Abstract
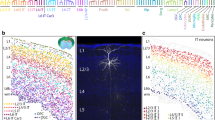
The inhibitory neurons in the brain play an essential role in neural network firing patterns by releasing γ-aminobutyric acid (GABA) as the neurotransmitter. In the mouse brain, based on the protein molecular markers, inhibitory neurons are usually to be divided into three non-overlapping groups: parvalbumin (PV), neuropeptide somatostatin (SST), and vasoactive intestinal peptide (VIP)-expressing neurons. Each neuronal group exhibited unique properties in molecule, electrophysiology, circuitry, and function. Calbindin 1 (Calb1), a ubiquitous calcium-binding protein, often acts as a “divider” in excitatory neuronal classification. Based on Calb1 expression, the excitatory neurons from the same brain region can be classified into two subgroups with distinct properties. Besides excitatory neurons, Calb1 also expresses in part of inhibitory neurons. But, to date, little research focused on the intersectional relationship between inhibitory neuronal subtypes and Calb1. In this study, we genetically targeted Calb1-expression (Calb1+) and Calb1-lacking (Calb1−) subgroups of PV and SST neurons throughout the mouse brain by flexibly crossing transgenic mice relying on multi-recombinant systems, and the distribution patterns and electrophysiological properties of each subgroup were further demonstrated. Thus, this study provided novel insights and strategies into inhibitory neuronal classification.





Similar content being viewed by others
Data Availability
The datasets during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Zeng H, Sanes JR (2017) Neuronal cell-type classification: challenges, opportunities and the path forward. Nat Rev Neurosci 18(9):530–546. https://doi.org/10.1038/nrn.2017.85
Tsien JZ, Chen DF, Gerber D, Tom C, Mercer EH, Anderson DJ, Mayford M, Kandel ER et al (1996) Subregion- and cell type-restricted gene knockout in mouse brain. Cell 87(7):1317–1326. https://doi.org/10.1016/s0092-8674(00)81826-7
Tsien JZ, Huerta PT, Tonegawa S (1996) The essential role of hippocampal CA1 NMDA receptor-dependent synaptic plasticity in spatial memory. Cell 87(7):1327–1338. https://doi.org/10.1016/s0092-8674(00)81827-9
Frantz GD, Tobin AJ (1994) Cellular distribution of calbindin D28K mRNAs in the adult mouse brain. J Neurosci Res 37(3):287–302. https://doi.org/10.1002/jnr.490370302
Lewis DA, Hashimoto T, Volk DW (2005) Cortical inhibitory neurons and schizophrenia. Nat Rev Neurosci 6(4):312–324. https://doi.org/10.1038/nrn1648
Poulin JF, Caronia G, Hofer C, Cui Q, Helm B, Ramakrishnan C, Chan CS, Dombeck DA et al (2018) Mapping projections of molecularly defined dopamine neuron subtypes using intersectional genetic approaches. Nat Neurosci 21(9):1260–1271. https://doi.org/10.1038/s41593-018-0203-4
Sürmeli G, Marcu DC, McClure C, Garden DLF, Pastoll H, Nolan MF (2015) Molecularly defined circuitry reveals input-output segregation in deep layers of the medial entorhinal cortex. Neuron 88(5):1040–1053. https://doi.org/10.1016/j.neuron.2015.10.041
Blatow M, Rozov A, Katona I, Hormuzdi SG, Meyer AH, Whittington MA, Caputi A, Monyer H (2003) A novel network of multipolar bursting interneurons generates theta frequency oscillations in neocortex. Neuron 38(5):805–817. https://doi.org/10.1016/s0896-6273(03)00300-3
Kitamura T, Pignatelli M, Suh J, Kohara K, Yoshiki A, Abe K, Tonegawa S (2014) Island cells control temporal association memory. Science (New York, NY) 343(6173):896–901. https://doi.org/10.1126/science.1244634
Li Y, Xu J, Liu Y, Zhu J, Liu N, Zeng W, Huang N, Rasch MJ et al (2017) A distinct entorhinal cortex to hippocampal CA1 direct circuit for olfactory associative learning. Nat Neurosci 20(4):559–570. https://doi.org/10.1038/nn.4517
Masurkar AV, Srinivas KV, Brann DH, Warren R, Lowes DC, Siegelbaum SA (2017) Medial and lateral entorhinal cortex differentially excite deep versus superficial CA1 pyramidal neurons. Cell Rep 18(1):148–160. https://doi.org/10.1016/j.celrep.2016.12.012
Soltesz I, Losonczy A (2018) CA1 pyramidal cell diversity enabling parallel information processing in the hippocampus. Nat Neurosci 21(4):484–493. https://doi.org/10.1038/s41593-018-0118-0
Rudy B, Fishell G, Lee S, Hjerling-Leffler J (2011) Three groups of interneurons account for nearly 100% of neocortical GABAergic neurons. Dev Neurobiol 71(1):45–61. https://doi.org/10.1002/dneu.20853
Daigle TL, Madisen L, Hage TA, Valley MT, Knoblich U, Larsen RS, Takeno MM, Huang L et al (2018) A suite of transgenic driver and reporter mouse lines with enhanced brain-cell-type targeting and functionality. Cell 174(2):465–480.e422. https://doi.org/10.1016/j.cell.2018.06.035
Madisen L, Zwingman TA, Sunkin SM, Oh SW, Zariwala HA, Gu H, Ng LL, Palmiter RD et al (2010) A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat Neurosci 13(1):133–140. https://doi.org/10.1038/nn.2467
Pfeffer CK, Xue M, He M, Huang ZJ, Scanziani M (2013) Inhibition of inhibition in visual cortex: the logic of connections between molecularly distinct interneurons. Nat Neurosci 16(8):1068–1076. https://doi.org/10.1038/nn.3446
Tremblay R, Lee S, Rudy B (2016) GABAergic interneurons in the neocortex: from cellular properties to circuits. Neuron 91(2):260–292. https://doi.org/10.1016/j.neuron.2016.06.033
Markram H, Toledo-Rodriguez M, Wang Y, Gupta A, Silberberg G, Wu C (2004) Interneurons of the neocortical inhibitory system. Nat Rev Neurosci 5(10):793–807. https://doi.org/10.1038/nrn1519
Gouwens NW, Sorensen SA, Baftizadeh F, Budzillo A, Lee BR, Jarsky T, Alfiler L, Baker K et al (2020) Integrated morphoelectric and transcriptomic classification of cortical GABAergic cells. Cell 183(4):935–953.e919. https://doi.org/10.1016/j.cell.2020.09.057
Gibson JR, Beierlein M, Connors BW (1999) Two networks of electrically coupled inhibitory neurons in neocortex. Nature 402(6757):75–79. https://doi.org/10.1038/47035
Fanselow MS, Dong HW (2010) Are the dorsal and ventral hippocampus functionally distinct structures? Neuron 65(1):7–19. https://doi.org/10.1016/j.neuron.2009.11.031
Floriou-Servou A, von Ziegler L, Stalder L, Sturman O, Privitera M, Rassi A, Cremonesi A, Thöny B et al (2018) Distinct proteomic, transcriptomic, and epigenetic stress responses in dorsal and ventral hippocampus. Biol Psychiatry 84(7):531–541. https://doi.org/10.1016/j.biopsych.2018.02.003
Pelkey KA, Chittajallu R, Craig MT, Tricoire L, Wester JC, McBain CJ (2017) Hippocampal GABAergic inhibitory interneurons. Physiol Rev 97(4):1619–1747. https://doi.org/10.1152/physrev.00007.2017
Mann EO, Paulsen O (2007) Role of GABAergic inhibition in hippocampal network oscillations. Trends Neurosci 30(7):343–349. https://doi.org/10.1016/j.tins.2007.05.003
Liu YQ, Yu F, Liu WH, He XH, Peng BW (2014) Dysfunction of hippocampal interneurons in epilepsy. Neurosci Bull 30(6):985–998. https://doi.org/10.1007/s12264-014-1478-4
Laure-Kamionowska M, Maślińska D (2009) Calbindin positive Purkinje cells in the pathology of human cerebellum occurring at the time of its development. Folia Neuropathol 47(4):300–305
Schmidt H, Schwaller B, Eilers J (2005) Calbindin D28k targets myo-inositol monophosphatase in spines and dendrites of cerebellar Purkinje neurons. Proc Natl Acad Sci U S A 102(16):5850–5855. https://doi.org/10.1073/pnas.0407855102
Vig PJ, Subramony SH, Burright EN, Fratkin JD, McDaniel DO, Desaiah D, Qin Z (1998) Reduced immunoreactivity to calcium-binding proteins in Purkinje cells precedes onset of ataxia in spinocerebellar ataxia-1 transgenic mice. Neurology 50(1):106–113. https://doi.org/10.1212/wnl.50.1.106
Schwaller B, Meyer M, Schiffmann S (2002) ‘New’ functions for ‘old’ proteins: the role of the calcium-binding proteins calbindin D-28k, calretinin and parvalbumin, in cerebellar physiology. Studies with knockout mice. Cerebellum 1(4):241–258. https://doi.org/10.1080/147342202320883551
Brandenburg C, Smith LA, Kilander MBC, Bridi MS, Lin YC, Huang S, Blatt GJ (2021) Parvalbumin subtypes of cerebellar Purkinje cells contribute to differential intrinsic firing properties. Mol Cell Neurosci 115:103650. https://doi.org/10.1016/j.mcn.2021.103650
Thankachan S, Katsuki F, McKenna JT, Yang C, Shukla C, Deisseroth K, Uygun DS, Strecker RE et al (2019) Thalamic reticular nucleus parvalbumin neurons regulate sleep spindles and electrophysiological aspects of schizophrenia in mice. Sci Rep 9(1):3607. https://doi.org/10.1038/s41598-019-40398-9
Steullet P, Cabungcal JH, Bukhari SA, Ardelt MI, Pantazopoulos H, Hamati F, Salt TE, Cuenod M et al (2018) The thalamic reticular nucleus in schizophrenia and bipolar disorder: role of parvalbumin-expressing neuron networks and oxidative stress. Mol Psychiatry 23(10):2057–2065. https://doi.org/10.1038/mp.2017.230
Ahrens S, Jaramillo S, Yu K, Ghosh S, Hwang GR, Paik R, Lai C, He M et al (2015) ErbB4 regulation of a thalamic reticular nucleus circuit for sensory selection. Nat Neurosci 18(1):104–111. https://doi.org/10.1038/nn.3897
Abrahams S, Morris RG, Polkey CE, Jarosz JM, Cox TC, Graves M, Pickering A (1999) Hippocampal involvement in spatial and working memory: a structural MRI analysis of patients with unilateral mesial temporal lobe sclerosis. Brain Cogn 41(1):39–65. https://doi.org/10.1006/brcg.1999.1095
Ben-Simon Y, Kaefer K, Velicky P, Csicsvari J, Danzl JG, Jonas P (2022) A direct excitatory projection from entorhinal layer 6b neurons to the hippocampus contributes to spatial coding and memory. Nat Commun 13(1):4826. https://doi.org/10.1038/s41467-022-32559-8
Décarie-Spain L, Liu CM, Lauer LT, Subramanian K, Bashaw AG, Klug ME, Gianatiempo IH, Suarez AN et al (2022) Ventral hippocampus-lateral septum circuitry promotes foraging-related memory. Cell Rep 40(13):111402. https://doi.org/10.1016/j.celrep.2022.111402
Netsyk O, Hammoud H, Korol SV, Jin Z, Tafreshiha AS, Birnir B (2020) Tonic GABA-activated synaptic and extrasynaptic currents in dentate gyrus granule cells and CA3 pyramidal neurons along the mouse hippocampal dorsoventral axis. Hippocampus 30(11):1146–1157. https://doi.org/10.1002/hipo.23245
Li X, Chen W, Yu Q, Zhang Q, Zhang T, Huang X, Li H, He A et al (2021) A circuit of mossy cells controls the efficacy of memory retrieval by Gria2I inhibition of Gria2. Cell Rep 34(7):108741. https://doi.org/10.1016/j.celrep.2021.108741
K PGaF (2019) The mouse brain in stereotaxic coordinates. Elsevier
Li X, Yu H, Zhang B, Li L, Chen W, Yu Q, Huang X, Ke X et al (2022) Molecularly defined and functionally distinct cholinergic subnetworks. Neuron 110(22):3774–3788.e3777. https://doi.org/10.1016/j.neuron.2022.08.025
Li X, Chen W, Pan K, Li H, Pang P, Guo Y, Shu S, Cai Y et al (2018) Serotonin receptor 2c-expressing cells in the ventral CA1 control attention via innervation of the Edinger-Westphal nucleus. Nat Neurosci 21(9):1239–1250. https://doi.org/10.1038/s41593-018-0207-0
Jing W, Zhang T, Liu J, Huang X, Yu Q, Yu H, Zhang Q, Li H et al (2021) A circuit of COCH neurons encodes social-stress-induced anxiety via MTF1 activation of Cacna1h. Cell Rep 37(13):110177. https://doi.org/10.1016/j.celrep.2021.110177
He A, Zhang C, Ke X, Yi Y, Yu Q, Zhang T, Yu H, Du H et al (2022) VGLUT3 neurons in median raphe control the efficacy of spatial memory retrieval via ETV4 regulation of VGLUT3 transcription. Sci China Life Sci 65(8):1590–1607. https://doi.org/10.1007/s11427-021-2047-8
Acknowledgements
We thank Prof. Miao He of Fudan University for Ai65-F and Ai65 mice.
Funding
This work was supported by the National Natural Science Foundation of China (Grants: 82271486 to XL; 81800133 to AH).
Author information
Authors and Affiliations
Contributions
XL, AH, and YL conceived and designed the studies. BZ, LL, and XT carried out the experiments, including cell typing, staining and image acquisition, cell counting, and analysis of the data. JZ, YS, ZH, TM, and HL performed the genotyping, electrophysiology, and data analysis experiments. The manuscript was written by HA, AH, and XL. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Ethics Approval and Consent to Participate
Mice were bred and reared under the same conditions in accordance with institutional guidelines and the Animal Care and Use Committee of the Animal Core Facility at Huazhong University of Science and Technology. All animal experiments were reviewed and approved by the Medical Ethics Committee at Huazhong University of Science and Technology.
Consent for Publication
All involved parties consent to publication.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, B., Li, L., Tang, X. et al. Distribution Patterns of Subgroups of Inhibitory Neurons Divided by Calbindin 1. Mol Neurobiol 60, 7285–7296 (2023). https://doi.org/10.1007/s12035-023-03542-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-023-03542-1