shire NJ, Klein AB, Golozar A, Collins JM, Fraeman KH, Nordstrom BL, et al. STK11 (LKB1) mutations in metastatic NSCLC: prognostic value in the real world. PLoS ONE. 2020;15(9): e0238358.
Gleeson FC, Kipp BR, Levy MJ, Voss JS, Campion MB, Minot DM, et al. Somatic STK11 and concomitant STK11/KRAS mutational frequency in stage IV lung adenocarcinoma adrenal metastases. J Thorac Oncol. 2015;10(3):531–4.
Cancer Genome Atlas Research N. Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511(7511):543–50.
Nie W, Gan L, Wang X, Gu K, Qian FF, Hu MJ, et al. Atezolizumab prolongs overall survival over docetaxel in advanced non-small-cell lung cancer patients harboring STK11 or KEAP1 mutation. Oncoimmunology. 2021;10(1):1865670.
Jaramillo MC, Zhang DD. The emerging role of the Nrf2-Keap1 signaling pathway in cancer. Genes Dev. 2013;27(20):2179–91.
Goeman F, De Nicola F, Scalera S, Sperati F, Gallo E, Ciuffreda L, et al. Mutations in the KEAP1-NFE2L2 pathway define a molecular subset of rapidly progressing lung adenocarcinoma. J Thorac Oncol. 2019;14(11):1924–34.
Jeong Y, Hellyer JA, Stehr H, Hoang NT, Niu X, Das M, et al. Role of KEAP1/NFE2L2 mutations in the chemotherapeutic response of patients with non-small cell lung cancer. Clin Cancer Res. 2020;26(1):274–81.
Skoulidis F, Arbour KC, Hellmann M, Patil PD, Marmarelis ME, Awad MM, et al. Association of STK11/LKB1 genomic alterations with lack of benefit from the addition of pembrolizumab to platinum doublet chemotherapy in non-squamous non-small cell lung cancer. J Clin Oncol. 2019;37(suppl 15):102.
Hellyer JA, Padda SK, Diehn M, Wakelee HA. Clinical implications of KEAP1-NFE2L2 mutations in NSCLC. J Thorac Oncol. 2021;16(3):395–403.
Skoulidis F, Goldberg ME, Greenawalt DM, Hellmann MD, Awad MM, Gainor JF, et al. STK11/LKB1 mutations and PD-1 inhibitor resistance in KRAS-mutant lung adenocarcinoma. Cancer Discov. 2018;8(7):822–35.
Koyama S, Akbay EA, Li YY, Aref AR, Skoulidis F, Herter-Sprie GS, et al. STK11/LKB1 deficiency promotes neutrophil recruitment and proinflammatory cytokine production to suppress T cell activity in the lung tumor microenvironment. Cancer Res. 2016;76(5):999–1008.
Kadara H, Choi M, Zhang J, Parra ER, Rodriguez-Canales J, Gaffney SG, et al. Whole-exome sequencing and immune profiling of early-stage lung adenocarcinoma with fully annotated clinical follow-up. Ann Oncol. 2017;28(1):75–82.
Galan-Cobo A, Sitthideatphaiboon P, Qu X, Poteete A, Pisegna MA, Tong P, et al. LKB1 and KEAP1/NRF2 pathways cooperatively promote metabolic reprogramming with enhanced glutamine dependence in KRAS-mutant lung adenocarcinoma. Cancer Res. 2019;79(13):3251–67.
Skoulidis F, Byers LA, Diao L, Papadimitrakopoulou VA, Tong P, Izzo J, et al. Co-occurring genomic alterations define major subsets of KRAS-mutant lung adenocarcinoma with distinct biology, immune profiles, and therapeutic vulnerabilities. Cancer Discov. 2015;5(8):860–77.
Facchinetti F, Bluthgen MV, Tergemina-Clain G, Faivre L, Pignon JP, Planchard D, et al. LKB1/STK11 mutations in non-small cell lung cancer patients: Descriptive analysis and prognostic value. Lung Cancer. 2017;112:62–8.
Mok TSK, Lopes G, Cho BC, Kowalski DM, Kasahara K, Wu YL, et al. Associations of tissue tumor mutational burden and mutational status with clinical outcomes in KEYNOTE-042: pembrolizumab versus chemotherapy for advanced PD-L1-positive NSCLC. Ann Oncol. 2023;34(4):377–88.
Zhao R, Choi BY, Lee MH, Bode AM, Dong Z. Implications of genetic and epigenetic alterations of CDKN2A (p16(INK4a)) in cancer. EBioMedicine. 2016;8:30–9.
Cheng YL, Lee SC, Harn HJ, Chen CJ, Chang YC, Chen JC, et al. Prognostic prediction of the immunohistochemical expression of p53 and p16 in resected non-small cell lung cancer. Eur J Cardiothorac Surg. 2003;23(2):221–8.
Reck M, Carbone DP, Garassino M, Barlesi F. Targeting KRAS in non-small-cell lung cancer: recent progress and new approaches. Ann Oncol. 2021;32(9):1101–10.
Yang H, Liang SQ, Schmid RA, Peng RW. New horizons in KRAS-mutant lung cancer: dawn after darkness. Front Oncol. 2019;9:953.
Goulding RE, Chenoweth M, Carter GC, Boye ME, Sheffield KM, John WJ, et al. KRAS mutation as a prognostic factor and predictive factor in advanced/metastatic non-small cell lung cancer: a systematic literature review and meta-analysis. Cancer Treat Res Commun. 2020;24: 100200.
Nakajima EC, Ren Y, Vallejo JJ, Akinboro O, Mishra-Kalyani PS, Larkins EA, et al. Outcomes of first-line immune checkpoint inhibitors with or without chemotherapy according to KRAS mutational status and PD-L1 expression in patients with advanced NSCLC: FDA pooled analysis. J Clin Oncol. 2022;40(16 suppl):9001.
Acknowledgements
This study was funded by Roche Spain. The authors would like to thank Ánchel González Barriga, from Medical Science Consulting (Valencia, Spain), for providing medical writing services.
Author information
Authors and Affiliations
Contributions
MPP conceptualization; investigation; methodology; project administration; supervision; validation; visualization; writing—review and editing. DPP: conceptualization; data curation; formal analysis; investigation; methodology; project administration; supervision; validation; visualization; writing — original draft; writing — review and editing. SO: conceptualization; data curation; formal analysis; investigation; methodology; project administration; supervision; validation; visualization; writing — original draft; writing — review and editing. HH: data curation; formal analysis; methodology; software; writing — review and editing. BCB: investigation; validation; visualization; writing — review and editing. DRA: investigation; validation; visualization; writing — review and editing. MLBP: investigation; validation; visualization; writing — review and editing. NP: data curation; formal analysis; methodology; software; writing — review and editing. SW: data curation; formal analysis; software; writing — review and editing. EV: conceptualization; methodology; writing — review and editing. PRG: conceptualization; methodology; writing — review and editing. MCD: conceptualization; investigation; methodology; project administration; supervision; validation; visualization; writing — review and editing.
Corresponding author
Ethics declarations
Conflict of interest
MPP: Advisory board (Roche, Merck, BMS, AstraZeneca, Lilly, Pfizer, Bayer, Amgen, Janssen, Takeda), Speaker (Roche, Merck, BMS, Takeda, Sanofi), Research grant (Roche, BMS, Takeda). BCB: Speaker (Roche, Merck, BMS, AstraZeneca, Sanofi), Travel/accommodation/expenses (Roche, BMS, Lilly, Pfizer, Boehringer). DRA: Advisory board (Roche, Merck, BMS, AstraZeneca, Pfizer, Boehringer, Takeda), Speaker (Roche, Merck, BMS, AstraZeneca, Boehringer, Takeda), Travel/accommodation/expenses (Roche, Merck, BMS, AstraZeneca). MLBP: Advisory board (Roche, Merck, BMS, Takeda), Speaker (Roche, Merck, BMS, AstraZeneca, Pfizer, Takeda). MCD: Advisory board (Roche, BMS, AstraZeneca, Boehringer), Travel/accommodation/expenses (Roche, BMS, AstraZeneca). DPP, SO, HH, NPl, SW, EV, and PRG are Roche Farma SA employees.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
For this type of study, formal consent is not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Provencio-Pulla, M., Pérez-Parente, D., Olson, S. et al. Identification of non-actionable mutations with prognostic and predictive value in patients with advanced or metastatic non-small cell lung cancer. Clin Transl Oncol 26, 1384–1394 (2024). https://doi.org/10.1007/s12094-023-03362-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-023-03362-8