Abstract
Purpose
Genetic factors play an important role in our predisposition to cancer. Genome-wide association studies have linked the chromosome 15q25.1 locus to lung cancer susceptibility and implicated proteasome subunit alpha type-4 (PSMA4) as a candidate gene. In this case–control study, pathologically confirmed lung cancer patients and controls from the Chinese Han population were investigated to determine the effect of variant genotypes within the PSMA4 locus on susceptibility to lung cancer and sensitivity to cisplatin-based chemotherapy.
Methods
We identified validated tagged single nucleotide polymorphisms (tSNPs) with minor allele frequency >5 % in the HapMap Chinese Han Beijing population and genotyped seven SNPs within the PSMA4 locus. Their correlation with lung cancer risks and treatment response were examined using χ 2 test and haplotype analysis. Multivariate logistic regression analysis tested the association between the polymorphisms and chemotherapy response.
Results
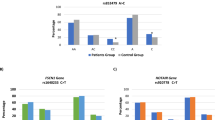
rs12901682 is associated with lung cancer risks (OR = 1.45, 95 % CI, 1.04–2.02; P = 0.029). Using SNPStats software, we found rs12901682 (OR = 6.30, 95 % CI, 1.31–30.26; P = 0.0073) associated with lung cancer risks in the recessive model. Haplotype analysis showed that “CAGAATC” conferred an increased risk of lung cancer (OR = 1.50, 95 % CI, 1.07–2.11; P = 0.019). After adjustment for age, this association was pronounced in the male gender (OR = 6.30, 95 % CI, 1.31–30.26; P = 0.0073). PSMA4 polymorphisms did not affect the tumor sensitivity to cisplatin combination chemotherapy.
Conclusions
Our study suggests a potential association between PSMA4 variants and lung cancer risk in Chinese Han population.
Similar content being viewed by others
References
Smith RA, Cokkinides V, Brooks D, Saslow D, Shah M, Brawley OW. Cancer screening in the United States, 2011: a review of current American Cancer Society guidelines and issues in cancer screening. CA Cancer J Clin. 2011;61:8–30.
Pirozynski M. 100 years of lung cancer. Respir Med. 2006;100:2073–84.
Chen WQ, Zeng HM, Zheng RS, Zhang SW, He J. Cancer incidence and mortality in China. Chin J Cancer Res. 2007;24:1–8.
Chen WQ, Zhang SW, Zou XN. Evaluation on the incidence, mortality and tendency of lung cancer in China. Thorac Cancer. 2010;1:35–40.
Wang Y, Liu J, Li S, Wang Q. Comparison and analysis of the incidence and mortality rate of cancer in developed and developing countries. Chin J Clin Oncol. 2012;2012(39):679–82.
Lichtenstein P, Holm NV, Verkasalo PK, Iliadou A, Kaprio J, Koskenvuo M, et al. Environmental and heritable factors in the causation of cancer—analyses of cohorts of twins from Sweden, Denmark, and Finland. N Engl J Med. 2000;343:78–85.
Shields PG. Molecular epidemiology of smoking and lung cancer. Oncogene. 2002;21:6870–6.
Taioli E, Benhamou S, Bouchardy C, Cascorbi I, Cajas-Salazar N, Dally H, et al. Myeloperoxidase g-463a polymorphism and lung cancer: a huge genetic susceptibility to environmental carcinogens pooled analysis. Genet Med. 2007;9:67–73.
Yokota J, Shiraishi K, Kohno T. Genetic basis for susceptibility to lung cancer: recent progress and future directions. Adv Cancer Res. 2010;109:51–72.
Liu PY, Vikis HG, Lu Y, Wang YA, Schwartz AG, Pinney SM, et al. Cumulative effect of multiple loci on genetic susceptibility to familial lung cancer. Cancer Epidemiol Biomarkers. 2010;19:517–24.
Brahmer JR. Genetic susceptibility to lung cancer in women. J Thorac Oncol. 2009;4:S61–2.
Wang W, Spitz MR, Yang HS, Lu C, Stewart DJ, Wu XF. Genetic variants in cell cycle control pathway confer susceptibility to lung cancer. Clin Cancer Res. 2007;13:5974–81.
Hansen HM, Xiao Y, Rice T, Bracci PM, Wrensch MR, Sison JD, et al. Fine mapping of chromosome 15q25.1 lung cancer susceptibility in African–Americans. Hum Mol Genet. 2010;19(18):3652–3661. doi:10.1093/hmg/ddq268.
D’Addario G, Pintilie M, Leighl NB, Feld R, Cerny T, Shepherd FA. Platinum-based versus non-platinum-based chemotherapy in advanced non-small-cell lung cancer: a meta-analysis of the published literature. J Clin Oncol. 2005;23:2926–36.
Chang A. Chemotherapy, chemoresistance and the changing treatment landscape for NSCLC. Lung Cancer. 2011;71(1):3–10. doi:10.1016/j.lungcan.2010.08.022.
Seng KC, Seng CK. The success of the genome-wide association approach: a brief story of a long struggle. Eur J Hum Genet. 2008;16:554–64.
Therasse P, Arbuck SG, Eisenhauer EA, Wanders J, Kaplan RS, Rubinstein L, et al. New guidelines to evaluate the response to treatment in solid tumors. J Natl Cancer I. 2000;92:205–16.
Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R, et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer. 2009;45:228–47.
Ramirez JL, Rosell R, Taron M, Gupta J, Alberola V, de las Penas R, et al. 14-3-3 sigma (sigma) methylation (M) in pre-treatment serum DNA of cisplatin (cis)/gemcitabine (gem)-treated non-small-cell lung cancer (NSCLC). J Clin Oncol. 2005;23:631s.
Hansen HM, Xiao YY, Rice T, Bracci PM, Wrensch MR, Sison JD, et al. Fine mapping of chromosome 15q25.1 lung cancer susceptibility in African–Americans. Hum Mol Genet. 2010;19:3652–61.
Amos CI, Gorlov IP, Dong QO, Wu XF, Zhang HF, Lu EY, et al. Nicotinic acetylcholine receptor region on chromosome 15q25 and lung cancer risk among African Americans: a case–control study. J Natl Cancer I. 2010;102:1199–205.
Liu Y, Liu P, Wen W, James MA, Wang Y, Bailey-Wilson JE, et al. Haplotype and cell proliferation analyses of candidate lung cancer susceptibility genes on chromosome 15q24-25.1. Cancer Res. 2009;69:7844–50.
Amos CI, Wei CJ, Wun I, Dong Q, Spitz MR, Frazier ML. Exploring rare variants and lung cancer risk from the nicotinic acetylcholine receptor region on chromosome 15q25 in African–Americans. Genet Epidemiol. 2009;33:766.
Liu PY, Vikis HG, Wang DL, Lu Y, Wang Y, Schwartz AG, et al. Familial aggregation of common sequence variants on 15q24-25.1 in lung cancer. J Natl Cancer I. 2008;100:1326–30.
Amos CI, Wu XF, Broderick P, Gorlov IP, Gu J, Eisen T, et al. Genome-wide association scan of tag SNPS identifies a susceptibility locus for lung cancer at 15q25.1. Nat Genet. 2008;40:616–22.
Gabriel S, Ziaugra L, Tabbaa D. SNP genotyping using the Sequenom massARRAY iPLEX platform. Current Protoc Hum Genet. 2009;Vol 1 Chapter 2: Unit 2 12.
Thomas RK, Baker AC, Debiasi RM, Winckler W, Laframboise T, Lin WM, et al. High-throughput oncogene mutation profiling in human cancer. Nat Genet. 2007;39:347–51.
Li Z, Bao S, Xu X, Bao Y, Zhang Y. Polymorphisms of CHRNA5-CHRNA3-CHRNB4 gene cluster and NSCLC risk in Chinese population. Transl Oncol. 2012;5(6):448–452.
Speedy HE, Di Bernardo MC, Sava GP, Dyer MJ, Holroyd A, Wang Y, et al. A genome-wide association study identifies multiple susceptibility loci for chronic lymphocytic leukemia. Nat Genet. 2014;46(1):56–60. doi:10.1038/ng.2843.
Bland JM, Altman DG. Statistics notes. The odds ratio. BMJ. 2000;320:1468.
Shi YY, He L. SHEsis, a powerful software platform for analyses of linkage disequilibrium, haplotype construction, and genetic association at polymorphism loci. Cell Res. 2005;15:97–8.
Sole X, Guino E, Valls J, Iniesta R, Moreno V. SNPStats: a web tool for the analysis of association studies. Bioinformatics. 2006;22:1928–9.
Tsai YY, McGlynn KA, Cassidy AB, Hu Y, Arnold J, Engstrom PF, et al. Dietary patterns and genetic susceptibility to lung cancer risk. Am J Hum Genet. 2001;69:206.
Quinones L, Berthou F, Varela N, Simon B, Gil L, Lucas D. Ethnic susceptibility to lung cancer: differences in CYP2E1, CYP1A1 and GSTM1 genetic polymorphisms between French Caucasian and Chilean populations. Cancer Lett. 1999;141:167–71.
Guillot M, Martin C, Manzano JL, Balana C, Font A, Barnadas A, et al. Genetic susceptibility to lung cancer associated with rare HRAS1 VNTR alleles in Spanish lung cancer patients. Eur J Cancer. 1999;35:S68.
Zhang Y, Zhang A, Shi H, Kong X. Association between TGM5, PPAP2B and PSMA4 polymorphisms and NSCLC in never-smoking Chinese population. J Can Res Ther. 2013;9:660–3.
Perneger TV. What’s wrong with Bonferroni adjustments. BMJ. 1998;316:1236–8.
Acknowledgments
This work was supported by the National Science and Technology Major Project of the Ministry of Science and Technology of China (No. 2012ZX09506001) and Collaboration and Communication Project of International Science and Technology of Shaanxi Province (Grant No. 2014KW24-03).
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
T. Wang and T. Chen contributed equally.
Rights and permissions
About this article
Cite this article
Wang, T., Chen, T., Thakur, A. et al. Association of PSMA4 polymorphisms with lung cancer susceptibility and response to cisplatin-based chemotherapy in a Chinese Han population. Clin Transl Oncol 17, 564–569 (2015). https://doi.org/10.1007/s12094-015-1279-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-015-1279-x