Abstract
Since more than a century, neuroscientists have distinguished excitatory (glutamatergic) neurons with long-distance projections from inhibitory (GABAergic) neurons with local projections and established layer-dependent schemes for the ~ 80% excitatory (principal) cells as well as the ~ 20% inhibitory neurons. Whereas, in the early days, mainly morphological criteria were used to define cell types, later supplemented by electrophysiological and neurochemical properties, nowadays. single-cell transcriptomics is the method of choice for cell type classification. Bringing recent insight together, we conclude that despite all established layer- and area-dependent differences, there is a set of reliably identifiable cortical cell types that were named (among others) intratelencephalic (IT), extratelencephalic (ET), and corticothalamic (CT) for the excitatory cells, which altogether comprise ~ 56 transcriptomic cell types (t-types). By the same means, inhibitory neurons were subdivided into parvalbumin (PV), somatostatin (SST), vasoactive intestinal polypeptide (VIP), and “other (i.e. Lamp5/Sncg)” subpopulations, which altogether comprise ~ 60 t-types. The coming years will show which t-types actually translate into “real” cell types that show a common set of multimodal features, including not only transcriptome but also physiology and morphology as well as connectivity and ultimately function. Only with the better knowledge of clear-cut cell types and experimental access to them, we will be able to reveal their specific functions, a task which turned out to be difficult in a part of the brain being so much specialized for cognition as the cerebral cortex.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
General introduction
The mammalian brain is a complex organ which comprises billions of neurons that altogether were recently suggested to consist of several thousand types of neurons [28]. The neurons of the cerebral cortex collectively shape the computations within networks that represent mental activities and govern behavior. The most practical means of navigating this complexity and thus unraveling the mysteries of neuronal diversity that contribute to cortical circuit structure and function is by classifying neurons into distinct types [39,40,41]. Therefore, the classification of neuronal cell types has been recognized as the pivotal pursuit within the realm of neuroscience research.
Historically, by means of histological stains, pioneers like Ramon y Cajal and Lorente de Nó relied on morphological criteria to identify cortical cell types [39]. In recent decades, the field has witnessed a transformative shift toward more sophisticated methods. Electrophysiological techniques brought functional insights into the classification, while neurochemical markers provided additional layers of characterization. However, it was the groundbreaking development of single-cell transcriptomics that overhauled laborious and low-throughput conventional approaches and revolutionized our ability to profile individual neurons at the most basic genetic level. This approach enables the identification of previously unknown cell types, revealing their unique genetic signatures and paving the way for a more comprehensive and nuanced classification of cortical neuronal cell types [40, 41].
Although strong consistency is observed among categories established through morphological, molecular, and physiological criteria [9, 26, 36], no single approach exists to fully encompass the inherently multimodal attributes of cell phenotypes and establish a universal standard for classification. In this review, we will mainly use the mouse brain as an example to shed light on the current status of cortical cell type classification.
Current status of excitatory cell type classification
In the six-layered mouse cortex, excitatory neurons make up the majority of the neuronal population, constituting approximately 80–85% of all neurons. Presently, they are categorized into 9 subclasses based on their soma localization within specific layers (L) and their patterns of projection [2, 17, 42] (Figs. 1 and 2 and Table 1) intratelencephalic projecting (IT), extratelencephalic projecting (ET), near-projecting (NP), and corticothalamic projecting (CT): L2/3 IT, L4/5 IT, L5 IT, L6 IT, Car3 IT, L5 ET, L5/6NP, L6 CT, and L6b. These neuron types can further vary according to their transcription profiles.
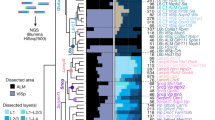
Comprehensive account of cortical cell types exemplified in the mouse primary motor cortex (MOp) by cluster analysis and spatial transcriptomics (MERFISH). a Glutamatergic, GABAergic neuronal and non-neuronal cells cluster dendrogram giving an overview of major groups and cell types, which are colored for their identity. Please note that GABAergic neurons not only cluster by major molecular markers but also by developmental origin (MGE: medial ganglionic eminence; CGE: caudal ganglionic eminence). b Spatial distribution of cell types identified by clustering (in a) shown with the same color code by spatial transcriptomics. Please note that due to layer identity being a major factor for the majority cell type of pyramidal neurons, a laminar pattern becomes apparent. Here, a section through MOp is displayed whose location is shown in the upper right corner. c This “six-layered” neocortical laminar pattern becomes even more crisp when the distribution of glutamatergic neurons alone is depicted. Inset: a biocytin-filled L2/3 IT (pyramidal) neuron of unknown t-type is shown, which has a typical spiny dendritic tree with an apical dendritic tuft in L1 and an axon running toward the white matter (Preuss, Witte, Staiger; unpublished material). Scale bars, 200 μm for a and b and 100 μm for the inset. For neuronal cell names, see Tables; for non-neuronal: OGC: oligodendrocyte; OPC: oligodendrocyte precursor cell; PVM: perivascular macrophage; SMC: smooth muscle cell; VLMC: vascular leptomeningeal cell. Modified from BRAIN Initiative Cell Census Network, Nature 2021; with permission
In-depth multimodal account of cortical neuronal cell types. a Hierarchical cluster tree showing the consensus transcriptomic taxonomy on mouse neocortical neurons. Major cellular divisions and class and subclass labels are shown above major branches and cluster labels are shown below each leaf node. Using Patch-seq and connectivity studies, many transcriptomic neuron types or subtypes are annotated and correlated with previously established cortical neuron types (see also Table 2). Relative proportions of all cell types are calculated from the snRNA-seq 10x v3 data (bar graphs below cluster tree leaves). The box to the right of the tree presents a UMAP representation of the mouse transcriptomic–epigenomic integrated molecular taxonomy (SCF version). b Exemplar candidates for somatodendritic (dark hues, often hidden by axon) and axonal arborizations (brighter hues) of various GABAergic and glutamatergic cell types as derived from Patch-seq studies. b Different morphologies of “classical” neocortical cell types. Left: Two L2/3 basket cells (fast spiking; not shown) are labeled in primary somatosensory cortex and visualized by high-resolution confocal imaging. For the upper large basket cell, a reconstruction is overlaid in which the somatodendritic compartment is shown in cyan and the axonal compartment in red. Please not the dense local axons with some descending collaterals (Preuss, Witte, Staiger; unpublished material), which also exist in the more compact small basket not superimposed by a reconstruction. Scale bar: 100 μm. Intermediate: 5 superimposed L5 IT neurons are visualized on a surface reconstruction of the mouse brain. Their axonal arbors project to ipsilateral and contralateral cortex as well as to striatum. Right: Superimpositions of 6 myelencephalon/medulla oblongata-projecting (L5 ET MY-projecting ) versus 6 non-myelencephalon-projecting (L5 ET non-MY-projecting) pyramidal cells are contrasted for their different target regions. Modified from BRAIN Initiative Cell Census Network, Nature 2021; with permission
IT neurons
They were previously called cortico-cortical, cortico-callosal, cortico-striatal, or callosal projection neurons, which are primarily located in layers 2/3, 4, and 5, with fewer numbers found in layer 6. Now, we know that these are not separate neuron types but that the same individual IT neurons project to multiple cortical areas both ipsilaterally and contralaterally through the corpus callosum or the anterior commissure as well as to the striatum [13]. Featuring thin-tufted apical dendrites, L2/3 IT neurons send their major descending interlaminar axonal projection into L5 (Fig. 1C, inset) [11]. They fire sparsely as a consequence of a hyperpolarized resting membrane potential. L4 IT neurons encompass various morphological subclasses, including pyramidal, star-pyramidal, and spiny stellate forms. Interestingly, although primary motor cortex (MOp) lacks a cytoarchitectonic layer 4, neurons resembling those traditionally defined as L4 neurons in sensory cortical regions are also observed [2, 37]. L5 IT neurons are more active than L2/3 IT and have broader projections including more extensive connections to striatum [31]. L6 ITs preferentially establish reciprocal connections with local deep layer neurons [11]. L6 Car3 neurons, having the highest number of targets, possess extensive intracortical axonal projections like other IT neurons, yet lack collateral projections into the striatum [24].
L5 ET neurons
They are alternatively known as pyramidal tract (PT) or subcerebral projection neurons (SCPN), exhibit thick-tufted dendrites and project to multiple subcortical regions, including the “higher-order” thalamus, midbrain, hindbrain, and the spinal cord (Fig. 2c). They typically display the electrophysiological characteristics of bursts of action potentials leading to a strong impact on their target neurons. Compared to IT neurons, L5 ETs often present with a more depolarized resting membrane potential, lower input resistance, a faster effective membrane time constant, and less spike frequency adaption [1].
Other neuronal types
L5/6 NP neurons project only sparsely to neighboring regions [32]. Found in layer 6, CT neurons provide feedback projections to the thalamic “relay/first-order” nuclei. In line with their scarcity of local inputs, the majority are notably silent in vivo [11]. L6b subplate neurons reveal projections to L1 within resident and adjacent cortical areas [32], their function in the adult brain remaining enigmatic.
Attempts have been made to classify cortical excitatory neurons using molecular profiles. A varied combination of layer-specific marker genes provided early evidence for the correlation between gene expression and target specificity at the subclass level [13]. Recent advances in single-cell transcriptomics have empowered researchers not only to unveil new markers such as Deptor for L5 IT, Slc17a8 for L5/6 NP [23], Osr1 for L6 IT, and Fam84b for L5 ET [32], but also to discern finer distinctions within the framework of major subclasses (see also Table 1). Then, 19 transcriptomic cell types were identified in primary visual cortex (V1) [31], whereas unsupervised clustering analysis of MERFISH-derived profiles revealed 39 types in primary motor cortex (Mop) [42] (Fig. 1). A combined total of 56 t-types have been discovered in both V1 and the anterior lateral motor cortex (ALM) [32]. The difference is in part due to regional variation but also possibly owns to more extensive cell sampling in ALM. Notably, cortical area differences do not exhibit uniformity across various subclasses, as is shown by L5 ET and L4/5 IT types, which display more pronounced distinctions across areas compared to types in other subclasses. However, it is believed that most types are shared among multiple areas [38]. An apparent difference in t-types between 2 distant areas (primary motor versus primary visual cortex) changes to a more continuous pattern on the global cortical scale. This graded feature is also found across the cortical depth [2, 13, 42]. L2/3-L6 IT types exhibit a continuum marked by gradual changes in gene expression profiles and their positioning within cortical layers. Types with more similar transcriptomes are located at neighboring cortical depths but do not necessarily respect laminar boundaries (Fig. 1c).
Similarly, gradual variation is present in both electrophysiological and morphological properties. In L2/3 of somatosensory and visual cortex, the dendritic tree structure, electrophysiological characteristics and connectional patterns continuously co-vary with pial depth [29, 34]. Results from Patch-seq indicate that properties can exhibit significant variations even within a specific electrophysiological type (e-type) or morphological type (m-type) [9, 26]. Morpho-electric properties exhibit continuous variation across the transcriptomic landscape, suggesting the absence of a clear point to subdivide the cluster. As a result, a one-to-several relationship was observed between t-types like L4-IT–Rspo1 and morpho-electric (me) types, and vice versa, although the integration of both morphological and electrical characteristics exhibited stronger correlations with t-type than examining each modality individually [10].
The exact relationship between projections and t-types remains to be clarified. Nevertheless, a strong alignment exists between major transcriptomic and projection neuron types, as evidenced by the Cre-defined projection mapping strategy [16, 19]. For example, Scnn1a-L4/5 and Ntsr1-L6 neurons exclusively represent L4/5 IT and L6 CT, respectively. Yet, within each subclass, there is no guarantee of a one-to-one correspondence between the finely graded projection pattern and the cell’s t-type, although certain instances like Medulla (MY)-projecting and non-MY-projecting types within L5 ET subclass do map to distinct taxonomic clusters [6]. By contrast, V1 neurons projecting to PM (posteromedial) and V1 neurons projecting to LM (lateromedial) (V1/PM and V1/LM) only form a single genetic cluster [14]. Another noteworthy observation was that neurons of the same t-type may exhibit distinct sets of projection targets specific to their respective home region [24]. This reinforces the notion that transcriptomic classification alone may be insufficient to capture all the heterogeneity within excitatory neuronal types. Therefore, future research endeavors should prioritize the investigation of how molecular, electrophysiological, morphological characteristics, and projection targets interact when defining cell types.
Current status of inhibitory cell type classification
While GABAergic interneurons comprise only 15–20% of cortical neurons, they exhibit the most extensive diversity in terms of morphology, electrophysiology, and neurochemical characteristics. By the 2005 Petilla Convention, a consensus nomenclature was put forth to improve the characterization of interneurons, yet failed to live up to expectations. Frequently, the same name was assigned to neurons with differing morphologies, and various terminologies were inconsistently adopted across different laboratories to describe the same cell classification [5, 25]. Decades later, single-cell transcriptomics has undertaken the challenge of resolving this long-standing issue and now stands as the anchor for defining cell types. Although similar to excitatory neurons, a degree of continuous variation exists within each t-type and the correspondence between t-types and electrophysiological or morphological features is not strictly one-to-one [9, 26, 32], 28 morphological/electrophysiological/transcriptomic types (met-types) of cortical interneurons with congruent properties were identified in V1 [9]. Here, we will highlight several widely accepted types, ignoring putative projection neurons and the still poorly studied Sncg neurons (Figs. 1, and 2 and Table 2).
The initial branches of the hierarchical t-type tree diagram distinguish between excitatory glutamatergic and inhibitory GABAergic classes, probably underlining their distinct developmental origins in the embryonic pallium and subpallial regions (Fig. 1a). The GABAergic branch further separates into 5 subclasses originating from the medial and caudal ganglionic eminence (MGE and CGE) based on marker molecules: parvalbumin (PV) and somatostatin (SST) in MGE and vasoactive intestinal polypeptide (VIP), Lamp5, and Sncg in CGE [32, 42].
Within the PV population, two well established anatomical subtypes stand out: fast-spiking basket cells (PV-BCs) and chandelier cells (ChCs). BCs target the perisomatic domain of pyramidal cells [7]. Contributing significantly to the cortical excitation-inhibition balance, they are credited with various perception-related functions, such as the generation of gamma-band oscillations of neuronal ensembles, as well as modulating gain control [30]. Found in cortical layers 2–6, layer 4 PV-BCs are mainly mapped to Pvalb Reln [9], exhibiting local axonal arborizations. However, PV-BCs in supra- and infragranular layers possess more extensive axonal projections which extend into neighboring columns and layers (Fig. 2c) [33]. ChCs (also called axo-axonic cells) are characterized by axonal arbors that resemble the candlesticks of a chandelier and specialize in targeting the axon initial segment of pyramidal cells, imposing an even stronger output control on them than PV-BCs. ChCs belong to the Pvalb Vipr2 t-type [9, 26].
The SST subclass encompasses diverse cell types, including Martinotti cells (MCs) and non Martinotti cells (NMCs). MCs are considered to primarily target distal apical dendrites of pyramidal cells through an extensive axonal arbor in layer 1, playing a vital role in feedback inhibition [21]. However, they also possess extensive axonal arborization outside of L1, the targeting of which is still unknown. Besides SST-Calb2 MCs in layer 2/3, 2 types of MCs exist in layer 5. While L5 T-shaped MCs primarily innervate layer 1 apical tufts, exhibiting low-threshold spiking, L5 fanning-out MCs innervate layer 2/3 and the lower half of layer 1 with adapting firing patterns [22]. They are defined by Myh8 and Etv1 expressions [9, 11, 35], respectively. Quasi-fast-spiking L4/L5A SST-Hpse [20, 26, 35] NMCs, with axon extensively targeting L4, relay information from the lemniscal thalamus [22]. The L5/6 NMCs possess, next to local axon collaterals, an ascending projection mostly targeting L4. They are associated with SST-Crhr2 and SST-C1ql3 t-types [9, 26].
The majority of VIP interneurons preferentially target other GABAergic subclasses, notably SST cells, consequently disinhibiting local pyramidal neurons [15]. They receive inputs from higher-order cortical areas and neuromodulatory inputs from subcortical regions [21]. Bipolar/bitufted VIP interneurons feature vertically oriented dendrites that extend into layer 1. Layer 2/3 VIP cells possess axonal arborizations that extend downward to the layer 6/white matter boundary [22], with Vip-Rspo1-Itga4 displaying regular spiking and Vip-Ptprt-Pkp2 (mostly associated with Chat+ cells) displaying irregular spiking [9, 10]. In contrast, Vip-Lmo1-Fam159b cells residing in layers 5 and 6 exhibit a more limited pattern of axonal arborizations, mainly confined to these layers [9]. Small basket cells, another (assumed) subtype of VIP interneuron, exhibit a predominantly localized axonal arbor and are characterized by the expression of CCK. These neurons possess small soma and are primarily distributed in layer 2/3. Unlike other VIP interneuron subtypes, they establish perisomatic basket terminals akin to PV-BCs [33]. There is some debate whether the Sncg subclass (at least partially) corresponds to VIP/CCK cells [32] or whether they form separate subclasses. They display regular, adapting firing patterns.
Neurogliaform cells (NGCs) with multipolar dendritic arbors and the notable feature of a highly dense perisomatic axonal arborization correspond to several Lamp5 types. Found in every layer, being especially prominent in layer 1, they are (partially) known for their late-spiking behavior, utilizing volume transmission with many thousands of boutons in their axonal arbor. Their activation of postsynaptic metabotropic GABAB receptors potentially serves as a source of slow inhibition [30]. Additionally, they exhibit gap junctional coupling with other inhibitory neuron types [31]. Intriguingly, the Lamp5 Lhx6 type (associated with deep L5/L6 NGCs) is believed to originate from the MGE, distinguishing it from all other Lamp5 types [32]. Layer 1 harbors two other subtypes: Alpha7 cells and canopy cells. Both of these subtypes display a non-late-spiking adapting firing pattern. Alpha7 cells are aligned with Lamp5-Chrna7 markers and are characterized by a prominent hyperpolarizing sag, along with axon collaterals that extend into L5A. Conversely, canopy cells, identified as Lamp5-Fam19a1-Tmem182 types, feature horizontally elongated axonal arbors primarily confined to the upper half of L1 [3, 27].
The question of whether neuronal classification can be based on underlying biological mechanisms or remains phenomenological has been a longstanding debate. However, recent evidence has demonstrated that distinct subtypes perform specific circuit functions, implying that these subtypes indeed exist rather than being a numbers game. For example, SST-Calb2 MCs selectively innervate L5-PT, while SST-Nmbr NMCs predominantly target L5-IT neurons [35]. During whisking, the activity of L2/3 MCs decreased. Conversely, L4-6 NMCs became more active [18]. Another interesting case was the subtypes in V1, of which the modulation state was observed to be associated with their positions along the main axis of transcriptomic variation (transcriptomic principal component 1). Subtypes at the negative end of the axis were most prominently active during synchronized states, while those that occupied the most positive end were most active during desynchronized and running states [3]. The notable variations of modulation state among subtypes seem to be another indication of continuous transcriptomic variation rather than discrete subtypes. The smoothly varying neuronal activity along the transcriptomic continuum offers an alternative viewpoint on the knotty situation of continuity within/across t-types. We believe there is an urgent need for more research to establish connections between cell type classification and the functional characteristics of their cortical area-specific circuits.
Conclusions and outlook
The journey from intuitive morphological criteria to cutting-edge transcriptomic profiling to multimodal classification represents a remarkable evolution in our quest to unravel the intricacies of the brain’s cellular landscape [12]. This transformation has not only expanded our understanding of neuronal diversity but also paved the way for more precise and comprehensive cell classifications.
One of the key strategies employed in this pursuit is the integration of multiple criteria for classification, a shift from flat to hierarchical systems, which inherently incorporates relationships between types into the classification and offers a versatile approach for adapting the system in response to new information [41]. Nevertheless, challenges persist in addressing the overlay of discrete cell type distinctions with graded properties. Though neurons feature discrete, non-overlapping branches at the family level, they have the capacity to create continuous transcriptomic and morpho-electrical landscapes within these families [2, 9, 26, 38, 42]. Finding a ‘sweet spot’ between splitter and lumper perspectives requires theoretical breakthrough for a principled framework on conceptualization of cell variability, which should to the largest extent possible reflect the functional consequences associated with different cell rather than reducing the classification process to a subjective exercise. Flexibility is paramount; the framework must adapt to the dynamic nature of individual cell entities, recognizing that these entities are not static but evolving components of intricate biological systems. In essence, the emphasis should not solely be on the sheer quantity of identified cell types but on the meaningful understanding of how these cell types carry relevance in terms of their specific contributions to information processing and network dynamics. While acknowledging the significance of the transcriptomic signature, it should primarily function as the bedrock for unveiling specific functional properties and establishing a causal link between gene expression and cellular function. Caution is urged against overprioritizing transcriptomics for classification purposes. Instead, this perspective encourages a more purpose-driven methodology in single-cell transcriptomics. It advocates for a thoughtful and context-driven approach, which ensures that the classification process is not a detached enumeration but remains intimately connected to the understanding of how molecular characteristics manifest in observable and functionally relevant features of neurons.
Furthermore, the road ahead involves the development of a unified nomenclature for neural cell types and fostering collaboration among research groups and analysis platforms. Initiatives like the Brain Initiative Cell Census Consortium (BICCC) [2, 17] embody these goals, aiming to generate a census of cell types and facilitate cross-areal and cross-species comparisons. These collective efforts are designed to provide open-access resources and tools that can benefit the broader research community.
The classification will aid in precise cell targeting and unlock invaluable insights into the developmental mechanisms driving neuronal identity and diversity also in the cerebral cortex, a task that was already successfully achieved in areas like the hypothalamus [8] or brainstem/spinal cord [4]. With the help of different recombinase-specific driver lines that replicate the activity of crucial transcription factors involved in specification and differentiation [11], fate-mapping of biologically important subtypes of neurons will enable the exploration of spatiotemporal determinants shaping their developmental trajectories. This could potentially be the linchpin for programming and reprogramming cortical neurons, offering innovative therapeutic approaches for diseases that predominantly affect specific neuronal populations.
Future efforts must explore additional dimensions, including local connectivity, multi-omics, in vivo functional characterization, brain states, network dynamics on shorter and longer time scales, and role of neuromodulators. Breakthrough techniques like automated morphological reconstruction and bona-fide measurements of dynamic changes, instead of snapshots of a cell, will also open new frontiers in research. This journey into the multidimensional realm holds the potential to usher in a new era of data-driven cell classification and a deeper understanding of the brain.
Data availability
Not applicable
References
Baker A, Kalmbach B, Morishima M, Kim J, Juavinett A, Li N, Dembrow N (2018) Specialized subpopulations of deep-layer pyramidal neurons in the neocortex: bridging cellular properties to functional consequences. J Neurosci 38:5441–5455. https://doi.org/10.1523/JNEUROSCI.0150-18.2018
BRAIN Initiative Cell Census Network (BICCN) (2021) A multimodal cell census and atlas of the mammalian primary motor cortex. Nature 598:86–102. https://doi.org/10.1038/s41586-021-03950-0
Bugeon S, Duffield J, Dipoppa M, Ritoux A, Prankerd I, Nicoloutsopoulos D, Orme D, Shinn M, Peng H, Forrest H, Viduolyte A, Reddy CB, Isogai Y, Carandini M, Harris KD (2022) A transcriptomic axis predicts state modulation of cortical interneurons. Nature 607:330–338. https://doi.org/10.1038/s41586-022-04915-7
Capelli P, Pivetta C, Soledad Esposito M, Arber S (2017) Locomotor speed control circuits in the caudal brainstem. Nature 551:373–377. https://doi.org/10.1038/nature24064
DeFelipe J, López-Cruz PL, Benavides-Piccione R, Bielza C, Larrañaga P, Anderson S, Burkhalter A, Cauli B, Fairén A, Feldmeyer D, Fishell G, Fitzpatrick D, Freund TF, González-Burgos G, Hestrin S, Hill S, Hof PR, Huang J, Jones EG et al (2013) New insights into the classification and nomenclature of cortical GABAergic interneurons. Nat Rev Neurosci 14:202–216. https://doi.org/10.1038/nrn3444
Economo MN, Viswanathan S, Tasic B, Bas E, Winnubst J, Menon V, Graybuck LT, Nguyen TN, Smith KA, Yao Z, Wang L, Gerfen CR, Chandrashekar J, Zeng H, Looger LL, Svoboda K (2018) Distinct descending motor cortex pathways and their roles in movement. Nature 563(7729):79–84. https://doi.org/10.1038/s41586-018-0642-9
Feldmeyer D, Qi G, Emmenegger V (2018) Staiger JF (2018). Inhibitory interneurons and their circuit motifs in the many layers of the barrel cortex. Neuroscience 368:132–151. https://doi.org/10.1016/j.neuroscience.2017.05.027
Fong H, Zheng J, Kurrasch D (2023) The structural and functional complexity of the integrative hypothalamus. Science 382:388–394. https://doi.org/10.1126/science.adh8488
Gouwens NW, Sorensen SA, Baftizadeh F, Budzillo A, Lee BR, Jarsky T, Alfiler L, Baker K, Barkan E, Berry K, Bertagnolli D, Bickley K, Bomben J, Braun T, Brouner K, Casper T, Crichton K, Daigle TL, Dalley R et al (2020) Integrated morphoelectric and transcriptomic classification of cortical GABAergic cells. Cell 183:935–953.e19. https://doi.org/10.1016/j.cell.2020.09.057
Gouwens NW, Sorensen SA, Berg J, Lee C, Jarsky T, Ting J, Sunkin SM, Feng D, Anastassiou CA, Barkan E, Bickley K, Blesie N, Braun T, Brouner K, Budzillo A, Caldejon S, Casper T, Castelli D, Chong P et al (2019) Classification of electrophysiological and morphological neuron types in the mouse visual cortex. Nat Neurosci 22:1182–1195. https://doi.org/10.1038/s41593-019-0417-0
Harris KD, Shepherd GM (2015) The neocortical circuit: themes and variations. Nat Neurosci 18:170–181. https://doi.org/10.1038/nn.3917
Hodge RD, Bakken TE, Miller JA, Smith KA, Barkan ER, Graybuck LT, Close JL, Long B, Johansen N, Penn O, Yao Z, Eggermont J, Höllt T, Levi BP, Shehata SI, Aevermann B, Beller A, Bertagnolli D, Brouner K et al (2019) Conserved cell types with divergent features in human versus mouse cortex. Nature 573:61–68. https://doi.org/10.1038/s41586-019-1506-7
Huang ZJ (2014) Toward a genetic dissection of cortical circuits in the mouse. Neuron 83:1284–1302. https://doi.org/10.1016/j.neuron.2014.08.041
Kim EJ, Zhang Z, Huang L, Ito-Cole T, Jacobs MW, Juavinett AL, Senturk G, Hu M, Ku M, Ecker JR, Callaway EM (2020) Extraction of distinct neuronal cell types from within a genetically continuous population. Neuron 107:274–282.e6. https://doi.org/10.1016/j.neuron.2020.04.018
Kullander K, Topolnik L (2021) Cortical disinhibitory circuits: cell types, connectivity and function. Trends Neurosci 44:643–657. https://doi.org/10.1016/j.tins.2021.04.009
Matho KS, Huilgol D, Galbavy W, He M, Kim G, An X, Lu J, Wu P, Di Bella DJ, Shetty AS, Palaniswamy R, Hatfield J, Raudales R, Narasimhan A, Gamache E, Levine JM, Tucciarone J, Szelenyi E, Harris JA et al (2021) Genetic dissection of the glutamatergic neuron system in cerebral cortex. Nature 598:182–187. https://doi.org/10.1038/s41586-021-03955-9
Miller JA, Gouwens NW, Tasic B, Collman F, van Velthoven CT, Bakken TE, Hawrylycz MJ, Zeng H, Lein ES, Bernard A (2020) Common cell type nomenclature for the mammalian brain. Elife 9:e59928. https://doi.org/10.7554/eLife.59928
Muñoz W, Tremblay R, Levenstein D, Rudy B (2017) Layer-specific modulation of neocortical dendritic inhibition during active wakefulness. Science 355:954–959. https://doi.org/10.1126/science.aag2599 PMID: 28254942
Muñoz-Castañeda R, Zingg B, Matho KS, Chen X, Wang Q, Foster NN, Li A, Narasimhan A, Hirokawa KE, Huo B, Bannerjee S, Korobkova L, Park CS, Park YG, Bienkowski MS, Chon U, Wheeler DW, Li X, Wang Y et al (2021) Cellular anatomy of the mouse primary motor cortex. Nature 598:159–166. https://doi.org/10.1038/s41586-021-03970-w
Naka A, Veit J, Shababo B, Chance RK, Risso D, Stafford D, Snyder B, Egladyous A, Chu D, Sridharan S, Mossing DP, Paninski L, Ngai J, Adesnik H (2019) Complementary networks of cortical somatostatin interneurons enforce layer specific control. Elife 8:e43696. https://doi.org/10.7554/eLife.43696
Naskar S, Qi J, Pereira F, Gerfen CR, Lee S (2021) Cell-type-specific recruitment of GABAergic interneurons in the primary somatosensory cortex by long-range inputs. Cell Rep 34:108774. https://doi.org/10.1016/j.celrep.2021.108774
Nigro MJ, Hashikawa-Yamasaki Y, Rudy B (2017) Diversity and connectivity of layer 5 somatostatin-expressing interneurons in the mouse barrel cortex. J Neurosci 38:1622–1633. https://doi.org/10.1523/JNEUROSCI.2415-17.2017
Patiño M, Lagos WN, Patne NS, Tasic B, Zeng H, Callaway EM (2022) Single-cell transcriptomic classification of rabies-infected cortical neurons. Proc Natl Acad Sci USA 119:e2203677119. https://doi.org/10.1073/pnas.2203677119
Peng H, Xie P, Liu L, Kuang X, Wang Y, Qu L, Gong H, Jiang S, Li A, Ruan Z, Ding L, Yao Z, Chen C, Chen M, Daigle TL, Dalley R, Ding Z, Duan Y, Feiner A et al (2021) Morphological diversity of single neurons in molecularly defined cell types. Nature 598:174–181. https://doi.org/10.1038/s41586-021-03941-1
Petilla Interneuron Nomenclature Group (2008) Petilla terminology: nomenclature of features of GABAergic interneurons of the cerebral cortex. Nat Rev Neurosci 9:557–568. https://doi.org/10.1038/nrn2402
Scala F, Kobak D, Bernabucci M, Bernaerts Y, Cadwell CR, Castro JR, Hartmanis L, Jiang X, Laturnus S, Miranda E, Mulherkar S, Tan ZH, Yao Z, Zeng H, Sandberg R, Berens P, Tolias AS (2021) Phenotypic variation of transcriptomic cell types in mouse motor cortex. Nature 598:144–150. https://doi.org/10.1038/s41586-020-2907-3
Schuman B, Machold RP, Hashikawa Y, Fuzik J, Fishell GJ, Rudy B (2019) Four unique interneuron populations reside in neocortical layer 1. J Neurosci 39:125–139. https://doi.org/10.1523/JNEUROSCI.1613-18.2018
Siletti K, Hodge R, Mossi Albiach A, Lee KW, Ding SL, Hu L, Lönnerberg P, Bakken T, Casper T, Clark M, Dee N, Gloe J, Hirschstein D, Shapovalova NV, Keene CD, Nyhus J, Tung H, Yanny AM, Arenas E et al (2023) Transcriptomic diversity of cell types across the adult human brain. Science 382(6667):eadd7046. https://doi.org/10.1126/science.add7046
Staiger JF, Bojak I, Miceli S, Schubert D (2015) A gradual depth-dependent change in connectivity features of supragranular pyramidal cells in rat barrel cortex. Brain Struct Funct 220:1317–1337. https://doi.org/10.1007/s00429-014-0726-8
Staiger JF, Petersen CCH (2021) Neuronal circuits in barrel cortex for whisker sensory perception. Physiol Rev 101:353–415. https://doi.org/10.1152/physrev.00019.2019
Tasic B, Menon V, Nguyen TN, Kim TK, Jarsky T, Yao Z, Levi B, Gray LT, Sorensen SA, Dolbeare T, Bertagnolli D, Goldy J, Shapovalova N, Parry S, Lee C, Smith K, Bernard A, Madisen L, Sunkin SM et al (2016) Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat Neurosci 19:335–346. https://doi.org/10.1038/nn.4216
Tasic B, Yao Z, Graybuck LT, Smith KA, Nguyen TN, Bertagnolli D, Goldy J, Garren E, Economo MN, Viswanathan S, Penn O, Bakken T, Menon V, Miller J, Fong O, Hirokawa KE, Lathia K, Rimorin C, Tieu M et al (2018) Shared and distinct transcriptomic cell types across neocortical areas. Nature 563:72–78. https://doi.org/10.1038/s41586-018-0654-5
Tremblay R, Lee S, Rudy B (2016) GABAergic Interneurons in the neocortex: from cellular properties to circuits. Neuron 91:260–292. https://doi.org/10.1016/j.neuron.2016.06.033
Weiler S, Guggiana Nilo D, Bonhoeffer T, Hübener M, Rose T, Scheuss V (2023) Functional and structural features of L2/3 pyramidal cells continuously covary with pial depth in mouse visual cortex. Cereb Cortex 33:3715–3733. https://doi.org/10.1093/cercor/bhac303
Wu SJ, Sevier E, Dwivedi D, Saldi GA, Hairston A, Yu S, Abbott L, Choi DH, Sherer M, Qiu Y, Shinde A, Lenahan M, Rizzo D, Xu Q, Barrera I, Kumar V, Marrero G, Prönneke A, Huang S et al (2023) Cortical somatostatin interneuron subtypes form cell-type-specific circuits. Neuron 111:2675–2692.e9. https://doi.org/10.1016/j.neuron.2023.05.032
Yao S, Wang Q, Hirokawa KE, Ouellette B, Ahmed R, Bomben J, Brouner K, Casal L, Caldejon S, Cho A, Dotson NI, Daigle TL, Egdorf T, Enstrom R, Gary A, Gelfand E, Gorham M, Griffin F, Gu H et al (2023) A whole-brain monosynaptic input connectome to neuron classes in mouse visual cortex. Nat Neurosci 26:350–364. https://doi.org/10.1038/s41593-022-01219-x
Yao Z, Liu H, Xie F, Fischer S, Adkins RS, Aldridge AI, Ament SA, Bartlett A, Behrens MM, Van den Berge K, Bertagnolli D, de Bézieux HR, Biancalani T, Booeshaghi AS, Bravo HC, Casper T, Colantuoni C, Crabtree J, Creasy H et al (2021a) A transcriptomic and epigenomic cell atlas of the mouse primary motor cortex. Nature 598:103–110. https://doi.org/10.1038/s41586-021-03500-8
Yao Z, van Velthoven CTJ, Nguyen TN, Goldy J, Sedeno-Cortes AE, Baftizadeh F, Bertagnolli D, Casper T, Chiang M, Crichton K, Ding SL, Fong O, Garren E, Glandon A, Gouwens NW, Gray J, Graybuck LT, Hawrylycz MJ, Hirschstein D et al (2021b) A taxonomy of transcriptomic cell types across the isocortex and hippocampal formation. Cell 184:3222–3241.e26. https://doi.org/10.1016/j.cell.2021.04.021
Yuste R, Hawrylycz M, Aalling N, Aguilar-Valles A, Arendt D, Armañanzas R, Ascoli GA, Bielza C, Bokharaie V, Bergmann TB, Bystron I, Capogna M, Chang Y, Clemens A, de Kock CPJ, DeFelipe J, Dos Santos SE, Dunville K, Feldmeyer D et al (2020) A community-based transcriptomics classification and nomenclature of neocortical cell types. Nat Neurosci 23:1456–1468. https://doi.org/10.1038/s41593-020-0685-8
Zeng H (2022) What is a cell type and how to define it? Cell 185:2739–2755. https://doi.org/10.1016/j.cell.2022.06.031
Zeng H, Sanes JR (2017) Neuronal cell-type classification: challenges, opportunities and the path forward. Nat Rev Neurosci 18:530–546. https://doi.org/10.1038/nrn.2017.85
Zhang M, Eichhorn SW, Zingg B, Yao Z, Cotter K, Zeng H, Dong H, Zhuang X (2021) Spatially resolved cell atlas of the mouse primary motor cortex by MERFISH. Nature 598:137–143. https://doi.org/10.1038/s41586-021-03705-x
Acknowledgements
We would like to thank all colleagues worldwide for their great insight though all the many publications, many of which could not be implemented here due to space restrictions and specific focus. Further thanks are due to Hongkui Zeng and Xiaowei Zhuang for the support with the figures. Felix Preuss and Pavel Truschow are also acknowledged for their contributions to the figures.
Funding
Open Access funding enabled and organized by Projekt DEAL. Work of JFS pertinent to this review is supported by the Deutsche Forschungsgemeinschaft (STA 431/14-1; 17-1; 21-1).
Author information
Authors and Affiliations
Contributions
XM and JFS have both conceived, written, and illustrated the review together.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mao, X., Staiger, J.F. Multimodal cortical neuronal cell type classification. Pflugers Arch - Eur J Physiol 476, 721–733 (2024). https://doi.org/10.1007/s00424-024-02923-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00424-024-02923-2