Abstract
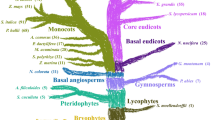
The lotus genome (Nelumbo nucifera (Gaertn.)) lacks the paleo-triplication found in other eudicots and has evolved remarkably slowly with fewer nucleotide mutations. It is thought to have greater retention of duplicated genes than other angiosperms. We evaluated the potential genes involved in cell wall synthesis and its modification, and ethylene synthesis and response. In many cell wall transferases and hydrolases families, lotus had fewer members in most families when compared to Arabidopsis. Lotus had similar or fewer members in each family as found in poplar, grape and papaya. The exceptions were in the sialyl and beta-glucuronsyl transferases where similar number were found as in the core eudicots. Lotus had similar numbers of polygalacturonase and pectin methyl esterases as found in Arabidopsis but fewer in all other hydrolases families. For starch degradation, lotus had only two alpha amylases predicted genes versus eight to ten in other eudicots, with similar numbers of beta amylase genes predicted. Lotus also had less than half the number of genes predicted for the enzymes involved in lignin and tannin synthesis compared to Arabidopsis. The stress plant growth regulator ethylene’s synthesis, reception and response predicted genes were fewer in lotus than other eudicots. Only two ethylene receptor genes were predicted in lotus with five reported for Arabidopsis and six for tomato. Our analysis does not supports the conclusion that this species has greater retention of duplicated genes though our data does support the conclusion that lotus split occurred at the base of the eudicots.

Similar content being viewed by others
References
Adams-Phillips L, Barry C, Kannan P, Lecercq J, Bouzayen M, Giovannoni JJ (2004) Ctr1-mediated ethylene signal transduction in tomato is encoded by a multigene family whose members display distinct regulatory features. Plant Mol Biol 54:387–404
Barakat A, Bagniewska-Zadworna A, Choi A, Plakkat U, DiLoreto DS, Yellanki P, Carlson JE (2009) The cinnamyl alcohol dehydrogenase gene family in Populus: phylogeny, organization, and expression. BMC Plant Biol 9:26
Barakat A, Yassin NBM, Park JS, Choi A, Herr J, Carlson JE (2011) Comparative and phylogenomic analyses of cinnamoyl-CoA reductase and cinnamoyl-CoA-reductase-like gene family in land plants. Plant Sci 181:249–257
Binder BM, Walker JM, Gagne JM, Emborg TJ, Hemmann G, Bleecker AB, Vierstra RD (2007) The Arabidopsis EIN3 binding F-Box proteins EBF1 and EBF2 have distinct but overlapping roles in ethylene signaling. Plant Cell 19:509–523
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546
Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B (2009) The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res 37:D233–D238
Carpita N, Gibeaut DM (1993) Structural models of primary cell walls in flowering plants: consistency of molecular structure with physical properties of the walls during growth. Plant J 3:1–30
Carpita NC (2011) Update on mechanisms of plant cell wall biosynthesis: how plants make cellulose and other (1-4)-ß-d-Glycans. Plant Physiol 155:171–184
Coutinho PM, Henrissat B (1999) Carbohydrate-active enzymes: an integrated database approach. In: Gilbert HJ, Davies G, Henrissat B, Svensson B (eds) Recent advances in carbohydrate bioengineering. The Royal Society of Chemistry, Cambridge, pp 3–12
Chervin C, Deluc L (2010) Ethylene signalling receptors and transcription factors over the grape berry development: gene expression profiling. Vitis 49:129–136
Chiang VL (2006) Monolignol biosynthesis and genetic engineering of lignin in trees, a review. Environ Chem Lett 4:143–146
Choi D, Cho DT, Lee Y (2006) Expansins: expanding importance in plant growth and development. Physiol Plant 126:511–518
Claus H (2004) Laccases: structure, reactions, distribution. Micron 35:93–96
Constabel CP, Yip L, Patton JJ, Christopher ME (2000) Polyphenol oxidase from hybrid poplar. cloning and expression in response to wounding and herbivory. Plant Physiol 124:285–296
Cosgrove DJ (1999) Enzymes and other agents that enhance cell wall extensibility. Annu Rev Plant Physiol Plant Mol Biol 50:391–417
Cosgrove DJ (2001) Wall structure and wall loosening. A look backwards and forwards. Plant Physiol 125:131–134
Cosgrove DJ (2012) Expansin Home Page. http://www.bio.psu.edu/expansins/index.htm. Accessed 2012 April 03.
Emanuelsson O, Nielsen H, Brunak S, von Heijne G (2000) Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol 300:1005–1016
Emanuelsson O, Brunak S, von Heijne G, Nielsen H (2007) Locating proteins in the cell using TargetP, SignalP, and related tools. Nat Protoc 2:953–971
Fry SC (2004) Primary cell wall metabolism: tracking the careers of wall polymers in living plant cells. New Phytol 161:641–675
Goubet F, Barton CJ, Mortimer JC, Yu X, Zhang Z, Miles GP, Richens J, Liepman AH, Seffen K, Dupree P (2009) Cell wall glucomannan in Arabidopsis is synthesised by CSLA glycosyltransferases, and influences the progression of embryogenesis. Plant J 60:527–538
Henrissat B, Coutinho PM, Davies GJ (2001) A census of carbohydrate-active enzymes in the genome of Arabidopsis thaliana. Plant Mol Biol 47:55–72
Hua J, Meyerowitz EM (1998) Ethylene responses are negatively regulated by a receptor gene family in Arabidopsis thaliana. Cell 94:261–271
Imsabai W, Ketsa S, van Doorn WG (2010) Role of ethylene in the lack of floral opening and in petal blackening of cut lotus (Nelumbo nucifera) flowers. Postharvest Biol Technol 58:57–64
Klee H, Tieman D (2002) The tomato ethylene receptor gene family: form and function. Physiol Plant 115:336–341
Lin ZF, Zhong SL, Grierson D (2009) Recent advances in ethylene research. J Exp Bot 60:3311–3336
Madden TL, Tatusov RL, Zhang J (1996) Application of network BLAST server. Meth Enzymol 266:131–141
Marusek CM, Trobaugh NM, Flurkey WH, Inlow JK (2006) Comparative analysis of polyphenol oxidase from plant and fungal species. J Inorg Biochem 100:108–123
Mayer AM, Staples RC (2002) Laccase: new functions for an old enzyme. Phytochem 60:551–565
Ming R. et al. (2013) Genome of the long-living sacred lotus (Nelumbo nucifera Gaertn.). Genom Biol 14:R41
Molina I, Li-Beisson Y, Beisson F, Ohlrogge JB, Pollard M (2009) Identification of an Arabidopsis feruloyl-coenzyme A transferase required for suberin synthesis. Plant Physiol 151:1317–1328
Nakano T, Suzuki K, Fujimura T, Shinshi H (2006) Genome-wide analysis of the ERF gene family in Arabidopsis and Rice. Plant Physiol 140:411–432
Nothnagel AL, Nothnagel E (2007) Primary cell wall structure in the evolution of land plants. J Integr Plant Biol 49:1271–1278
Park BH, Karpinets TV, Syed MH, Leuze MR, Uberbacher EC (2010) CAZymes Analysis Toolkit (CAT): web-service for searching and analyzing carbohydrate-active enzymes in a newly sequenced organism using CAZy database. Glycobiology 20:1574–1584, http://cricket.ornl.gov/cgi-bin/cat.cgi
Passardi F, Longet D, Penel C, Dunand C (2004) The class III peroxidase multigenic family in rice and its evolution in land plants. Phytochem 65:1879–1893
Popper ZA (2008) Evolution and diversity of green plant cell walls. Curr Opin Plant Biol 11:286–292
Pourcel L, Routaboul JM, Cheynier V, Lepiniec L, Debeaujon I (2006) Flavonoid oxidation in plants: from biochemical properties to physiological functions. Trends Plant Sci 12:29–36
Rodríguez FI, Esch JJ, Hall AE, Binder BM, Schaller GE, Bleecker AB (1999) A copper cofactor for the ethylene receptor ETR1 from Arabidopsis. Science 283:996–998
Saito N, Nei M (1987) The neighbor-joining method: a new method for constructing phylogenetic trees. Molec Biol Evol 4:406–425
Sampedro J, Cosgrove DJ (2005) The expansin superfamily. Genom Biol 6:242
Sampedro J, Lee Y, Carey RE, de Pamphilis C, Cosgrove DJ (2005) Use of genomic history to improve phylogeny and understanding of births and deaths in a gene family. Plant J 44:409–419
Sarkar P, Bosneaga E, Auer M (2009) Plant cell walls throughout evolution: towards a molecular understanding of their design principles. J Exp Bot 60:3615–3635
Schaller GE, Kieber JJ (2002) Ethylene. In: Somerville C, Meyerowitz E (eds) The arabidopsis book. American Society of Plant Biologists, Rockville
Shaw MF (1929) A microchemical study of the fruit coat of Nelumbo lutea. Am J Bot 16:259–276
Shen-Miller J, Mudgett MB, Schopf JW, Clarke S, Berger R (1995) Exceptional seed longevity and robust growth: ancient sacred lotus from China. Am J Bot 82:1367–1380
Shen-Miller J, Schopf JW, Harbottle G, Cao R-J, Ouyang S, Zhou K-S, Southon JR, Liu G-H (2002) Long-living lotus: germination and soil γ-irradiation of centuries-old fruits, and cultivation, growth, and phenotypic abnormalities of offspring. Am J Bot 89:236–247
Small I, Peeters N, Legeai F, Lurin C (2004) Predotar: a tool for rapidly screening proteomes for N-terminal targeting sequences. Proteomics 4:1581–1590
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Molec Biol Evol 24:1596–1599
Thipyapong P, Joel DM, Steffens JC (1997) Differential expression and turnover of the tomato polyphenol oxidase gene family during vegetative and reproductive development. Plant Physiol 113:707–718
Tompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Ac Res 22:4673–4680
Turlapati PV, Kim KW, Davin LB, Lewis NG (2011) The laccase multigene family in Arabidopsis thaliana: towards addressing the mystery of their gene function(s). Planta 233:439–470
Valerio L, De Meyer M, Penel C, Dunand C (2004) Expression analysis of the Arabidopsis peroxidase multigenic family. Phytochem 65:1331–1342
Van Bergen PF, Hatcher PG, Boon JJ, Collinson ME, de Leeuw JW (1997) Macromolecular composition of the propagule wall of Nelumbo nucifera. Phytochemistry 45:601–610
Vanholme R, Morreel K, Ralph J, Boerjan W (2008) Lignin engineering. Curr Opin Plant Biol 11:278–285
Vanholme R, Demedts B, Morreel K, Ralph J, Boerjan W (2010) Lignin biosynthesis and structure. Plant Physiol 153:895–905
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by: Graham Bonnett
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
Lotus Cell wall Carbohydrate Enzymes (XLSX 92 kb)
Supplementary Table 2
Lotus Expansins (XLSX 34 kb)
Supplementary Table 3
Lotus Ethylene Biosynthesis (XLSX 12 kb)
Supplementary Table 4
Lotus Laccase, PPO, Peroxidase (XLSX 15 kb)
Supplementary Table 5
Lotus Terpene synthesis (XLSX 51 kb)
Supplementary Table 6
Lotus Tannin Synthesis (XLSX 14 kb)
Supplementary Figure 1
(DOCX 1203 kb)
Supplementary Table 7
(PDF 132 kb)
Rights and permissions
About this article
Cite this article
Paull, R.E., Carroll, A. & Chen, N.J. Lotus Cell Walls and the Genes Involved in its Synthesis and Modification. Tropical Plant Biol. 6, 152–160 (2013). https://doi.org/10.1007/s12042-013-9129-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12042-013-9129-x