Abstract
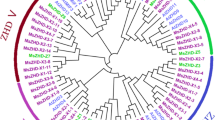
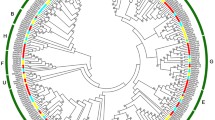
Banana is an important tropical fruit belonging to the Musa family, which is susceptible to nutritional deficiency symptoms during cultivation. Due to magnesium deficiency, the growth of bananas (Musa paradisiaca AA) is inhibited, and become prone to diseases and pests, which decreases the yield and quality of bananas. Basic helix-loop-helix(bHLH) is one of the largest transcription factor(TF) families in plants. It plays an important role in plant growth, development as well as resistance against abiotic stresses. In this research, 177 MabHLH proteins with four conserved subdomains were obtained via 166 AtbHLH proteins were applied to homologous comparisons from the banana genome database, which carried five highly conserved key amino acid residues consist of Arg-8, Arg-9, Leu-19, Leu-45 and Leu-52. Then, 12 subfamilies were divided according to the phylogenetic tree of 177 MabHLHs with wholly distributed in 11 chromosomes of banana. Functional prediction substantiated that subfamilies of 177 MabHLHs were involved in different biological and abiotic stresses, homeostasis regulation, cell signaling, plant growth and development and so on. Transcriptome analysis showed that the expression of 45 MabHLHs existed a significant difference in response to magnesium deficiency stress, among 16 MabHLHs were part of Group III and IVa. The qRT-PCR experiment of 18 MabHLHs confirmed that MabHLHs did play an important role in magnesium deficiency stress. Above all, we studied the MabHLH gene family of bananas to dig out the potential regulatory genes against magnesium deficiency stress, which are holpful to further improve the stress resistance level of bananas under magnesium deficiency.





Similar content being viewed by others
Data Availability
The data that support the findings of this study are available from Hainan Key Laboratory for Sustainable Utilization of Tropical Bioresources, which were used under license for this study. Data are available in https://doi.org/10.1007/s00425-021-03769-z.
References
Al-Sady B, Ni W, Kircher S et al (2006) Photoactivated phytochrome induces rapid PIF3 phosphorylation prior to proteasome-mediated degradation. Mol Cell 23(3):439–446. https://doi.org/10.1016/j.molcel.2006.06.011. PMID: 16885032.
Atchley WR, Terhalle W, Dress A (1999) Positional dependence, cliques, and predictive motifs in the bHLH protein domain. J Mol Evol 48(5):501–516. https://doi.org/10.1007/pl00006494
Bailey TL, Johnson J, Grant CE et al (2015) The MEME suite. Nucleic Acids Res 43(W1):W39–W49. https://doi.org/10.1093/nar/gkv416
Bauer P, Ling H-Q, Guerinot ML (2007) FIT, the fer-like iron deficiency induced transcription factor in arabidopsis. Plant Physiol Biochem PPB 45(5):260–261. https://doi.org/10.1016/j.plaphy.2007.03.006
Berardini TZ, Reiser L, Li D et al (2015) The arabidopsis information resource: Making and mining the “gold standard” annotated reference plant genome. Genesis 53(8):474–485. https://doi.org/10.1002/dvg.22877
Billard V, Maillard A, Coquet L et al (2016) Mg deficiency affects leaf Mg remobilization and the proteome in Brassica napus. Plant Physiol Biochem 107:337–343. https://doi.org/10.1016/j.plaphy.2016.06.025
Bruessow F, Bautor J, Hoffmann G et al (2021) Natural variation in temperature-modulated immunity uncovers transcription factor bHLH059 as a thermoresponsive regulator in Arabidopsis thaliana. Plos Genet 17(1):e1009290. https://doi.org/10.1371/journal.pgen.1009290
Cammarano P, Felsani A, Gentile M et al (1972) Formation of active hybrid 80-S particles from subunits of pea seedlings and mammalian liver ribosomes. Biochem Biophys Acta 281(4):625–642. https://doi.org/10.1016/0005-2787(72)90160-8
Carretero-Paulet L, Galstyan A, Roig-Villanova I et al (2010) Genome-wide classification and evolutionary analysis of the bHLH family of transcription factors in arabidopsis, poplar, rice, moss, and algae. Plant Physiol 153(3):1398–1412. https://doi.org/10.1104/pp.110.153593
Chen C, Chen H, Zhang Y et al (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Chen H-B, Fan X-L (2018) Effects of magnesium remobilization and allocation on banana plant growth. J Plant Nutr 41(10):1312–1320. https://doi.org/10.1080/01904167.2018.1450422
Chinnusamy V, Ohta M, Kanrar S et al (2003) ICE1: a regulator of cold-induced transcriptome and freezing tolerance in Arabidopsis. Genes Dev 17(8):1043–1054. https://doi.org/10.1101/gad.1077503
Crooks GE, Hon G, Chandonia JM (2004) Brenner SE WebLogo: A sequence logo generator. Genome Res 2004(14):1188–1190
D’Hont A, Denoeud F, Aury J-M et al (2012) The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 488(7410):213–+. https://doi.org/10.1038/nature11241
Droc G, Lariviere D, Guignon V et al (2013). The Banana Genome Hub Database. https://doi.org/10.1093/database/bat035
Du H, Liang Z, Zhao S et al (2015) The evolutionary history of R2R3-MYB proteins across 50 eukaryotes: New insights into subfamily classification and expansion. Sci Rep 5:11037. https://doi.org/10.1038/srep11037
Duek PD, Fankhauser C (2005) bHLH class transcription factors take centre stage in phytochrome signalling. Trends Plant Sci 10(2):51–54. https://doi.org/10.1016/j.tplants
Eddy SR (1998) Profile hidden Markov models. Bioinformatics 14(9):755–763. https://doi.org/10.1093/bioinformatics/14.9.755. PMID: 9918945
Feng X-M, Zhao Q, Zhao L-L et al (2012) The cold-induced basic helix-loop-helix transcription factor gene MdCIbHLH1 encodes an ICE-like protein in apple. Bmc Plant Biol 12:22. https://doi.org/10.1186/1471-2229-12-22
Gransee A, Fuehrs H (2013) Magnesium mobility in soils as a challenge for soil and plant analysis, magnesium fertilization and root uptake under adverse growth conditions. Plant Soil 368(1–2):5–21. https://doi.org/10.1007/s11104-012-1567-y
Guo W, Nazim H, Liang Z et al (2016) Magnesium deficiency in plants: An urgent problem. Crop Journal 4(2):83–91. https://doi.org/10.1016/j.cj.2015.11.003
Han X, Zhang M, Yang M et al (2020) Arabidopsis JAZ Proteins Interact with and Suppress RHD6 Transcription Factor to Regulate Jasmonate-Stimulated Root Hair Development. Plant Cell 32(4):1049–1062. https://doi.org/10.1105/tpc.19.00617
Hao Y, Zong X, Ren P et al (2021) Basic helix-loop-helix (bHLH) transcription factors regulate a wide range of functions in Arabidopsis. Int J Mol Sci 22(13):7152. https://doi.org/10.3390/ijms22137152. PMID: 34281206
Heim MA, Jakoby M, Werber M et al (2003) The basic helix-loop-helix transcription factor family in plants: a genome-wide study of protein structure and functional diversity. Mol Biol Evol 20(5):735–747. https://doi.org/10.1093/molbev/msg088
Hermans C, Chen J, Coppens F et al (2011) Low magnesium status in plants enhances tolerance to cadmium exposure. New Phytol 192(2):428–436
Huang JH, Xu J, Ye X et al (2019) Magnesium deficiency affects secondary lignification of the vascularsystem in Citrus sinensis seedlings. Trees 33:171–182. https://doi.org/10.1007/s00468-018-1766-0
Huang X-S, Wang W, Zhang Q et al (2013) A basic helix-loop-helix transcription factor, PtrbHLH, of poncirus trifoliata confers cold tolerance and modulates peroxidase-mediated scavenging of hydrogen peroxide. Plant Physiol 162(2):1178–1194. https://doi.org/10.1104/pp.112.210740
Krzywinski M, Schein J, Birol I et al (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645. https://doi.org/10.1101/gr.092759.109
Kurt F, Filiz E (2018) Genome-wide and comparative analysis of bHLH38, bHLH39, bHLH100 and bHLH101 genes in Arabidopsis, tomato, rice, soybean and maize: insights into iron (Fe) homeostasis. Biometals 4:489–504. https://doi.org/10.1007/s10534-018-0095-5
Lalitha S (2004) Primer premier 5. Biotech Softw Internet Rep 270–272. https://doi.org/10.1089/152791600459894
Lassoudière A (2007) Bananier et sa culture(le) (p.383). Versailles CEDEX. France: Editions Quae. 383 p. (Savoir-faire) ISBN 978–2–7592–0046–7
Lau OS, Song Z, Zhou Z et al (2018) Direct control of SPEECHLESS by PIF4 in the high-temperature response of stomatal development. Curr Biol 28(8):1273–+. https://doi.org/10.1016/j.cub.2018.02.054
Leivar P, Tepperman JM, Cohn MM et al (2012) Dynamic antagonism between phytochromes and PIF family basic helix-loop-helix factors induces selective reciprocal responses to light and shade in a rapidly responsive transcriptional network in Arabidopsis. Plant Cell 24(4):1398–1419. https://doi.org/10.1105/tpc.112.095711. Epub 2012 Apr 18 PMID: 22517317
Letunic I, Khedkar S, Bork P (2021) SMART: recent updates, new developments and status in 2020. Nucleic Acids Res 49(D1):D458–D460. https://doi.org/10.1093/nar/gkaa937. PMID: 33104802; PMCID: PMC7778883
Li C, Yan C, Sun Q et al (2021) The bHLH transcription factor AhbHLH112 improves the drought tolerance of peanut. BMC Plant Biol 21(1):540. https://doi.org/10.1186/s12870-021-03318-6. PMID: 34784902
Li X, Duan X, Jiang H et al (2006) Genome-wide analysis of basic/helix-loop-helix transcription factor family in rice and Arabidopsis. Plant Physiol 141(4):1167–1184. https://doi.org/10.1104/pp.106.080580
Liang B, Wan S, Ma Q et al (2022) A novel bHLH Transcription factor ptrbhlh66 from trifoliate orange positively regulates plant drought tolerance by mediating root growth and ROS scavenging. Int J Mol Sci 23(23):15053. https://doi.org/10.3390/ijms232315053. PMID: 36499381
Liu Y, Lin-Wang K, Espley RV et al (2019) StMYB44 negatively regulates anthocyanin biosynthesis at high temperatures in tuber flesh of potato. J Exp Bot 70(15):3809–3824. https://doi.org/10.1093/jxb/erz194
Lorenzo O, Chico JM, Sanchez-Serrano JJ et al (2004) JASMONATE-INSENSITIVE1 encodes a MYC transcription factor essential to discriminate between different jasmonate-regulated defense responses in Arabidopsis. Plant Cell 16(7):1938–1950. https://doi.org/10.1105/tpc.022319
Lu R, Li Y, Zhang J et al (2022) The bHLH/HLH transcription factors GhFP2 and GhACE1 antagonistically regulate fiber elongation in cotton. Plant Physiol 189(2):628–643. https://doi.org/10.1093/plphys/kiac088. PMID: 35226094
Ludwig SR, Habera LF, Dellaporta SL et al (1989) (1989) Lc, a member of the maize R gene family responsible for tissue-specific anthocyanin production, encodes a protein similar to transcriptional activators and contains the myc-homology region. Proc Natl Acad Sci USA 86(18):7092–7096. https://doi.org/10.1073/pnas.86.18.7092
Madden T (2008) Threading by query. 2008 Jun 23 [Updated 2021 Jun 28]. In: BLAST command line applications user manual [Internet]. Bethesda (MD): National Center for Biotechnology Information (US). Available from: https://www.ncbi.nlm.nih.gov/books/NBK571452/. Accessed Mar 2022
Maguire ME, Cowan JA (2002) Magnesium chemistry and biochemistry. Biometals Int J Role Metal Ions Biol Biochem Med 15(3):203–210. https://doi.org/10.1023/a:1016058229972
Mao K, Dong Q, Li C et al (2017) Genome wide identification and characterization of Apple bHLH transcription factors and expression analysis in response to drought and salt stress. Front Plant Sci 8:480. https://doi.org/10.3389/fpls.2017.00480
Mistry J, Chuguransky S, Williams L et al (2021) Pfam: The protein families database in 2021. Nucleic Acids Res 49(D1):D412–D419. https://doi.org/10.1093/nar/gkaa913. PMID: 33125078
Moreno JE, Moreno-Piovano G, Chan RL (2018) The antagonistic basic helix-loop-helix partners BEE and IBH1 contribute to control plant tolerance to abiotic stress. Plant Sci 271:143–149. https://doi.org/10.1016/j.plantsci.2018.03.024
Nakata M, Mitsuda N, Herde M et al (2013) A bHLH-type transcription factor, ABA-INDUCIBLE BHLH-TYPE TRANSCRIPTION FACTOR/JA-ASSOCIATED MYC2-LIKE1, acts as a repressor to negatively regulate jasmonate signaling in Arabidopsis. Plant Cell 25(5):1641–1656. https://doi.org/10.1105/tpc.113.111112
Niu X, Guan Y, Chen S et al (2017) Genome-wide analysis of basic helix-loop-helix (bHLH) transcription factors in Brachypodium distachyon. BMC Genom 18:619
Ogo Y, Itai RN, Nakanishi H et al (2007) The rice bHLH protein OsIRO2 is an essential regulator of the genes involved in Fe uptake under Fe-deficient conditions. Plant J Cell Mol Biol 51(3):366–377. https://doi.org/10.1111/j.1365-313X.2007.03149.x
Paul J-Y, Khanna H, Kleidon J et al (2017) Golden bananas in the field: elevated fruit pro-vitamin A from the expression of a single banana transgene. Plant Biotechnol J 15(4):520–532. https://doi.org/10.1111/pbi.12650
Raissig MT, Matos JL, Gil MXA et al (2017) Mobile MUTE specifies subsidiary cells to build physiologically improved grass stomata. Science 355(6330):1215–1218. https://doi.org/10.1126/science.aal3254
Riechmann JL, Heard J, Martin G et al (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science (New York, N.Y.) 290(5499):2105–2110. https://doi.org/10.1126/science.290.5499.2105
Salih H, Tan L, Htet NNW (2021) Genome-wide identification, characterization of bHLH transcription factors in mango. Trop Plant Biol 14(1):72–81. https://doi.org/10.1007/s12042-020-09277-w
Samira R, Li B, Kliebenstein D et al (2018) The bHLH transcription factor ILR3 modulates multiple stress responses in Arabidopsis. Plant Mol Biol 97(4–5):297–309. https://doi.org/10.1007/s11103-018-0735-8
Sasaki-Sekimoto Y, Saito H, Masuda S et al (2014) Comprehensive analysis of protein interactions between JAZ proteins and bHLH transcription factors that negatively regulate jasmonate signaling. Plant Signal Behav 9(1):e27639–e27639. https://doi.org/10.4161/psb.27639
Song S, Qi T, Fan M et al (2013) The bHLH subgroup IIId factors negatively regulate jasmonate-mediated plant defense and development. Plos Genet 9(7):e1003653. https://doi.org/10.1371/journal.pgen.1003653
Sun X, Kay AD, Kang H et al (2013) Correlated biogeographic variation of magnesium across trophic levels in a terrestrial food chain. Plos One 8(11):e78444. https://doi.org/10.1371/journal.pone.0078444
Tamura K, Dudley J, Nei M et al (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599. https://doi.org/10.1093/molbev/msm092
Toledo-Ortiz G, Huq E, Quail PH (2003) The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 15(8):1749–1770. https://doi.org/10.1105/tpc.013839
Van de Peer Y, Mizrachi E, Marchal K (2017) The evolutionary significance of polyploidy. Nat Rev Genet 18(7):411–424. https://doi.org/10.1038/nrg.2017.26
Verbruggen N, Hermans C (2013) Physiological and molecular responses to magnesium nutritional imbalance in plants. Plant Soil 368(1–2):87–99. https://doi.org/10.1007/s11104-013-1589-0
Wang D, Zhang Y, Zhang Z et al (2010) KaKs_Calculator 2.0: a toolkit incorporating gamma-series methods and sliding window strategies. Genom Proteom Bioinform 8(1):77–80. https://doi.org/10.1016/S1672-0229(10)60008-3. PMID: 20451164; PMCID: PMC5054116
Wang J, Hu Z, Zhao T et al (2015) Genome-wide analysis of bHLH transcription factor and involvement in the infection by yellow leaf curl virus in tomato (Solanum lycopersicum). BMC Genom 16(1):39. https://doi.org/10.1186/s12864-015-1249-2
Wilkins MR, Gasteiger E, Bairoch A et al (1999) Protein identification and analysis tools in the ExPASy server. Methods Mol Biol 1999(112):531–552. https://doi.org/10.1385/1-59259-584-7:531
Xie X-B, Li S, Zhang R-F et al (2012) The bHLH transcription factor MdbHLH3 promotes anthocyanin accumulation and fruit colouration in response to low temperature in apples. Plant Cell Environ 35(11):1884–1897. https://doi.org/10.1111/j.1365-3040.2012.02523.x
Xu Y, Zhang H, Zhong Y et al (2022) Comparative genomics analysis of bHLH genes in cucurbits identifies a novel gene regulating cucurbitacin biosynthesis. Hortic Res 9:uhac038. https://doi.org/10.1093/hr/uhac038
Yan H, Jiang G, Wu F et al (2021). Sulfoxidation regulation of transcription factor NAC42 influences its functions in relation to stress-induced fruit ripening in banana. J Exp Bot 72(2):682–699. https://doi.org/10.1093/jxb/eraa474. PMID: 33070185
Yang J, Chen Y, Gao M et al (2022) Comprehensive identification of bHLH transcription factors in Litsea cubeba reveals candidate gene involved in the monoterpene biosynthesis pathway. Front Plant Sci 13:1081335. https://doi.org/10.3389/fpls.2022.1081335
Yang N, Jiang J, Xie H et al (2017) Metabolomicsreveals distinct carbon and nitrogen metabolic responses tomagnesium deficiency in leaves and roots of soybean [Glycine max (Linn.) Merr.]. Front Plant Sci 8:2091. https://doi.org/10.3389/fpls.2017.02091
Yao X, Cai Y, Yu D et al (2018) bHLH104 confers tolerance to cadmium stress in Arabidopsis thaliana. J Integr Plant Biol 60(8):691–702. https://doi.org/10.1111/jipb.12658
Ye K, Li H, Ding Y et al (2019) BRASSINOSTEROID-INSENSITIVE2 negatively regulates the stability of transcription factor ICE1 in response to cold stress in arabidopsis. Plant Cell 31(11):2682–2696. https://doi.org/10.1105/tpc.19.00058
Zander M, Lewsey MG, Clark NM et al (2020) Integrated multi-omics framework of the plant response to jasmonic acid. Nat Plants 6(3):290–302. https://doi.org/10.1038/s41477-020-0605-7
Zhang C, Lei Y, Lu C et al (2020) MYC2, MYC3, and MYC4 function additively in wounding-induced jasmonic acid biosynthesis and catabolism. J Integr Plant Biol 62(8):1159–1175. https://doi.org/10.1111/jipb.12902
Zorrilla-Fontanesi Y, Rouard M, Cenci A et al (2016) Differential root transcriptomics in a polyploid non-model crop: the importance of respiration during osmotic stress. Sci Rep 7:25683. https://doi.org/10.1038/srep22583
Funding
This research was financially supported by the Hainan Natural Science Foundation of High-Level Talents Project (no. 320RC494), the Research Foundation for Advanced Talents of Hainan University [no. KYQD(ZR)1849].
Author information
Authors and Affiliations
Contributions
H.Y.H. conceptualized and designed the experiments. P.L. and X.P.L. performed most of the experiments under the supervision of H.Y.H., provided help with bio-informatic analysis and wrote the manuscript with the support of all other authors. S.H.Z. helped to collect the samples. P.L, X.P.L and H.Y.H. critically revised the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by: Ray Ming
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, P., Li, X., Zhou, S. et al. Genome-Wide Identification of MabHLH Transcription Factors Family Response to Magnesium Deficiency Stress in Banana (Musa paradisiaca AA). Tropical Plant Biol. 16, 156–169 (2023). https://doi.org/10.1007/s12042-023-09336-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12042-023-09336-y