Abstract
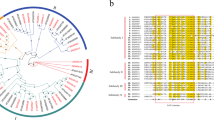
GOLDEN 2-LIKE (GLK) transcription factors are members of the GARP superfamily and are involved in chloroplast development and stress tolerance. These transcription factors have been studied at the genome level across several plant species. However, no study has comprehensively analyzed the GLK family in foxtail millet (Setaria italica L.). The present study discovered 59 GLK genes in foxtail millet. Multiple sequence alignment and gene motif analysis revealed the presence of Myb-SHAQKYF and Myb-CC-LHEQLE motifs in all the members. In addition, the gene promoters mainly had light- and stress-responsive cis-regulatory elements. Syntenic analysis showed that whole-genome duplications (WGD) and dispersed duplications contributed to the expansion of the SiGLK gene family. Gene expression analysis revealed that SiGLKs exhibited preferential expression in bundle sheath than in the mesophyll cells. Meanwhile, analyzing these genes in foxtail millet under abiotic stress demonstrated a significant role for SiGLK30, 38, 48, and 52 in regulating salt, cold, and drought stress responses. Finally, qRT-PCR confirmed that cold stress increased SiGLK52 and SiGLK31 expression. The study thus provides an overview of GLK gene evolution in foxtail millet and a clue to its function in bundle sheath and mesophyll cell and stress tolerance.










Similar content being viewed by others
Data Availability
All data used in this study were obtained from publicly accessible databases, and all generated data were included in this published article and its supplementary information file.
Abbreviations
- GLK:
-
GOLDEN 2-LIKE
- pI:
-
Isoelectric points
- MW:
-
Molecular weight
- NJ:
-
Neighbor-joining
- BS:
-
Bundle sheath
- M:
-
Mesophyll
- WGD:
-
Whole genome duplication
- mya:
-
Million years ago
References
Barton L, Newsome SD, Chen FH, Wang H, Guilderson TP, Bettinger RL (2009) Agricultural origins and the isotopic identity of domestication in northern China. Proc Natl Acad Sci USA 106(14):5523–5528. https://doi.org/10.1073/pnas.0809960106
Bettinger RL, Barton L, Morgan C (2010) The origins of food production in north China: a different kind of agricultural revolution. Evol Anthropol: Issues News Rev 19(1):9–21. https://doi.org/10.1002/evan.20236
Blanc G, Wolfe KH (2004) Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell 16(7):1667–1678. https://doi.org/10.1105/tpc.021345
Bravo-Garcia A, Yasumura Y, Langdale JA (2009) Specialization of the Golden2-like regulatory pathway during land plant evolution. New Phytol 183(1):133–141. https://doi.org/10.1111/j.1469-8137.2009.02829.x
Brutnell TP, Wang L, Swartwood K, Goldschmidt A, Jackson D, Zhu XG, Kellogg E, Van Eck J (2010) Setaria viridis: a model for C4 photosynthesis. Plant Cell 22(8):2537–2544. https://doi.org/10.1105/tpc.110.075309
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Chen M, Ji M, Wen B, Liu L, Li S, Chen X, Gao D, Li L (2016) GOLDEN 2-like transcription factors of plants. Front Plant Sci 7. https://doi.org/10.3389/fpls.2016.01509
Donald RG, Cashmore AR (1990) Mutation of either G box or I box sequences profoundly affects expression from the Arabidopsis rbcS-1A promoter. Embo j 9(6):1717–1726
Dong P, Tu X, Chu PY, Lü P, Zhu N, Grierson D, Du B, Li P, Zhong S (2017) 3D chromatin architecture of large plant genomes determined by local A/B compartments. Mol Plant 10(12):1497–1509. https://doi.org/10.1016/j.molp.2017.11.005
Doust AN, Kellogg EA, Devos KM, Bennetzen JL (2009) Foxtail millet: a sequence-driven grass model system. Plant Physiol 149(1):137–141. https://doi.org/10.1104/pp.108.129627
Edwards GE, Franceschi VR, Ku MS, Voznesenskaya EV, Pyankov VI, Andreo CS (2001) Compartmentation of photosynthesis in cells and tissues of C(4) plants. J Exp Bot 52(356):577–590
Emms DM, Covshoff S, Hibberd JM, Kelly S (2016) Independent and parallel evolution of new genes by gene duplication in two origins of C4 photosynthesis provides new insight into the mechanism of phloem loading in C4 species. Mol Biol Evol 33(7):1796–1806. https://doi.org/10.1093/molbev/msw057
Fitter DW, Martin DJ, Copley MJ, Scotland RW, Langdale JA (2002) GLK gene pairs regulate chloroplast development in diverse plant species. Plant J 31(6):713–727. https://doi.org/10.1046/j.1365-313X.2002.01390.x
Gang H, Li R, Zhao Y, Liu G, Chen S, Jiang J (2019) Loss of GLK1 transcription factor function reveals new insights in chlorophyll biosynthesis and chloroplast development. J Exp Bot 70(12):3125–3138. https://doi.org/10.1093/jxb/erz128
Giuliano G, Pichersky E, Malik VS, Timko MP, Scolnik PA, Cashmore AR (1988) An evolutionarily conserved protein binding sequence upstream of a plant light-regulated gene. Proc Natl Acad Sci USA 85(19):7089–7093. https://doi.org/10.1073/pnas.85.19.7089
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2011) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40(D1):D1178–D1186. https://doi.org/10.1093/nar/gkr944
Gowik U, Bräutigam A, Weber KL, Weber AP, Westhoff P (2011) Evolution of C4 photosynthesis in the genus Flaveria: how many and which genes does it take to make C4? Plant Cell 23(6):2087–2105. https://doi.org/10.1105/tpc.111.086264
Hall LN, Rossini L, Cribb L, Langdale JA (1998) GOLDEN 2: a novel transcriptional regulator of cellular differentiation in the maize leaf. Plant Cell 10(6):925–936. https://doi.org/10.1105/tpc.10.6.925
Han X-Y, Li P-X, Zou L-J, Tan W-r, Zheng T, Zhang D-W, Lin H-H (2016) GOLDEN2-LIKE transcription factors coordinate the tolerance to Cucumber mosaic virus in Arabidopsis. Biochem Biophys Res Commun 477(4):626–632. https://doi.org/10.1016/j.bbrc.2016.06.110
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Kumar S, Stecher G, Suleski M, Hedges SB (2017) timetree: a resource for timelines, timetrees, and divergence times. Mol Biol Evol 34(7):1812–1819. https://doi.org/10.1093/molbev/msx116
Lata C, Gupta S, Prasad M (2013) Foxtail millet: a model crop for genetic and genomic studies in bioenergy grasses. Crit Rev Biotechnol 33(3):328–343. https://doi.org/10.3109/07388551.2012.716809
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1):325–327. https://doi.org/10.1093/nar/30.1.325
Li P, Brutnell TP (2011) Setaria viridis and Setaria italica, model genetic systems for the Panicoid grasses. J Exp Bot 62(9):3031–3037. https://doi.org/10.1093/jxb/err096
Liu F, Xu Y, Han G, Zhou L, Ali A, Zhu S, Li X (2016) Molecular evolution and genetic variation of G2-like transcription factor genes in maize. PLoS ONE 11(8):e0161763–e0161763. https://doi.org/10.1371/journal.pone.0161763
Liu J, Mehari TG, Xu Y, Umer MJ, Hou Y, Wang Y, Peng R, Wang K, Cai X, Zhou Z, Liu F (2021) GhGLK1 a key candidate gene from GARP family enhances cold and drought stress tolerance in cotton. Front Plant Sci 12. https://doi.org/10.3389/fpls.2021.759312
Liu Y et al (2022) The Cycas genome and the early evolution of seed plants. Nat Plants 8(4):389–401. https://doi.org/10.1038/s41477-022-01129-7
Lu S, Wang J, Chitsaz F, Derbyshire MK, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Marchler GH, Song JS, Thanki N, Yamashita RA, Yang M, Zhang D, Zheng C, Lanczycki CJ, Marchler-Bauer A (2020) CDD/SPARCLE: the conserved domain database in 2020. Nucleic Acids Res 48(D1):D265–D268. https://doi.org/10.1093/nar/gkz991
Lundgren MR, Osborne CP, Christin P-A (2014) Deconstructing Kranz anatomy to understand C4 evolution. J Exp Bot 65(13):3357–3369. https://doi.org/10.1093/jxb/eru186
Lynch M, Conery JS (2000) The evolutionary fate and consequences of duplicate genes. Science 290(5494):1151–1155. https://doi.org/10.1126/science.290.5494.1151
Meng X, Liang Z, Dai X, Zhang Y, Mahboub S, Ngu DW, Roston RL, Schnable JC (2021) Predicting transcriptional responses to cold stress across plant species. Proc Natl Acad Sci USA 118(10):e2026330118. https://doi.org/10.1073/pnas.2026330118
Nakamura H, Muramatsu M, Hakata M, Ueno O, Nagamura Y, Hirochika H, Takano M, Ichikawa H (2009) Ectopic overexpression of the transcription factor OsGLK1 induces chloroplast development in non-green rice cells. Plant Cell Physiol 50(11):1933–1949. https://doi.org/10.1093/pcp/pcp138
Nakashima K, Yamaguchi-Shinozaki K (2013) ABA signaling in stress-response and seed development. Plant Cell Rep 32(7):959–970. https://doi.org/10.1007/s00299-013-1418-1
Nguyen CV, Vrebalov JT, Gapper NE, Zheng Y, Zhong S, Fei Z, Giovannoni JJ (2014) Tomato GOLDEN2-LIKE transcription factors reveal molecular gradients that function during fruit development and ripening. Plant Cell 26(2):585–601. https://doi.org/10.1105/tpc.113.118794
Osborne CP, Freckleton RP (2009) Ecological selection pressures for C4 photosynthesis in the grasses. Proc Biol Sci 276(1663):1753–1760. https://doi.org/10.1098/rspb.2008.1762
Panchy N, Lehti-Shiu M, Shiu SH (2016) Evolution of Gene Duplication in Plants. Plant Physiol 171(4):2294–2316. https://doi.org/10.1104/pp.16.00523
Qin M, Zhang B, Gu G, Yuan J, Yang X, Yang J, Xie X (2021) Genome-wide analysis of the G2-Like transcription factor genes and their expression in different senescence stages of tobacco (Nicotiana tabacum L.). Front Genet 12. https://doi.org/10.3389/fgene.2021.626352
Riechmann JL, Heard J, Martin G, Reuber L, Jiang C-Z, Keddie J, Adam L, Pineda O, Ratcliffe OJ, Samaha RR, Creelman R, Pilgrim M, Broun P, Zhang JZ, Ghandehari D, Sherman BK, Yu G-L (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290(5499):2105–2110. https://doi.org/10.1126/science.290.5499.2105
Rossini L, Cribb L, Martin DJ, Langdale JA (2001) The maize golden2 gene defines a novel class of transcriptional regulators in plants. Plant Cell 13(5):1231–1244. https://doi.org/10.1105/tpc.13.5.1231
Sage RF, Sage TL, Kocacinar F (2012) Photorespiration and the evolution of C4 photosynthesis. Annu Rev Plant Biol 63:19–47. https://doi.org/10.1146/annurev-arplant-042811-105511
Savitch LV, Subramaniam R, Allard GC, Singh J (2007) The GLK1 ‘regulon’ encodes disease defense related proteins and confers resistance to Fusarium graminearum in Arabidopsis. Biochem Biophys Res Commun 359(2):234–238. https://doi.org/10.1016/j.bbrc.2007.05.084
Schindler U, Menkens AE, Beckmann H, Ecker JR, Cashmore AR (1992) Heterodimerization between light-regulated and ubiquitously expressed Arabidopsis GBF bZIP proteins. Embo j 11(4):1261–1273
Schreiber KJ, Nasmith CG, Allard G, Singh J, Subramaniam R, Desveaux D (2011) Found in Translation: high-throughput chemical screening in arabidopsis thaliana identifies small molecules that reduce fusarium head blight disease in wheat. Mol Plant-Microbe Interact® 24(6):640–648. https://doi.org/10.1094/mpmi-09-10-0210
Sedelnikova OV, Hughes TE, Langdale JA (2018) Understanding the Genetic Basis of C(4) Kranz Anatomy with a View to Engineering C(3) Crops. Annu Rev Genet 52:249–270. https://doi.org/10.1146/annurev-genet-120417-031217
Shi W, Yue L, Guo J, Wang J, Yuan X, Dong S, Guo J, Guo P (2020) Identification and evolution of C(4) photosynthetic pathway genes in plants. BMC Plant Biol 20(1):132. https://doi.org/10.1186/s12870-020-02339-x
Soltis PS, Soltis DE (2016) Ancient WGD events as drivers of key innovations in angiosperms. Curr Opin Plant Biol 30:159–165. https://doi.org/10.1016/j.pbi.2016.03.015
Swigonová Z, Lai J, Ma J, Ramakrishna W, Llaca V, Bennetzen JL, Messing J (2004) Close split of sorghum and maize genome progenitors. Genome Res 14(10a):1916–1923. https://doi.org/10.1101/gr.2332504
Wang P, Fouracre J, Kelly S, Karki S, Gowik U, Aubry S, Shaw MK, Westhoff P, Slamet-Loedin IH, Quick WP, Hibberd JM, Langdale JA (2013) Evolution of GOLDEN2-LIKE gene function in C3 and C4 plants. Planta 237(2):481–495. https://doi.org/10.1007/s00425-012-1754-3
Wang Y, Tang H, DeBarry JD, Tan X, Li J, Wang X, Lee T-h, Jin H, Marler B, Guo H, Kissinger JC, Paterson AH (2012) MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res 40(7):e49–e49. https://doi.org/10.1093/nar/gkr1293
Wood TE, Takebayashi N, Barker MS, Mayrose I, Greenspoon PB, Rieseberg LH (2009) The frequency of polyploid speciation in vascular plants. Proc Natl Acad Sci 106(33):13875–13879. https://doi.org/10.1073/pnas.0811575106
Wu S, Han B, Jiao Y (2020) Genetic contribution of paleopolyploidy to adaptive evolution in angiosperms. Mol Plant 13(1):59–71. https://doi.org/10.1016/j.molp.2019.10.012
Xiong H, Hua L, Reyna-Llorens I, Shi Y, Chen K-M, Smirnoff N, Kromdijk J, Hibberd JM (2021) Photosynthesis-independent production of reactive oxygen species in the rice bundle sheath during high light is mediated by NADPH oxidase. Proc Natl Acad Sci USA 118(25):e2022702118. https://doi.org/10.1073/pnas.2022702118
Yasumura Y, Moylan EC, Langdale JA (2005) A conserved transcription factor mediates nuclear control of organelle biogenesis in anciently diverged land plants. Plant Cell 17(7):1894–1907. https://doi.org/10.1105/tpc.105.033191
Yi F, Huo M, Li J, Yu J (2022) Time-series transcriptomics reveals a drought-responsive temporal network and crosstalk between drought stress and the circadian clock in foxtail millet. Plant J 110:1213–1228. https://doi.org/10.1111/tpj.15725
Zhang G et al (2012) Genome sequence of foxtail millet (Setaria italica) provides insights into grass evolution and biofuel potential. Nat Biotechnol 30(6):549–554. https://doi.org/10.1038/nbt.2195
Zhang J (2003) Evolution by gene duplication: an update. Trends Ecol Evol 18(6):292–298. https://doi.org/10.1016/S0169-5347(03)00033-8
Zhang X, Li X, Zhao R, Zhou Y, Jiao Y (2020) Evolutionary strategies drive a balance of the interacting gene products for the CBL and CIPK gene families. New Phytol 226(5):1506–1516. https://doi.org/10.1111/nph.16445
Funding
This work was supported by grants from the National Natural Science Foundation of China (31901552 and 32172013).
Author information
Authors and Affiliations
Contributions
HFC, XLW designed the study, HFC performed the research, analyzed the data and wrote the paper. LQ, JT and XLW modified the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Communicated by: Yuan Qin
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, H., Qin, L., Tian, J. et al. Identification and Evolutionary Analysis of the GOLDEN 2-LIKE Gene Family in Foxtail Millet. Tropical Plant Biol. 15, 301–318 (2022). https://doi.org/10.1007/s12042-022-09324-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12042-022-09324-8