Abstract
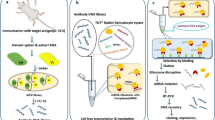
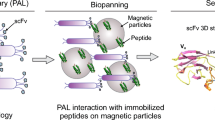
The aim of this study was to construct a ribosome display library of single chain variable fragments (scFvs) associated with hepatocarcinoma and screen such a library for hepatocarcinoma-binding scFvs. mRNA was isolated from the spleens of mice immunized with hepatocellular carcinoma cell line HepG2. Heavy and k chain genes (VH and k) were amplified separately by RT-PCR, and an anti-HepG2 VH/k chain ribosome display library was constructed by assembling VH and k into the VH/k chain with a specially constructed linker by SOE-PCR. The VH/k chain library was transcribed and translated in vitro using a rabbit reticulocyte lysate system. In order to isolate specific scFvs, recognizing HepG2 negative selection on a normal hepatocyte line WRL-68 was carried out before three rounds of positive selection on HepG2. After three rounds of panning, cell enzyme-linked immunosorbent assay (ELISA) showed that one of the scFvs had high affinity for the HepG2 cell and lower affinity for the WRL-68 cell. In this study, we successfully constructed a native ribosome display library. Such a library would prove useful for direct intact cell panning using ribosome display technology. The selected scFv had a potential value for hepatocarcinoma treatment.
Similar content being viewed by others
Abbreviations
- ARM:
-
antibody-ribosome-mRNA
- BSA:
-
bovine serum albumin
- CDR:
-
complementarity-determining region
- DMEM:
-
Dulbecco modified Eagle high glucose medium
- ELISA:
-
enzyme-linked immunosorbent assay
- FBS:
-
foetal bovine serum
- IPTG:
-
isopropyl-b-D-thiogalactopyranoside
- PBS:
-
phosphate buffered saline
- RT-PCR:
-
reverse transcriptase-polymerase chain reaction
- scFv:
-
single-chain variable fragment
- SDS-PAGE:
-
sodium dodecyl sulphate-polyacrylamide gel electrophoresis
- VH:
-
variable region of heavy chain
References
Bradford M M 1976 A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding; Anal. Biochem. 72 248–254
Gao C, Mao S, Ronca F, Zhuang S, Quaranta V, Wirsching P and Janda K D 2003 De novo identification of tumor-specific internalizing human antibody-receptor pairs by phage-display methods; J. Immunol. Methods 274 185–197
Hanes J, Jermutus L, Weber-Bornhauser S, Bosshard H R and Plückthun A 1998 Ribosome display efficiently selects and evolves high-affinity antibodies in vitro from immune libraries; Proc. Natl. Acad. Sci. USA 95 14130–14135
He M and Taussig M J 1997 Antibody-ribosome-mRNA (ARM) complexes as efficient selection particles for in vitro display and evolution of antibody combining sites; Nucleic Acids Res. 25 5132–5134
He M and Taussig M J 2005 Ribosome display of antibodies: expression, specificity and recovery in a eukaryotic system; J. Immunol. Methods 297 73–82
He M, Menges M, Groves M A, Corps E, Liu H, Brüggemann M and Taussig M J 1999 Selection of a human anti-progesterone antibody fragment from a transgenic mouse library by ARM ribosome display; J. Immunol. Methods 231 105–117
Irving R A, Coia G, Roberts A, Nuttall S D and Hudson P J 2001 Ribosome display and affinity maturation: from antibodies to single V-domains and steps towards cancer therapeutics; J. Immunol. Methods 248 31–45
Lee M S, Kwon M H, Kim K H, Shin H J, Park S and Kim H I 2005 Selection of scFvs specific for HBV DNA polymerase using ribosome display; J. Immunol. Methods 284 147–157
Melén K, Keskinen P, Lehtonen A and Julkunen I 2000 Interferon-induced gene expression and signaling in human hepatoma cell lines; J. Hepatol. 33 764–772
Rothe A, Nathanielsz A, Hosse R J, Oberhauser F, Strandmann E P, Engert A, Hudson P R and Power B E 2007 Selection of human anti-CD28 scFvs from a T-NHL related scFv library using ribosome display; J. Biotechnol. 130 448–454
Taussig M J, Stoevesandt O, Borrebaeck C A, Bradbury A R, Cahill D, Cambillau C, de Daruvar A, Dübel S, et al. 2007 Proteome binders: planning a European resource of affinity reagents for analysis of the human proteome; Nat. Methods 4 13–17
van Dijk M A and van de Winkel J G 2001 Human antibodies as next generation therapeutics; Curr. Opin. Chem. Biol. 5 368–374
Verhaar M J, Chester K A, Keep P A, Robson L, Pedley R B, Boden J A, Hawkins R E and Begent R H 1995 A single chain Fv derived from a filamentous phage library has distinct tumor targeting advantages over one derived from a hybridoma; Int. J. Cancer 61 497–501
Yau K Y, Groves M A, Li S, Sheedy C, Lee H, Tanha J, MacKenzie C R, Jermutus L and Hall JC 2003 Selection of hapten-specific single-domain antibodies from a non-immunized llama ribosome display library; J. Immunol. Methods 281 161–175
Zhu X, Wu H, Luo S, Xianyu Z Q and Zhu D 2008 Screening and identification of a novel hepatocellular carcinoma cell binding peptide by using a phage display library; J. Huazhong Univ. Sci. Technol. Med. Sci. 28 299–303
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhou, L., Mao, WP., Fen, J. et al. Selection of scFvs specific for the HepG2 cell line using ribosome display. J Biosci 34, 221–226 (2009). https://doi.org/10.1007/s12038-009-0026-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12038-009-0026-2