Abstract
The six proteins of the CCN family have important roles in development, angiogenesis, cell motility, proliferation, and other fundamental cell processes. To date, CCN5 distribution in developing rodents and humans has not been mapped comprehensively. CCN5 strongly inhibits adult smooth muscle cell proliferation and motility. Its anti-proliferative action predicts that CCN5 would not be present in developing tissues until the proliferation phase of tissue morphogenesis is complete. However, estrogen induces CCN5 expression in epithelial and smooth muscle cells, suggesting that CCN5 might be widely expressed in embryonic tissues exposed to high levels of estrogen. 9–16 day murine embryos and fetuses and 3–7 month human fetal tissues were analyzed by immunohistochemistry. CCN5 was detected in nearly all developing tissues. CCN5 protein expression was initially present in most tissues, and at later times in development tissue-specific expression differences were observed. CCN5 expression was particularly strong in vascular tissues, cardiac muscle, bronchioles, myotendinous junctions, and intestinal smooth muscle and epithelium. CCN5 expression was initially absent in bone cartilaginous forms but was increasingly expressed during bone endochondral ossification. Widespread CCN5 mRNA expression was detected in GD14.5 mice. Although CCN2 and CCN5 protein expression patterns in some adult pathologic conditions are inversely expressed, this expression pattern was not found in developing mouse and human tissues. The widespread expression pattern of CCN5 in most embryonic and fetal tissues suggests a diverse range of functions for CCN5.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
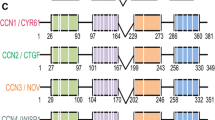
CCN5 is a member of the cysteine-rich 61/connective tissue growth factor/nephroblastoma-overexpressed (CCN) family of genes (Perbal and Takigawa 2005b). The six members of this family are matricellular proteins that have important functions in numerous cell and physiologic processes, including embryonic development, cell motility and proliferation, angiogenesis, and extracellular matrix biology (Perbal and Takigawa 2005a; Leask and Abraham 2006; Perbal 2004; Rachfal and Brigstock 2005). CCN5, previously known as WISP-2 (Pennica et al. 1998), rCop-1 (Zhang et al. 1998), COP-1 (Delmolino et al, 2001), HICP (Delmolino et al, 1997), and CTGF-L (Kumar et al. 1999), is highly conserved among vertebrates and is the only CCN protein that lacks the fourth C-terminal domain (Gray and Castellot 2005). In adult rodents, CCN5 mRNA is highly expressed in the aorta, heart, brain, spleen, lung, skeletal muscle, and uterus, and at lower levels in the spleen, liver, kidney, and testis (Delmolino et al. 2001; Mason et al. 2004a).
Originally discovered as a heparin-induced gene in vascular smooth muscle cells (SMC), CCN5 behaves as a growth-arrest-specific gene in this cell type. CCN5 is highly expressed in quiescent, non-proliferating rat aortic SMC and expression levels drop rapidly as cells are stimulated to re-enter the cell cycle. SMC CCN5 expression decreases in uterine fibroids and after vascular injury, two in vivo models of SMC proliferation (Delmolino et al. 2001; Lake et al. 2003; Mason et al. 2004b). CCN5 over-expression inhibits SMC proliferation, motility, and invasiveness in vitro, and CCN5 knock-down causes opposite effects (Lake et al. 2003; Lake and Castellot 2003; Mason et al. 2004b). CCN5 is strongly up-regulated by estrogen in both SMC and epithelial cells (Fritah et al. 2006; Gray and Castellot 2005; Inadera et al. 2000; Mason et al. 2004a).
There are several pathologic conditions in which CCN2 and CCN5 have opposite expression patterns, including asthma (unpublished observations; van den Brule S et al. 2007) uterine leiomyoma (fibroids) (De Falco et al. 2006; Mason et al. 2004b), and hepatocellular carcinoma (Cervello et al. 2004; Hirasaki et al. 2001). However, in other diseases, including arthritis (Manns et al. 2006; Tanaka et al. 2005) and viral hepatitis (Fukutomi et al. 2005; Shin et al. 2005), CCN2 and CCN5 are similarly expressed. CCN2 and CCN5 have opposite expression patterns in vascular SMC. CCN2 over-expression induces vascular SMC proliferation and increases MMP-2 expression, and CCN5 over-expression reduces proliferation and MMP-2 expression. CCN5 expression decreases and CCN2 expression increases in vascular SMC during the proliferative phase of balloon angioplasty injury (Ando et al. 2004; Fan et al. 2000; Fan and Karnovsky 2002; Lake et al. 2003; Lake and Castellot 2003). Opposite expression patterns of CCN2 and CCN5 in vascular SMC and in some proliferative disease states suggest that they might have complementary or opposite expression patterns in the developing embryo.
The temporospatial embryonic expression patterns of CCN1, CCN2, CCN3, and CCN4 have been comprehensively analyzed in developing mammals, (French et al. 2004; Friedrichsen et al. 2003; Ivkovic et al. 2003; Kireeva et al. 1997; Kocialkowski et al. 2001; Lopes et al. 2004; Natarajan et al. 2000; O’Brien and Lau 1992; Surveyor et al. 1998; Surveyor and Brigstock 1999) however, the expression pattern of CCN5 has not been explored. The strong anti-proliferative and anti-motility activity of CCN5 against cultured SMC (Delmolino et al. 2001; Lake et al. 2003; Mason et al. 2004b) predicts that, at least in SMC, CCN5 expression should be low in early development when cell proliferation and motility is high, and CCN5 expression should increase in some embryonic tissues later in development as cells stop proliferating and enter a more differentiated state. On the other hand, embryos develop in a very estrogen-rich environment, and CCN5 is strongly induced by estrogen (Banerjee et al. 2003; Fritah et al. 2006; Gray and Castellot 2005; Inadera et al. 2000; Mason et al. 2004a, b), thus predicting that CCN5 should be present in most or all embryonic tissues in response to this stimulus.
To resolve the alternative outcomes predicted by the anti-proliferative and estrogen-responsive nature of CCN5, we examined the expression patterns of CCN5 protein and mRNA and compared them to CCN2 protein expression patterns in developing mouse and human tissues. We observed widespread expression of CCN5 and CCN2 early in development that becomes increasingly restricted close to parturition. In general, we found that while CCN2 has a more restricted expression pattern than CCN5, many developing cell types and tissues express both proteins. These findings have important implications for the biologic and physiologic roles of CCN5.
Materials and methods
Animals
Animal protocols were reviewed and approved by The Institutional Animal Care and Use Committee at Tufts University (Boston, MA). C57BL/6J mice were obtained from Charles River Laboratories (Wilmington, MA). Food and water were available ad libitum. Mouse gestational ages were confirmed by comparison to a mouse development atlas (Kauffman 1992). Mice of gestational age 9–13 days post coitum are referred to as embryos (E9–E13). Older developing mice are referred to by gestational day (GD14–GD16).
Immunohistochemistry
Paraffin-embedded 7 μm sagittal sections of C57BL embryos and fetuses (FD Neurotechnologies, Baltimore, MD) and paraffin-embedded 5 μm sections from tissue cores in a human normal fetus tissue array (BE01014 and BE01015; US Biomax, Rockville, MD) were cleared with xylene. Endogenous peroxidase activity was quenched by treatment with two changes of 0.6% hydrogen peroxide (H2O2) in ethanol for 5 min. The slides were then rehydrated and treated with the Avidin/Biotin Blocking Kit (Vector Laboratories) in blocking serum (4% bovine serum albumin and 2% goat serum in phosphate-buffered saline) and then incubated in primary antibody in blocking serum overnight at 4°C. CCN5 protein was detected using a well characterized, highly specific, peptide affinity-purified rabbit polyclonal antibody to a polypeptide fragment from amino acids 103–117 of the von Willebrand Factor-C (VWC) domain of CCN5 (Gray and Castellot 2005; Lake et al. 2003; Lake and Castellot 2003; Mason et al. 2004a). This antibody is routinely used in the laboratory to follow expression of the full-length 27 kDa CCN5 protein on western blot (Gray and Castellot 2005; Lake et al. 2003; Lake and Castellot 2003; Mason et al. 2004a). CCN2 protein was detected using a well characterized, peptide affinity-purified rabbit polyclonal antibody to polypeptide fragment amino acids 223–348 from the thrombospondin-1 (TSP) and carboxy-terminal (CT) domains of mouse CCN2 (ab6992; Abcam, Cambridge, MA). Purified rabbit immunoglobulin IgG was used as a negative control (Biomeda, Foster City, CA). All negative controls completely lacked brown staining. The anti-CCN2 antibody has been used previously in numerous immunohistochemical studies (Candido et al. 2003; Dean et al. 2005; Finckenberg et al. 2003; Razzaque et al. 2003). Antigen retrieval by boiling slides in 10 mM citric acid pH 6.0 did not alter CCN5 or CCN2 staining or intensity in paraffin sections and thus was not performed in this study (data not shown). Previous reports (CCN2 reports listed above, Lake et al. 2003 for CCN5) of immunohistochemistry with these CCN2 and CCN5 antibodies on paraffin sections have also not employed antigen retrieval. Slides were developed using the Vectastain Elite ABC kit (Vector Laboratories, Burlingame, CA) and the 3,3′-diaminobenzidine (DAB) substrate kit (Vector Laboratories) and counterstained with Harris modified hematoxylin with acetic acid (Fisher). All slides were dehydrated and embedded in permanent mounting medium (#13510; DPX Mountant; Electron Microscopy Sciences; Hatfield, PA) and photographed using a microscope (Zeiss Axioscope) and a digital camera system (SPOT; Diagnostic Instruments). Antibody concentrations and substrate exposure times were carefully titrated to minimize artifacts and ensure that the staining intensities produced by both antibodies were similar. All directly compared images are from slides processed in a single experiment with a matched negative control (purified rabbit immunoglobulin IgG).
Reverse transcriptase PCR (RT-PCR)
Two pregnant female GD14.5 mice were sacrificed with carbon dioxide (CO2) overdose. Fetuses were dissected and immediately placed in RNA later (QIAGEN, Valencia, CA) and stored at −20°C. The dissected tissues were later removed from storage and 10 mg of each tissue was homogenized using a rotor/stator homogenizer (Fisher Scientific, Pittsburgh, PA). RNA isolation was performed using the RNeasy Mini kit (QIAGEN). DNA was removed using RQ1 RNase-Free DNase (Promega, Madison, WI), and reverse transcription was performed using the RETROscript kit (Ambion, Austin, TX). All assays were performed according to the manufacturer’s protocol. Control reactions with no reverse transcriptase were used to check for genomic DNA contamination in each sample. PCR was performed using the HotStarTaq Master Mix kit (QIAGEN) with 95°C 15 min polymerase activation step followed by 35 cycles of 94°C 30 sec/50 C 30 sec/72°C 1 min and final 72°C 10 min extension step and products were examined on a 1.5% agarose gel containing ethidium bromide. Primers were purchased from Integrated DNA Technologies (IDT, Coralville, IA). The sense CCN5 (GenBank Accession no. GI 4028578) primer consisted of the DNA sequence 5′-ATACAGGTGCCAGGAAGGTG-3′ (position 707–726), and the sequence of the anti-sense CCN5 primer was 5′-GTTGGATACTCGGGTGGCTA-3′ (position 913-932). After PCR, these primers produced a 225 bp (base pairs) DNA fragment that included the exon 4–5 (VWC-TSP) boundary in order to prevent amplification of genomic DNA sequence. The amplified DNA fragment was purifed by electrophoresis and the QIAquick Gel Extraction kit (QIAGEN) and sequenced by the Tufts University Core Facility (Boston, MA) to verify its identity. A plasmid containing mouse CCN5 cDNA was used as a positive control for PCR. Both water and mRNA not treated with reverse transcriptase enzyme were used as negative controls. PCR was also performed with the above conditions for the reference gene TATA box binding protein (Tbp) (GenBank Accession no. GI 2052376) with the following primers sense 5′-GCCTCTCAGAAGCATCACTA-3′ and anti-sense 5′-GCCAAGCCCTGAGCATAA-3′. PCR produced a 166 bp DNA fragment that included an exon–exon boundary (Willems et al. 2006).
Results
We determined the temporal and spatial expression pattern of CCN5 during embryonic and fetal development using a well-characterized anti-CCN5 antibody (Lake et al. 2003; Mason et al. 2004a). Mouse embryos and fetuses were obtained at time points ranging from E9 to GD16 and prepared as described in “Materials and methods”. Sections from human fetuses at 3 to 7 months of gestation were obtained and stained as described in “Materials and methods”. We compared the expression patterns of CCN5 with those of CCN2, a prototypical CCN family member containing all four domains. CCN2 exhibits a biological activity profile that is almost completely opposite that of CCN5 in smooth muscle cells (Ando et al. 2004; De Falco et al. 2006; Fan et al. 2000; Fan and Karnovsky 2002; Lake et al. 2003; Lake and Castellot 2003; Mason et al. 2002). No color reaction was detected when pooled rabbit IgG was used in place of primary antibody. Descriptions of staining intensity are intended for comparing protein levels detected for one protein only, not to compare the relative protein levels of CCN2 and CCN5. The two antibodies would not be expected to have identical affinities for their antigens and thus cannot be directly compared in this manner.
Early embryonic development
Immunohistochemistry results demonstrate that CCN5 is widely expressed throughout early embryonic development of the mouse. CCN5 was expressed in tissues of ectodermal, mesodermal, and endodermal origin (Fig. 1 and Supp. Fig. 1). In general, CCN2 and CCN5 staining was similar and detected in all embryonic organs at E9, E10, and E11 (Supp. Fig. 1 and data not shown; at high magnification specific staining is increased relative to negative control for both CCN2 and CCN5 in all organs including brain in E9 embryos). As organs further differentiated at E12 and later, some cell types increased and others decreased CCN5 expression. Beginning at E12, staining differences were detected. CCN5 was present in a larger number of cell types than CCN2 in GD14–16 mice.
CCN5 expression during E12–GD16 mouse development. Mouse embryonic and fetal sagittal sections from a E12, b GD14 and c GD16 developing mice. Negative control (c right) – pooled rabbit IgG applied in place of primary antibody. Bl = Bladder; Br = Brain; He = Heart; Int = Intestine; K = Kidney; Li = Liver; Lu = Lung; To = Tongue. a,b CCN2 left, CCN5 right c CCN2 left, CCN5 middle, Control right. Scale Bars = 1 mm. Scale Bars and labels on CCN5 panels also apply to CCN2 and IgG panels. CCN5 and CCN2 immunoreactivity shown by brown staining. Nuclei are shown by hematoxylin (blue) staining. Tissue sections treated with pooled rabbit IgG gave no detectable staining (c and data not shown)
In the sections below, we present comparisons of CCN5 and CCN2 expression patterns in mouse and human tissues of the major organ systems at several time points in embryonic and fetal development.
Cardiovascular system
Mouse
The cardiovascular system is the first functional organ system in developing mammalian embryos. CCN5 was abundantly expressed in the endothelium and smooth muscle of veins and arteries, while CCN2 was detectable at very low levels (Figs. 2a, 4b, and Supp. Fig. 2a,b). Higher levels of both CCN2 and CCN5 were detected in the apex of the developing heart during E12–GD14 when compared to myocardial expression levels. At GD16, CCN5 expression in myocardium was uniform. In contrast, at GD16 CCN2 expression was decreased throughout the myocardium relative to GD14 expression (Fig. 2b–d, Supp. Fig. 2c). Neither CCN2 nor CCN5 was concentrated at the apex at GD16 (data not shown). At E12, CCN5 was present at lower levels in the forming atrioventricular septum than in myocardium which demonstrated uniform levels of CCN5 (Fig. 2b). CCN5 levels were very low and CCN2 was absent in the developing cardiac valves (Fig. 2d). At GD14, CCN5 levels were lower in aorta when compared to the pulmonary arterial trunk (Fig. 2d). Mouse umbilical vessels expressed CCN5 and CCN2 (Fig. 2e).
CCN5 expression during cardiovascular development. a GD16 artery from lung. b E12 mouse heart. c,d GD14 mouse heart. e GD14 mouse umbilical vessels. f Human 5 month myocardium and coronary vessels. Negative control (f right) – pooled rabbit IgG applied in place of primary antibody. A=Artery; Ao = Aorta; CV = Coronary Vessels; PA = Pulmonary Artery; V=Vein; empty right arrow = Atrioventricular Septa; filled triangle = apex; empty triangle = Valve. a–e CCN2 left, CCN5 right. f CCN2 left, CCN5 middle, Control right. Scale Bar = 10 μm (a) Scale Bars = 100 μm (b–f). Tissue sections treated with pooled rabbit IgG gave no detectable staining (f and data not shown)
Human
CCN5 was detected throughout the human fetal myocardium, endothelium and smooth muscle of coronary arteries and veins and to a lesser extent in other large vessels at 4 months of gestation (Fig. 2f, Supp. Fig. 2d). CCN2 expression was low in these fetal cardiovascular tissues. The smooth muscle and endothelium of human umbilical vessels displayed a high level of CCN2 and a moderate level of CCN5 (Supp. Fig. 2e).
Respiratory system
Mouse
Lung development between GD14–GD16 is considered the pseudoglandular period, corresponding to weeks 5–17 in human development (Van Tuyl and Post 2003; Volpe et al. 2003). CCN5 was present in GD14 and GD16 bronchial epithelium and in GD16 mesenchymal cells surrounding branching bronchioles (Fig. 3a–c). In contrast, CCN2 was present only in the bronchiolar epithelium and not in the surrounding mesenchyme. Some terminal ends of actively branching bronchioles displayed very high levels of CCN2 at GD14 (Fig. 3a). At GD16, some bronchiolar terminal ends demonstrated little or no CCN2 and others expressed higher levels of CCN2, while all bronchiolar terminal ends expressed CCN5 (Fig. 3b). CCN5 was present in the epithelial and mesenchymal cell layers of larger bronchioles, and CCN2 was absent (Fig. 3c). CCN2 was present mostly in actively growing and branching bronchi, but not in some terminal ends of actively branching bronchioles and in the bronchiolar epithelium after larger bronchioles are established. In contrast, CCN5 was expressed both in the larger bronchioles and throughout actively branching bronchioles.
CCN5 expression during lung development. a GD14 mouse lung. Inset shows staining in epithelial cells of bronchiole. b GD16 mouse lung actively branching bronchioles. c GD16 larger bronchi that are no longer branching. d Human 5 month fetal lung. ep = Epithelial Cells me = Mesenchymal Cells. a–c CCN2 left, CCN5 right. Scale Bars = 50 μm (a,b,c). Scale Bars = 100 μm (d). Tissue sections treated with pooled rabbit IgG gave no detectable staining (data not shown)
Human
Fetal lung tissue at 5 months of development demonstrated CCN5 at low but uniform levels in epithelial and mesenchymal cells, and no CCN2 staining (Fig. 3d). At this time in development (the canalicular period), the lung has finished forming the bronchial tree and is forming the acini, capillary network, and alveolar type I and II cells (Wert 2004).
Musculoskeletal system
Mouse
CCN5 was detected in GD14–16 mouse skeletal muscle (Fig. 4a–b). CCN5 staining was particularly strong at myotendinous junctions, beginning in GD14 and becoming pronounced by GD16. In contrast, GD14–16 skeletal muscle displayed very little CCN2 and it did not accumulate at myotendinous junctions. Both CCN2 and CCN5 staining was present in chondrocytes during endochondral ossification (Fig. 4b–c). Hyaline cartilage in the resting zones displayed neither CCN2 nor CCN5. CCN5 was present in the chondrocytes of the proliferating, hypertrophic, and calcification zones, and CCN2 was seen primarily in the chondrocytes of the hypertrophic and calcification zones. In addition, perichondrial cells expressed CCN5 (Fig. 4c).
CCN5 expression in developing bones and skeletal muscle. a GD14 skeletal muscle. b GD16 skeletal muscle and bone. c GD16 rib. d Human 5 month fetal bone osteoclast. art = arteriole; CZ = Calcification Zone; HC = Hypertrophic Chondrocytes; MyoJxn = Myotendinous Junctions; PZ = Proliferating Zone; RZ = Resting Zones; SkMusc = Skeletal Muscle. a–d CCN2 left, CCN5 right. Scale Bars = 100 μm (a,b,c). Scale Bars = 20 μm (d). Tissue sections treated with pooled rabbit IgG gave no detectable staining (data not shown)
Human
In the human fetus at 5 months of gestation, CCN5 was detected at a low level in skeletal muscle (Supp. Fig. 3a). The human fetal samples analyzed did not include myotendinous junctions. CCN2 was not detected in human fetal osteocytes or osteoclasts (Fig. 4d and Supp. Fig. 3b). In contrast, CCN5 staining was detected in osteoclast nuclei and absent in osteocytes (Fig. 4d and Supp. Fig. 3b).
Gastrointestinal system
Mouse
In GD14–16 mouse liver, both CCN2 and CCN5 staining were detected in hepatocytes but not hematopoietic stem cells (Fig. 5a and data not shown). Neutrophils express CCN5 but not CCN2.
CCN5 expression during intestinal development. a GD16 mouse liver. b E12 mouse intestine. c GD14 mouse intestine. d GD16 mouse intestine (longitudinal section). e Human 4 month fetal liver. f Human 5 month fetal liver. g Human 6 month fetal esophagus. h Human 5 month fetal colon. EP = Intestinal Epithelium; Hep = Hepatocytes; HSC = Hematopoietic Stem Cells; Neu = Neutrophil; SM = Intestinal Smooth Muscle. a–h CCN2 left, CCN5 right. Scale Bars = 50 μm (a,e,f). Scale Bars = 100 μm (b–d, g–h). Tissue sections treated with pooled rabbit IgG gave no detectable staining (data not shown)
The developing small intestine of the mouse expressed both CCN2 and CCN5. At E12 (Fig. 5b) CCN2 staining was highly expressed in intestinal epithelium, lower at GD14 (Fig. 5c), and then much weaker by GD16 (Fig. 5d). CCN2 staining was very low in intestinal smooth muscle throughout all of the time points examined. CCN5 was detected at low levels in intestinal smooth muscle and epithelium beginning at E12 (Fig. 5b), at moderate levels at GD14 (Fig. 5c), and at intermediate levels in GD16 fetuses (Fig. 5d).
Human
Human fetal hepatocytes at 4 months of development revealed moderate CCN2 and CCN5 staining (Fig. 5e). However, at 5 months of gestation, fetal hepatocytes revealed lower levels of CCN2 and higher levels of CCN5 than observed at 4 months (Fig. 5f). Liver hematopoietic stem cells do not express CCN5 or CCN2.
Analysis of CCN2 and CCN5 protein expression patterns in the developing human intestinal tract revealed a gradation of expression among different sections of the digestive tract. CCN2 and CCN5 were present at moderate levels in the esophagus at 6 months (Fig. 5g); in this structure both proteins were detected in the muscularis mucosae and muscularis externa. CCN2 and CCN5 were also expressed in the esophageal epithelium, and absent in the lamina propria. Other regions of the human fetal intestine were examined at 5 months. CCN5 was detected in the stomach musculature and at low levels in the epithelium and lamina propria at 5 months; CCN2 staining was absent in the stomach (Supp. Fig. 4a). CCN5 staining, but not that of CCN2, was detected in the smooth muscle and intestinal epithelium of the small intestine at 5 months (Supp. Fig. 4b). CCN5 staining was present uniformly throughout the gallbladder smooth muscle and epithelium, and CCN2 staining was absent (Supp. Fig. 4c). CCN5 staining was detected in the smooth muscle of the colon (Fig. 5h), and CCN2 was low except for the first few layers of intestinal smooth muscle that face the lumen. Both CCN2 and CCN5 were absent in the base of the intestinal crypts (the location of proliferating colonic epithelium) and highly expressed in the tips of the villi (the cells that have moved away from the crypts and are no longer proliferating). CCN5 was detected at very low levels in the colon lamina propria, and CCN2 was absent (Fig. 5h). In the rectum, CCN5 was detected at low levels in smooth muscle, lamina propria, and non-proliferating epithelium; CCN2 was not present in these tissues (Supp. Fig. 4d).
Kidney and urogenital system
Mouse
In early mouse kidney development (E12), metanephric tubules express CCN5 and CCN2 (Fig. 6a) in tubule terminals rather than uniformly throughout the tubule cells. CCN5 was expressed in glomerular mesangial cells, tubules, blood vessels, and collecting ducts, while CCN2 was detected only in tubules at GD 16 (Fig. 6b–d, Supp. Fig. 2b). Glomerular endothelial cells do not display any CCN5, in contrast to endothelial cells of larger vessels. GD14 mouse bladder smooth muscle, mesenchyme, and urothelium displays CCN5 but CCN2 was only detected in the urothelium (Fig. 6d). In the mouse bladder at GD16, the urothelium and smooth muscle display CCN2 and CCN5 (Fig. 6e).
CCN5 expression in developing urogenital system. a E12 mouse kidney. b,c GD16 mouse kidney. d GD14 mouse bladder. e GD16 mouse bladder. f GD14 mouse ovary. g GD16 mouse ovary and reproductive tract. h Human 5 month fetal kidney. i Human 5 month placenta. j Human 5 month fetal testis. k Human 5 month fetal uterus. l Human 5 month fetal ovary. m Human 5 month fetal fallopian tube. n Human 5 month fetal epididymis. filled triangle = High or empty triangle = low CCN protein expression in metanephric tubules. Br = branch point of chorionic villi with absent CCN5 staining; CD = collecting ducts; DT = distal tubules; E = glomerular endothelial cells; GE = germinal epithelium; Le = Leydig cells; M = mesangial cells; Oo = oocytes; SM = smooth muscle; Tr = trophoblasts; Ur = urothelium. a–h CCN2 left, CCN5 right. i–n CCN5. Scale Bars = 50 μm. Tissue sections treated with pooled rabbit IgG gave no detectable staining (data not shown)
The germinal epithelium and ovary reveal CCN5 staining at GD14 (Fig. 6f). Higher levels of CCN5 staining and low levels of CCN2 staining were found in the ovary and epithelial cells and glands of the reproductive tract at GD16 (Fig. 6g).
Human
In the human fetal kidney at 5 months, CCN5 staining was detected in mesangial cells, tubules, and glomeruli (Fig. 6h). CCN2 expression was very low in these structures.
Low levels of CCN5 staining were detected in the developing human reproductive system at 5 months, with the exception of high levels in placental trophoblasts (Fig. 6i). CCN5 staining was absent in regions where the chorionic villi fused together and branched. Weak CCN5 staining was detected in testicular Leydig cells (Fig. 6j), uterus (Fig. 6k), ovarian stroma (Fig. 6l), fallopian tube (Fig. 6m), and epididymis (Fig. 6n). In contrast, CCN2 was absent in all developing human reproductive organs examined (data not shown).
Endocrine and immune organs
Mouse
In GD16 mouse thyroid gland (Fig. 7a), very low levels of CCN2 and moderate levels of CCN5 were observed. CCN5 staining was detected on the colloidal side of developing thyroid follicles, suggesting that the thyroid follicle cells produce and excrete CCN5. Both CCN5 and CCN2 were detected in the connective tissue of the adrenal capsule (Fig. 7b). CCN5 staining was observed in the nuclei of some adrenal cells (Fig. 7b, inset). The pancreas in developing mice displayed low levels of CCN5 and CCN2 in the exocrine ducts. CCN2 and CCN5 levels were extremely low or absent in thymus and spleen (data not shown).
CCN5 expression in endocrine organs. a Mouse GD16 thyroid gland. b Mouse GD16 adrenal gland. Insets magnified to show nuclear CCN5 staining in adrenal gland cells. c Human 5 month fetal thyroid gland. d Human 5 month fetal adrenal gland. F = thyroid follicle; ZF = zona fasciculata; ZG = zona glomerulosa. a–d CCN2 left, CCN5 right. Scale Bars = 100 μm (a,b,d). Scale Bars = 25 μm (c). Tissue sections treated with pooled rabbit IgG gave no detectable staining (data not shown)
Human
In 5 month human fetal thyroid, (Fig. 7c), moderate levels of CCN5 were observed, and CCN2 was not detectable. In the 5 month human fetal adrenal gland, CCN5 was present in the zona fasciculata, the region of the gland that makes cortisol, and very low in the zona glomerulosa, the region that produces aldosterone (Fig. 7d). No CCN2 staining was observed in the human fetal adrenal gland. Human fetal pancreas demonstrated only CCN5, and not CCN2, in the exocrine ducts. Very little CCN2 or CCN5 was detected in fetal human thymus and spleen (data not shown).
Nervous system and skin
Mouse
Both CCN2 and CCN5 were widely distributed in the brain of E12 mouse embryos (Fig. 8a). Expression of both proteins was much more restricted in the later GD14 (Fig. 8b) and GD16 fetuses (Fig. 8c). Many cells in the developing mouse brain and spinal cord demonstrated CCN2 and CCN5 predominantly and strongly in the nucleus of neurons (Fig. 8d), in contrast to most cell types where CCN5 is found primarily in cytoplasm or tightly bound to the outer surface of the cell membrane. Both CCN5 and CCN2 were present in the choroid plexi, particularly at GD16 (Fig. 8b–c).
CCN5 expression in developing brain, hair, skin, and eyes. a E12 mouse brain. b GD14 mouse brain. c GD16 mouse brain. d GD16 spinal cord. e GD16 mouse whisker. f GD16 mouse skin. g GD16 mouse eye. h Human 5 month fetal hair follicle. i Human 5 month fetal skin. j Human 4 month fetal retina Af = afferent nerve fibers; C = choroid; IRS = inner root sheath; ORS = outer root sheath; PR = photoreceptors. a–k CCN2 left, CCN5 right. Scale Bars = 100 μm (a–f, i,j). Scale Bars = 50 μm (g,h). Tissue sections treated with pooled rabbit IgG gave no detectable staining (data not shown)
Both CCN2 and CCN5 were present in the developing GD16 mouse vibrissae (Fig. 8e). CCN5 was observed in both the inner and outer root sheath layer of cells, and CCN2 was limited to the inner sheath. Both CCN2 and CCN5 were detected at high levels in the developing epidermis of GD16 mouse (Fig. 8f).
In the mouse eye at GD16, CCN5 and CCN2 were detected in the fused eyelid and corneal epithelium (Fig. 8g). CCN5, and not CCN2, was detected in the mouse corneal stroma.
Human
Neither CCN5 nor CCN2 was found in the human fetal brain at 4 months (data not shown). This may be due to species differences between mouse and human. Alternatively, because only cores of tissue were examined it is possible that CCN2 or CCN5 was present in the developing human brain in other regions or at other time points not examined in this study.
Both CCN2 and CCN5 were present in human fetal hair follicles at 5 months (Fig. 8i), with CCN5 found in both the inner and outer root sheath layer of cells and CCN2 limited to the inner sheath. Both CCN2 and CCN5 were detected a high levels in the developing epidermis of 5 month human fetal skin (Fig. 8i). In the human fetal eye at 4 months, the rod/cone photoreceptor segment of the retina displayed high levels of CCN5, but not CCN2 (Fig. 8j). Low levels of both CCN2 and CCN5 were detected in the afferent nerve fibers. We also noted very lightly stained material just under the retinal pigmented epithelium in what appears to be the cytosol of the retinal pigmented epithelia. This structure appears very early on in development (as do the retinal ganglia cells), and thus this observation fits with our hypothesis that CCN5 may limit proliferation of these two layers as the photoreceptors are usually in a highly proliferative stage at this point in development.
Expression of CCN5 mRNA
Immunohistochemical analysis provides the most accurate representation of the location of the CCN5 protein. It is possible that this proteins is translated, secreted, and then distributed to its target cells and tissues. We compared the sites of CCN5 protein translation and its final distribution by analyzing mouse fetal tissues for CCN5 mRNA. Organs from GD14.5 mice were collected and mRNA prepared for analysis by RT-PCR. CCN5 mRNA was found in every fetal organ examined, including lung, limbs and tails, umbilical cord, intestine, heart, liver, carcass, and head (Fig. 9). CCN5 mRNA was also demonstrated in the maternal placenta. We previously demonstrated CCN5 mRNA in the intact rat uterus and aorta and in cultured human uterine SMC and rat aortic SMC and endothelial cells (Delmolino et al. 2001; Mason et al. 2004a). These results suggest that the observed immunohistochemical findings in GD14 mice are the result of local synthesis of CCN5 mRNA, rather than synthesis in one organ and subsequent protein distribution to other sites.
CCN5 mRNA is expressed in all GD14.5 fetal mouse organs. RNA was harvested from pooled GD14.5 C57BL/6 mouse organs, reverse transcribed, and PCR was performed using primers specific to mouse CCN5 or Tbp (loading control) on mRNA treated with (+RT) or without (−RT) reverse transcriptase. (negative signs) no template control (positive signs) plasmid containing mouse CCN5 cDNA (a) maternal uterus (b) placenta (c) lung (d) limbs and tails (e) umbilical cord (f) intestine (g) heart (h) liver (i) carcass (j) head (k) maternal aorta
Discussion
In this report, we comprehensively examined the distribution pattern of CCN5 and CCN2 protein during embryonic and fetal development of the mouse, comparing it to the expression pattern in human fetal tissues. We examined CCN5 mRNA levels in selected organs in the GD14.5 mouse, and provide a comparison of all of the currently available data from similarly comprehensive studies on developing mammalian mRNA and protein distribution for all of the CCN family members (Table 1). We found that CCN5 protein is present in most or all cells in early embryos, and then tissue-specific CCN5 expression differences appear as embryos age. CCN5 expression was not restricted to cell types of a particular embryonic germ layer lineage.
While the hypothesis that high estrogen levels cause high CCN5 expression in embryonic tissues is attractive, at least two lines of evidence suggest that this is not the entire explanation. First, numerous cell types in GD16 fetuses produce little or no CCN5 protein when high estrogen levels are present (17-β estradiol levels in mouse pregnancy are stable at 15–25 pg/ml between days 2–16) (McCormac and Greenwal 1974). Second, despite several independent attempts by both our group and other laboratories to produce CCN5 knockout mice, no laboratory has produced either hemizygous or homozygous CCN5 knockout animals, suggesting that CCN5 is necessary for early development. Furthermore, primary human bone-marrow derived mesenchymal stem cells initially express high levels of CCN5, and then express CCN5 at high, medium, or low levels respectively as the cells undergo differentiation into osteogenic, chondrogenic, and adipogenic lineages (Schutze et al. 2005). This mirrors our observation that the less differentiated cells of early embryos express high levels of CCN5, and as cells further differentiate, CCN5 expression levels generally decline and vary among different cell types. Thus, CCN5 is not a general marker of quiescence and maturation as originally predicted due to our observations of SMC expression patterns, but increases with differentiation in some cell types and decreases in others.
Though this and other published reports generally agree on the distribution of CCN2 in most developing tissues, there are several discrepancies among published reports of CCN2 protein or mRNA expression that remain unresolved (Table 1). Discrepancies between ISH and IHC data can occur for a number of reasons, including post-transcriptional modifications that alter mRNA translation or stability, and post-translational processing that alters protein maturation, accumulation, or degradation. Discrepancies in mRNA or protein distribution may be due to differences in sensitivity of the methods. For example, Kireeva et al (1997) used a peroxidase-conjugated secondary antibody and found less CCN2 in many organ systems than we and Surveyor and Brigstock (1999) did using the more sensitive avidin–biotin technique. Discrepancies may also be due to differences in the affinity, avidity, or epitope availability of the specific CCN2 antibodies used, as well in the specific fixation techniques used. CCN2 distribution in our study was mapped with an antibody against amino acids 223–348 of the CCN2 protein, while previous efforts employed antibodies against amino acids 165–200 (Kireeva et al. 1997) and amino acids 80–93 (Surveyor and Brigstock 1999) of the CCN2 protein. We have also used this latter antibody and see the same CCN2 expression pattern as we found with the aa223–348 antibody testing identical tissues and fixation conditions.
We initially hypothesized that CCN2 and CCN5 would have complementary expression patterns. With a few exceptions, we found that cell types expressing CCN2 also express CCN5. However, CCN5 was highly expressed in many cells where CCN2 expression was low or absent, for example in developing skeletal muscle myotendinous junctions. We conclude that while in certain disease systems CCN2 increases in pathologies where CCN5 decreases that this pattern is not generally true for developing human and mouse tissues.
We observed CCN5 protein in the SMC and endothelial cells of large vessels, as was expected due to previous observations that both CCN5 protein and mRNA are present in vascular smooth muscle cells and endothelium of aorta and other larger vessels (Delmolino et al. 2001; Gray and Castellot 2005; Lake et al. 2003). Although we predicted that CCN5 levels would be low in developing vascular smooth muscle because the embryo is actively growing, we found that CCN5 was consistently expressed in VSMC, suggesting that the carefully controlled cell proliferation required for embryonic growth does not require an environment with low CCN5 levels. We also observed lower CCN5 expression in the aorta relative to that seen in the pulmonary artery, suggesting that CCN5 protein levels in VSMC may be modulated by exposure to different mechanical forces or arterial pressure as has been described for CCN1 (Grote et al. 2004).
In the developing lung, the variable expression of CCN2 in terminal bronchiolar segments may represent, respectively, terminal buds that are either actively growing and creating new segments or segments that have stopped proliferating. The presence of CCN5 in mesenchymal cells suggests that it may be important for signaling epithelial growth, such as defining barriers or edges of the bronchiole. Although CCN5 was expressed in both mesenchymal and epithelial cells of the developing airway, CCN2 was observed only in the epithelial cells. The interactions between epithelial and mesenchymal cells are critical in airway formation and differentiation (Evans et al. 1999; Van Tuyl and Post 2003). In addition, CCN5 was present in larger bronchi, and no CCN2 was present. The primary bronchioles and major airways form earlier in development and are thus relatively quiescent at this stage (Van Tuyl and Post 2003). The presence of high levels of CCN2 in some terminal bronchiolar buds but not others suggests that CCN2 might be required only in actively branching bronchioles, and the continued presence of CCN5 throughout both actively branching and stable bronchioles suggests it might be required throughout their development.
In the musculoskeletal system, it is interesting to note that the pattern of integrin αv protein expression is concentrated at the extremities of apical myotubes marking their shape (Hirsch et al. 1994; Tarone et al. 2000). The pattern of CCN5 expression was similar, suggesting that it may participate in the organization of the myotendinous junction. Integrin binding has been demonstrated in other CCN family members and has been implicated in angiogenesis and matrix attachment (reviewed in Lau and Lam 1999; Lau and Lam 2005; Leask and Abraham 2006). Our findings suggest that integrin αv may be important for CCN5 signaling.
In contrast to previous observations in 2 week old rats, we did not detect CCN2 in osteocytes, suggesting that the role of CCN2 in this cell type may not emerge until bones have fully formed (Safadi et al. 2003). This may represent a species or age difference in CCN2 expression; however, because only a small core of human fetal bone was examined, it cannot be concluded that CCN2 is not expressed in fetal human bone osteocytes. CCN2 expression in hypertrophic chondrocytes in both fetal and healing cartilage and bone is thought to feed back on proliferating chondrocytes to induce their maturation (Kubota and Takigawa 2007; Takigawa et al. 2003; Takigawa et al. 2005; Yamaai et al. 2005). The presence of CCN5 in the proliferating and calcification zones of ossifying bones suggests that it has an important role in ossification. The expression pattern we observe is similar to that reported by Kumar et al. (1999). The opposite expression patterns of CCN2 and CCN5 in hypertrophic chondrocytes suggest that these proteins may be coordinately regulated during ossification.
We observed a change from patchy to uniform CCN5 expression in the liver as gestational age increased. This may reflect the change of liver function as a hematopoietic organ in younger fetuses to a non-hematopoietic organ in 5 month old fetuses (Tavian and Peault 2005). At GD16 in mice, approximately 40% of the liver volume is dedicated to hematopoiesis (Dzierzak and Medvinsky 1995). It is noteworthy that liver hematopoiesis remains important throughout mouse fetal development and is present at much lower levels in the 5 month human fetus, consistent with the hypothesis that loss of hepatic CCN2 expression correlates with the loss of hematopoiesis in human fetal tissue. CCN2 protein and mRNA expression has been reported to be low but present in normal adult rat tissue (Li et al. 2006).
CCN2 protein and mRNA expression has been reported to be low in intestinal smooth muscle unless there is a fibrotic stimulus such as TGF-β, radiation treatment, or Crohn’s disease (Beddy et al. 2006; Haydont et al. 2005; Vozenin-Brotons et al. 2003). In our study CCN2 was initially highly expressed in intestinal development and expression was reduced to lower levels in older human and mouse intestine. Because of our previous studies of the differential expression of CCN5 relative to growth state of the cell, we expected to find that rapidly proliferating cells would not express high levels of CCN5, and quiescent or growth-arrested cells would express large amounts of CCN5. This pattern is present in the colonic epithelium, where the proliferating cells in the crypts of the villi did not express CCN5, but in non-proliferating cells in the villi tips CCN5 levels were high. However, in contrast to the situation observed for vascular SMC, in which CCN2 is expressed in proliferating but not quiescent cells and CCN5 is the opposite expression pattern (Fan et al. 2000; Lake et al. 2003), both CCN2 and CCN5 were expressed in a similar pattern in the colonic epithelium. This suggests that although CCN5 and CCN2 both are needed for cell growth control, they are not necessarily regulated in a complementary fashion in all cell types and key mechanistic differences may exist in CCN protein biology between mesenchymal and epithelial cells.
In the kidney and urogenital system, the terminal buds of kidney metanephric tubules express both CCN2 or CCN5 terminal ends of tubules but not throughout the tubules, suggesting that these proteins may work in concert to regulate terminal bud proliferation and branching. The lack of CCN5 in the endothelial cells of the glomerular tufts suggests that CCN5 has a different function in the tightly coupled endothelial cells of large vessels compared to those of the more permeable glomerular tufts. CCN2 mRNA and protein expression has been reported to be low but present in healthy adult kidney (Riser et al. 2005).
We detected high levels of CCN2 and CCN5 throughout GD16 skin and oral epithelium. Kireeva et al. detected CCN2 in skin during early gestation and in the dermis of tongue and tail, and Surveyor and Brigstock detected CCN2 only in oral epithelium (see Table 1) (Kireeva et al. 1997; Surveyor and Brigstock 1999). This discrepancy might be caused by differences in antibody sensitivity. The antibody that detected CCN2 both in mouse and human fetal skin in this report also detects CCN2 in normal adult human skin (Carulli et al. 2005).
CCN5 has been observed to be expressed in secretory structures, including in the endometrial glands (Mason et al. 2004a), thus it is not unexpected that CCN5 protein was found in most secretory glands including the exocrine pancreas and thyroid. CCN5 is expressed in many steroid-secreting cells during human development, including human testicular Leydig cells and the cortisol-secreting cells of the adrenal glands. The low expression levels of CCN5 in fetal uterine smooth muscle and glands was in agreement with the hypothesis that CCN5 protein increases as smooth muscle cells differentiate because it is expressed at high levels in both adult human and rat uterine tissue (Mason et al. 2004a, b). Although CCN5 is highly expressed in response to high estrogen levels in the adult, this tissue may not have the ability to regulate CCN5 in response to estrogen levels until sexual maturity is reached.
In most cell and tissue types, CCN5 expression was observed primarily in the cytoplasm and periphery of cells as previously observed in cultured vascular SMC (Lake et al. 2003). In addition, we also observed nuclear localization of CCN5 in some cells. Nuclear localization was particularly prominent in the cells of the spinal cord and adrenal gland. Both CCN2 and CCN3 have been detected in the nucleus and are likely to help regulate transcription (Gellhaus et al. 2004; Perbal 1999; Wahab et al. 2001). Amino-truncated but not full length CCN3 has been detected in cell nuclei, and CCN3–Gal4 fusion proteins can reduce transcription of reporter genes with Gal4 binding sites (Perbal 1999). CCN3 has been shown to interact with the rpb7 subunit of RNA polymerase in a yeast two hybrid system (Perbal 1999) However, the C-terminal domain of CCN3 has been identified as the responsible module for both nuclear localization because it contains an NLS and also transcription repression (Planque et al. 2006). As CCN5 lacks the C-terminal module this suggests that CCN5 may be locating to the nucleus by a novel mechanism (Planque et al. 2006). The variation in subcellular localization of CCN5 among different cell types supports the idea that CCN5 may have several different functions within the cell.
The unexpectedly broad expression pattern of CCN5 in most embryonic and fetal tissues, combined with its increasingly tissue-specific expression with developmental age, suggests cell- and tissue-specific functions for CCN5. Furthermore, we observe CCN5 expression on the cell periphery, in the cytoplasm, and in the nucleus. These findings point to a complex set of conditions and parameters that are likely to regulate CCN5 and its biological roles, including growth state, hormonal influences, extracellular milieu, cell cycle position, reproductive status, developmental stage, contact/density, cytokine exposure, adhesion, disease, and other influences. The finding that CCN5, like other CCN proteins, is localized in the nucleus represents another layer of complexity and leads to the hypothesis that CCN5 regulates transcription of genes controlling cell proliferation and motility. Furthermore, while our data rule out the original premise that CCN5 and CCN2 are generally expressed in coordinately regulated fashion, it still leaves open the possibility that in certain tissues, notably bone and cartilage, these CCN proteins may work in concert to regulate development and function.
Efforts to elucidate the mechanisms and molecules regulating the developmental and physiologic role(s) of CCN5 are underway in several laboratories. The results presented in this communication should provide a firm foundation for additional hypothesis generation and interpretation of data obtained by other experimental approaches.
Abbreviations
- CCN:
-
Cysteine-rich 61/Connective Tissue Growth Factor/Nephroblastoma-overexpressed family of proteins
- CT:
-
carboxy-terminal
- DAB:
-
3,3′-diaminobenzidine
- E9:
-
embryonic day 9
- GD16:
-
gestational day 16
- IHC:
-
immunohistochemistry
- ISH:
-
in situ hybridization
- RT-PCR:
-
reverse transcriptase polymerase chain reaction
- SMC:
-
smooth muscle cells
- Tbp:
-
TATA box binding protein
- TSP:
-
thrombospondin-1
- VWC:
-
von Willebrand Factor-C
References
Ando H, Fukuda N, Kotani M, Yokoyama S, Kunimoto S, Matsumoto K, Saito S, Kanmatsuse K, Mugishima H (2004) Chimeric DNA–RNA hammerhead ribozyme targeting transforming growth factor-[beta]1 mRNA inhibits neointima formation in rat carotid artery after balloon injury. Eur J Pharmacol 483(2–3):207–214
Banerjee S, Saxena N, Sengupta K, Tawfik O, Mayor MS, Banerjee SK (2003) WISP-2 gene in human breast cancer: estrogen and progesterone inducible expression and regulation of tumor cell proliferation. Neoplasia 5(1):63–73
Beddy D, Mulsow J, Watson RWG, Fitzpatrick JM, O’Connell PR (2006) Expression and regulation of connective tissue growth factor by transforming growth factor beta and tumour necrosis factor alpha in fibroblasts isolated from strictures in patients with Crohn’s disease. Br J Surg 93(10):1290–1296
Candido R, Forbes JM, Thomas MC, Thallas V, Dean RG, Burns WC, Tikellis C, Ritchie RH, Twigg SM, Cooper ME, Burrell LM (2003) A breaker of advanced glycation end products attenuates diabetes-induced myocardial structural changes. Circ Res 92(7):785–792
Carulli MT, Ong VH, Ponticos M, Shiwen X, Abraham DJ, Black CM, Denton CP (2005) Chemokine receptor CCR2 expression by systemic sclerosis fibroblasts: evidence for autocrine regulation of myofibroblast differentiation. Arthritis Rheum 52(12):3772–3782
Cervello M, Giannitrapani L, Labbozzetta M, Notarbartolo M, D’Alessandro N, Lampiasi N, Azzolina A, Montalto G (2004) Expression of WISPs and of their novel alternative variants in human hepatocellular carcinoma cells. Ann N Y Acad Sci 1028:432–439
Dean RG, Balding LC, Candido R, Burns WC, Cao ZM, Twigg SM, Burrell LM (2005) Connective tissue growth factor and cardiac fibrosis after myocardial infarction. J Histochem Cytochem 53(10):1245–1256
De Falco M, Staibano S, D’Armiento FP, Mascolo M, Salvatore G, Busiello A, Carbone IF, Pollio F, Di Lieto A (2006) Preoperative treatment of uterine leiomyomas: clinical findings and expression of transforming growth factor-beta 3 and connective tissue growth factor. J Soc Gynecol Investig 13(4):297–303
Delmolino LM, Stearns NA, Castellot JJ (1997) Heparin induces a member of the CCN family which has characteristics of a growth arrest specific gene. Mol Biol Cell 8:287a
Delmolino LM, Stearns NA, Castellot JJ (2001) COP-1, a member of the CCN family, is a heparin-induced growth arrest specific gene in vascular smooth muscle cells. J Cell Physiol 188(1):45–55
Dzierzak E, Medvinsky A (1995) Mouse embryonic hematopoiesis. Trends Genet 11(9):359–366
Evans MJ, Van Winkle LS, Fanucchi MV, Plopper CG (1999) The attenuated fibroblast sheath of the respiratory tract epithelial–mesenchymal trophic unit. Am J Resir Cell Mol Biol 21(6):655–657
Fan WH, Karnovsky MJ (2002) Increased MMP-2 expression in connective tissue growth factor over-expression vascular smooth muscle cells. J Biol Chem 277(12):9800–9805
Fan WH, Pech M, Karnovsky MJ (2000) Connective tissue growth factor (CTGF) stimulates vascular smooth muscle cell growth and migration in vitro. Eur J Cell Biol 79(12):915–923
Finckenberg P, Inkinen K, Ahonen J, Merasto S, Louhelainen M, Vapaatalo H, Muller D, Ganten D, Luft F, Mervaala E (2003) Angiotensin II induces connective tissue growth factor gene expression via calcineurin-dependent pathways. Am J Pathol 163(1):355–366
French DM, Kaul RJ, D’Souza AL, Crowley CW, Bao M, Frantz GD, Filvaroff EH, Desnoyers L (2004) WISP-1 is an osteoblastic regulator expressed during skeletal development and fracture repair. Am J Pathol 165(3):855–867
Friedrichsen S, Heuer H, Christ S, Winckler M, Brauer D, Bauer K, Raivich G (2003) CTGF expression during mouse embryonic development. Cell Tissue Res 312(2):175–188
Fritah A, Redeuilh G, Sabbah M (2006) Molecular cloning and characterization of the human WISP-2/CCN5 gene promoter reveal its upregulation by oestrogens. J Endocrinol 191(3):613–624
Fukutomi T, Zhou YH, Kawai S, Eguchi H, Wands JR, Li JS (2005) Hepatitis C virus core protein stimulates hepatocyte growth: correlation with upregulation of wnt-1 expression. Hepatology 41(5):1096–1105
Gellhaus A, Dong X, Propson S, Maass K, Klein-Hitpass L, Kibschull M, Traub O, Willecke K, Perbal B, Lye SJ, Winterhager E (2004) Connexin43 Interacts with NOV: a possible mechanism for negative regulation of cell growth in choriocarcinoma cells. J Biol Chem 279(35):36931–36942
Gray MR, Castellot JJ (2005) Function and regulation of CCN5. In: Perbal BV, Takigawa M (eds) CCN proteins: a new family of cell growth and differentiation regulators. Imperial College Press, London, pp 207–238
Grote K, Bavendiek U, Grothusen C, Flach I, Hilfiker-Kleiner D, Drexler H, Schieffer B (2004) Stretch-inducible expression of the angiogenic factor CCN1 in vascular smooth muscle cells is mediated by Egr-1. J Biol Chem 279(53):55675–55681
Haydont V, Mathe D, Bourgier C, Abdelali J, Aigueperse J, Bourhis J, Vozenin-Brotons MC (2005) Induction of CTGF by TGF-[beta]1 in normal and radiation enteritis human smooth muscle cells: Smad/Rho balance and therapeutic perspectives. Radiother Oncol 76(2):219–225
Hirasaki S, Koide N, Ujike K, Shinji T, Tsuji T (2001) Expression of Nov, CYR61 and CTGF genes in human hepatocellular carcinoma. Hepatol Res 19(3):294–305
Hirsch E, Gullberg D, Balzac F, Altruda F, Silengo L, Tarone G (1994) Alpha(V) integrin subunit is predominantly located in nervous-tissue and skeletal-muscle during mouse development. Dev Dyn 201(2):108–120
Inadera H, Hashimoto S, Dong HY, Suzuki T, Nagai S, Yamashita T, Toyoda N, Matsushima K (2000) WISP-2 as a novel estrogen-responsive gene in human breast cancer cells. Biochem Biophys Res Commun 275(1):108–114
Ivkovic S, Yoon BS, Popoff SN, Safadi FF, Libuda DE, Stephenson RC, Daluiski A, Lyons KM (2003) Connective tissue growth factor coordinates chondrogenesis and angiogenesis during skeletal development. Development 130(12):2779–2791
Kauffman MH (1992) Atlas of mouse development. Elsevier, San Diego, CA
Kireeva ML, Latinkic BV, Kolesnikova TV, Chen CC, Yang GP, Abler AS, Lau LF (1997) Cyr61 and Fisp12 are both ECM-associated signaling molecules: activities, metabolism, and localization during development. Exp Cell Res 233(1):63–77
Kocialkowski S, Yeger H, Kingdom J, Perbal B, Schofield PN (2001) Expression of the human NOV gene in first trimester fetal tissues. Anat Embryol 203(6):417–427
Kubota S, Takigawa M (2007) Role of CCN2/CTGF/Hcs24 in bone growth. Int Rev Cyt 257:1–41
Kumar S, Hand AT, Connor JR, Dodds RA, Ryan PJ, Trill JJ, Fisher SM, Nuttall ME, Lipshutz DB, Zou C, Hwang SM, Votta BJ, James IE, Rieman DJ, Gowen M, Lee JC (1999) Identification and cloning of a connective tissue growth factor-like cDNA from human osteoblasts encoding a novel regulator of osteoblast functions. J Biol Chem 274(24):17123–17131
Lake A, Castellot J (2003) CCN5 modulates the antiproliferative effect of heparin and regulates cell motility in vascular smooth muscle cells. Cell Commun Signal 1(1):5
Lake AC, Bialik A, Walsh K, Castellot JJ (2003) CCN5 is a growth arrest-specific gene that regulates smooth muscle cell proliferation and motility. Am J Pathol 162(1):219–231
Lau LF, Lam SC-T (1999) The CCN family of angiogenic regulators: the integrin connection. Exp Cell Res 248(1):44–57
Lau LM, Lam SC-T (2005) Integrin-mediated CCN functions. In: Perbal BV, Takigawa M (eds) CCN proteins: a new family of cell growth and differentiation regulators. Imperial College Press, London, pp 61–80
Leask A, Abraham DJ (2006) All in the CCN family: essential matricellular signaling modulators emerge from the bunker. J Cell Sci 119(23):4803–4810
Li GM, Xie Q, Shi Y, Li DG, Zhang MJ, Jiang S, Zhou HJ, Lu HM, Jin YX (2006) Inhibition of connective tissue growth factor by siRNA prevents liver fibrosis in rats. J Gene Med 8(7):889–900
Lopes SMCD, Feijen A, Korving J, Korchynskyi O, Larsson J, Karlsson S, Ten Dijke P, Lyons KM, Goldschmeding R, Doevendans P, Mummery CL (2004) Connective tissue growth factor expression and Smad signaling during mouse heart development and myocardial infarction. Dev Dyn 231(3):542–550
Manns JM, Uknis AB, Rico MC, Agelan A, Castaneda J, Arango I, Barbe MF, Safadi FF, Popoff SN, DeLa Cadena RA (2006) A peptide from thrombospondin 1 modulates experimental erosive arthritis by regulating connective tissue growth factor. Arthritis Rheum 54(8):2415–2422
Mason HR, Castellot JJ, Nowak RA (2002) CCN5 inhibits the proliferation and motility of human myometrial and leiomyoma smooth muscle cells. Mol Biol Cell 13:289A–289A
Mason HR, Grove-Strawser D, Rubin BS, Nowak RA, Castellot JJ (2004a) Estrogen induces CCN5 expression in the rat uterus in vivo. Endocrinology 145(2):976–982
Mason HR, Lake AC, Wubben JE, Nowak RA, Castellot JJ (2004b) The growth arrest-specific gene CCN5 is deficient in human leiomyomas and inhibits the proliferation and motility of cultured human uterine smooth muscle cells. Mol Hum Reprod 10(3):181–187
McCormac JT, Greenwal GS (1974) Progesterone and estradiol-17Beta concentrations in peripheral plasma during pregnancy in mouse. J Endocrinol 62(1):101–107
Natarajan D, Andermarcher E, Schofield PN, Boulter CA (2000) Mouse Nov gene is expressed in hypaxial musculature and cranial structures derived from neural crest cells and placodes. Dev Dyn 219(3):417–423
O’Brien TP, Lau LF (1992) Expression of the growth factor-inducible immediate early gene Cyr61 correlates with chondrogenesis during mouse embryonic-development. Cell Growth Differ 3(9):645–654
Pennica D, Swanson TA, Welsh JW, Roy MA, Lawrence DA, Lee J, Brush J, Taneyhill LA, Deuel B, Lew M, Watanabe C, Cohen RL, Melhem MF, Finley GG, Quirke P, Goddard AD, Hillan KJ, Gurney AL, Botstein D, Levine AJ (1998) WISP genes are members of the connective tissue growth factor family that are up-regulated in Wnt-1-transformed cells and aberrantly expressed in human colon tumors. Proc Natl Acad Sci U S A 95(25):14717–14722
Perbal B (1999) Nuclear localisation of NOVH protein: a potential role for NOV in the regulation of gene expression. Mol Pathol 52(2):84–91
Perbal B (2004) CCN proteins: multifunctional signalling regulators. Lancet 363(9402):62–64
Perbal BV, Takigawa M (eds) (2005a) CCN Proteins: a new family of cell growth and differentiation regulators. Imperial College Press, London
Perbal B, Takigawa M (2005b) The CCN family of proteins: an overview. In: Perbal BV, Takigawa M (eds) CCN proteins: a new family of cell growth and differentiation regulators. Imperial College Press, London, pp 1–18
Planque N, Li CL, Saule S, Bleau AM, Perbal B (2006) Nuclear addressing provides a clue for the transforming activity of amino-truncated CCN3 proteins. J Cell Biochem 99(1):105–116
Rachfal AW, Brigstock DR (2005) Structural and functional properties of CCN proteins. Vitam Horm 70:69–103
Razzaque MS, Foster CS, Ahmed AR (2003) Role of connective tissue growth factor in the pathogenesis of conjunctival scarring in ocular cicatricial pemphigoid. Investig Ophthalmol Vis Sci 44(5):1998–2003
Riser B, Karoor S, Peterson D (2005) CCN genes and the kidney. In: Perbal BV, Takigawa M (eds) CCN proteins: a new family of cell growth and differentiation regulators. Imperial College Press, London, pp 95–116
Safadi FF, Xu J, Smock SL, Kanaan RA, Selimf AH, Odgren PR, Marks SC, Owen TA, Popoff SN (2003) Expression of connective tissue growth factor in bone: its role in osteoblast proliferation and differentiation in vitro and bone formation in vivo. J Cell Physiol 196(1):51–62
Schutze N, Noth U, Schneidereit J, Hendrich C, Jakob F (2005) Differential expression of CCN-family members in primary human bone marrow-derived mesenchymal stem cells during osteogenic, chondrogenic and adipogenic differentiation. Cell Commun Signal 3(1):5
Shin JY, Hur W, Wang JS, Jang JW, Kim CW, Bae SH, Jang SK, Yang SH, Sung YC, Kwon OJ, Yoon SK (2005) HCV core protein promotes liver fibrogenesis via up-regulation of CTGF with TGF-beta 1. Exp Mol Med 37(2):138–145
Surveyor GA, Brigstock DR (1999) Immunohistochemical localization of connective tissue growth factor (CTGF) in the mouse embryo between days 7.5 and 14.5 of gestation. Growth Factors 17(2):115–124
Surveyor GA, Wilson AK, Brigstock DR (1998) Localization of connective tissue growth factor during the period of embryo implantation in the mouse. Biol Reprod 59(5):1207–1213
Takigawa M, Nakanishi T, Kubota S, Nishida T (2003) Role of CTGF/HCS24/ecogenin in skeletal growth control. J Cell Physiol 194(3):256–266
Takigawa M, Nishida T, Kubota S (2005) Roles of CCN2/CTGF in the control of growth and regeneration. In: Perbal BV, Takigawa M (eds) CCN proteins: a new family of cell growth and differentiation regulators. Imperial College Press, London, pp 19–60
Tanaka I, Morikawa M, Okuse T, Shirakawa M, Imai K (2005) Expression and regulation of WISP2 in rheumatoid arthritic synovium. Biochem Biophys Res Commun 334(4):973–978
Tarone G, Hirsch E, Brancaccio M, De Acetis M, Barberis L, Balzac F, Retta F, Botta C, Altruda F, Silengo L (2000) Integrin function and regulation in development. Int J Dev Biol 44(6):725–731
Tavian M, Peault B (2005) The changing cellular environments of hematopoiesis in human development in utero. Exp Hematol 33(9):1062–1069
van den Brule S, Heymans J, Havaux X, Renauld JC, Lison D, Huax F, Denis O (2007) Pro-fibrotic effect of IL-9 overexpression in a model of airway remodeling. Am J Resir Cell Mol Biol 37:202–209
Van Tuyl M, Post M (2003) Molecular mechanisms of lung development and lung branching morphogenesis. In: Polin RA, Fox WW, Abman SH (eds) Fetal and neonatal physiology, 3rd edn. Saunders, Harcourt Health Sciences, Philadelphia, PA, p. 812–821
Volpe MV, Nielsen HC, Archavachotikul K, Ciccone TJ, Chinoy MR (2003) Thyroid hormone affects distal airway formation during the late pseudoglandular period of mouse lung development. Molec Genet Metab 80(1–2):242–254
Vozenin-Brotons MC, Milliat F, Sabourin JC, de Gouville AC, Francois A, Lasser P, Morice P, Haie-Meder C, Lusinchi A, Antoun S, Bourhis J, Mathe D, Girinsky T, Aigueperse J (2003) Fibrogenic signals in patients with radiation enteritis are associated with increased connective tissue growth factor expression. Int J Radiat Oncol Biol Phys 56(2):561–572
Wahab NA, Brinkman H, Mason RM (2001) Uptake and intracellular transport of the connective tissue growth factor: a potential mode of action. Biochem J 359:89–97
Wert SE (2004) Normal and abnormal structural development of the lung. In: Polin RA, Fox WW, Abman SH (eds) Fetal and neonatal physiology, 3rd edn. W.B. Saunders Co., Philadelphia, PA, pp 783–793
Willems E, Mateizel I, Kemp C, Cauffman G, Sermon K, Leyns L (2006) Selection of reference genes in mouse embryos and in differentiating human and mouse ES cells. Int J Dev Biol 50(7):627–635
Yamaai T, Nakanishi T, Asano M, Nawachi K, Yoshimichi G, Ohyama K, Komori T, Sugimoto T, Takigawa M (2005) Gene expression of connective tissue growth factor (CTGF/CCN2) in calcifying tissues of normal and cbfa1-null mutant mice in late stage of embryonic development. J Bone Miner Metab 23(4):280–288
Zhang R, Averboukh L, Zhu WM, Zhang H, Jo H, Dempsey PJ, Coffey RJ, Pardee AB, Liang P (1998) Identification of rCop-1, a new member of the CCN protein family, as a negative regulator for cell transformation. Mol Cell Biol 18(10):6131–6141
Acknowledgements
This work was supported by NIH Grants HD046251 and HL49973 to JJC and by Fundacao para a Ciencia e Tecnologia (F.C.T.), no. SFRH/BD/1284/2000 to BEO. We are grateful to John A. Hamlin and Alexandra Sanfins for teaching JAM the embryo dissection, manipulation, and fixation techniques used in this report for both preliminary studies and also in collecting the tissue used in the RT-PCR studies. We also thank David R. Brigstock for generously providing anti-CCN2 antibody used to confirm expression patterns seen in our studies with the commercial antibody used in this report and also advice on antibody characterization. We thank MaryAnn Volpe, Josh Russo, Joan Lemire, Ron Myers, Kristina Cvitanovic, and Lan Wei for useful discussions and advice. We thank Po Kwok Tse for expert technical assistance.
Competing interests
None declared.
Authors’ contributions
All authors participated in the design and interpretation of the experiments presented, as well as in the preparation, editing, or approval of the manuscript. JJC discovered CCN5 mRNA and protein, provided overall direction and guidance, edited the manuscript, and provided the financial support. JAM provided intellectual input into experimental design and interpretation, performed most of the experiments, and wrote and edited the manuscript; MRG and MK provided intellectual and technical oversight of parts of the project. MBO provided intellectual and technical contributions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplemental Fig. 1
CCN5 expression during E9–E11 mouse development. Mouse embryonic sagittal sections from a E9 and b E11 developing mice. Negative control – pooled rabbit IgG applied in place of primary antibody. Br = Brain; He = Heart. a,b IgG top, CCN2 middle, CCN5 bottom. Scale Bars = 1 mm. Scale Bars and labels on CCN5 panels also apply to CCN2 and IgG panels. CCN5 and CCN2 immunoreactivity shown by brown staining. Nuclei are shown by hematoxylin (blue) staining. Tissue sections treated with pooled rabbit IgG gave no detectable staining (a,b) (DOC 3.09 MB).
Supplemental Fig. 2
CCN5 expression during cardiovascular development. a GD16 artery from lung. b GD16 artery from kidney. c GD16 mouse heart. d Human 5 month fetal blood vessel. e Human 5 month fetal umbilical cord. a–e CCN2 left, CCN5 right. Scale Bars = 10 μm (a,b). Scale Bars = 100 μm (c–e). Tissue sections treated with pooled rabbit IgG gave no detectable staining (data not shown) (DOC 3.08 MB).
Supplemental Fig. 3
CCN5 expression in developing bones and skeletal muscle. a Human 5 month fetal skeletal muscle. b Human 5 month fetal bone. Inset shows a magnified osteoclast. OCL = Osteoclast; OCT = Osteocyte. a,b CCN2 left, CCN5 right. Scale Bars = 100 μm (a,b,c). Scale Bars = 20 μm (a,b). Tissue sections treated with pooled rabbit IgG gave no detectable staining (data not shown) (DOC 2.85 MB).
Supplemental Fig. 4
CCN5 expression during intestinal development. a Human 5 month fetal stomach. b Human 5 month fetal intestine. c Human 5 month fetal gallbladder. d Human 5 month fetal rectum. a–d CCN2 left, CCN5 right. Scale Bars = 100 μm. Tissue sections treated with pooled rabbit IgG gave no detectable staining (data not shown) (DOC 2.86 MB).
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License ( https://creativecommons.org/licenses/by-nc/2.0 ), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Jones, J.A., Gray, M.R., Oliveira, B.E. et al. CCN5 expression in mammals. J. Cell Commun. Signal. 1, 127–143 (2007). https://doi.org/10.1007/s12079-007-0012-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12079-007-0012-0