Abstract
Human MYCN is an oncogene amplified in neuroblastoma and many other tumors. Both human MYCN and mouse Mycn genes are important in embryonic brain development, but their functions in adult healthy nerve system are completely unknown. Here, with Mycn-eGFP mice and quantitative RT-PCR, we found that Mycn was expressed in specific brain regions of young adult mice, including subventricular zone (SVZ), subgranular zone (SGZ), olfactory bulb (OB), subcallosal zone (SCZ), and corpus callosum (CC). With immunohistochemistry (IHC), we found that many Mycn-expressing cells expressed neuroblast marker doublecortin (DCX) and proliferation marker Ki67. With Dcx-creER and Mki67-creER mouse lines, we fate mapped Dcx-expressing neuroblasts and Mki67-expressing proliferation cells, along with deleting Mycn from these cells in adult mice. We found that knocking out Mycn from adult neuroblasts or proliferating cells significantly reduced cells in proliferation in SVZ, SGZ, OB, SCZ, and CC. We also demonstrated that the Mycn-deficient neuroblasts in SGZ matured quicker than wild-type neuroblasts, and that Mycn-deficient proliferating cells were more likely to survive in SVZ, SGZ, OB, SCZ, and CC compared to wild type. Thus, our results demonstrate that, in addition to causing tumors in the nervous system, oncogene Mycn has a crucial function in neurogenesis and oligodendrogenesis in adult healthy brain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MYCN, the human gene encoding N-Myc, was first identified as an oncogene amplified in human neuroblastoma [1], a tumor characterized as having undifferentiated neuroblasts [2]. The expression level of MYCN correlates with the prognosis of neuroblastoma [3], and overexpression of human MYCN gene produces neuroblastoma in mice [4]. MYCN overexpression is also observed in other nervous system tumors such as medulloblastoma, glioblastoma, retinoblastoma, and spinal ependymoma, as well as tumors outside of the nervous system such as neuroendocrine prostate cancer, nephroblastoma (Wilms’ tumor), and many others [5,6,7]. The nucleotide sequence of MYCN is very similar to that of oncogene MYC [8].
In addition to causing tumors, MYCN is also expressed in human fetal brain [9], and heterozygous MYCN mutation causes Feingold syndrome characterized as reduced brain size and learning disability [10]. Similarly, mouse Mycn gene is expressed in embryonic and neonatal forebrains and hindbrains [11], and in embryonic eye, spinal cord, dorsal root ganglion (DRG) (spinal ganglia) and trigeminal ganglia (cranial ganglion) [12]. A homozygous Mycn-deficient mouse dies between 10.5 and 12.5 days of gestation, with significant developmental abnormalities in central and peripheral nervous systems [13]. Conditional deletion of Mycn from Nestin-expressing neuronal progenitor cells (NPCs) in embryo results in dramatic reduction of NPC proliferation and disruption of neuronal differentiation, leading to a substantial decrease in brain mass and disorganization of both the cortex and cerebellum [14]. Mycn also plays an important role in the proliferation and survival of embryonic sympathetic neuroblasts [15]. Finally, Mycn is highly functional in pluripotent cells as well, because overexpression of Mycn, along with Oct4, Sox2, and Klf4, transforms mouse embryonic fibroblasts into induced pluripotent stem cells [16].
As Mycn was initially found to be not expressed in adult brain [11], the function of Mycn in the nervous system of an healthy adult has never been revealed. With Mycn-EGFP reporter mice [17], we found that Mycn is expressed in restricted regions in adult brain, such as subventricular zone (SVZ), subgranular zone (SGZ) of hippocampal dentate gyrus (DG), the two major sites for adult neurogenesis [18], and corpus callosum (CC) and subcallosal zone (SCZ), a site of adult oligodendrogenesis [19]. Our results further demonstrate that Mycn plays important roles in adult neurogenesis and oligodendrogenesis.
Materials and Method
Animals
All animal procedures were performed according to protocols approved by Institutional Animal Care and Use of Laboratory animals. Animals were housed in groups of five in a pathogen-free barrier facility. Young adult (8–12-week-old) male and female mice were used in all experiments. Tamoxifen or hydroxytamoxifen was injected in doses of 100 mg per kg intraperitoneally.
Mycn-EGFP mouse, STOCK Tg(Mycn-EGFP)ET250Gsat/Mmucd, RRID:MMRRC_010553-UCD, was obtained from the Mutant Mouse Resource and Research Center (MMRRC) at University of California at Davis, an NIH-funded strain repository, and was donated to the MMRRC by Nathaniel Heintz, Ph.D. at Rockefeller University to GENSAT.
Mycn fl/fl mice were obtained from Jackson Laboratory (B6,129-Mycn tm1Psk/J, stock No. 006933); Mki67-creER (Mki67 tm2.1(cre/ERT2)cle/J, stock No. 029803). Dcx-creER was a gift from professor Zhiqi Xiong. Rosa26-TdTomato-loxp-STOP-loxp mice were obtained from Jackson Laboratory (B6. Cg-Gt (ROSA)26Sor tm14(CAG−Td−Tomato)Hze/J, stock No. 007914).
Immunohistochemistry
Animals were sacrificed and then intracardially perfused with ice-cold PBS and 4% paraformaldehyde (PFA; Thermo Scientific, AC416785000). Brains were dissected, post-fixed in 4% paraformaldehyde overnight, and embedded in OCT compound (Tissue-Tek, Thermo Fisher, 4585). Coronal brain sections (20 µm thick) were collected by cryostat. The sections were washed with 1 × PBS and incubated with blocking buffer with 10% fetal bovine serum (FBS) (Thermo Fisher) and 0.3% Triton X-100 in PBS for 30 min at room temperature. Primary antibodies diluted in 1% FBS and 0.3% Triton X-100 in PBS were added to the slices and incubated overnight at 4 °C. Subsequently, conjugated secondary antibodies of Alexa Fluor 488, Alexa Fluor 555, or Alexa Fluor 647 (1:1000; Invitrogen) were added, and DAPI (1:1000; Invitrogen) was used for showing nucleuses.
The following primary antibodies and their dilutions were used: GFAP (rabbit, 1:1000; Dako, #Z0334), Ki67 (rabbit, 1:1000; Abcam), doublecortin (DCX) (rabbit, 1:500; Abcam), NeuN (mouse, 1:500; Millipore), oligodendrocyte transcription factor 2 (Olig2) (rabbit, 1:500; Millipore, #AB9610), and Tbr2 (rabbit, 1:500; Abcam). Images were captured using a Zeiss LSM700 confocal microscope as a single plain and were processed with Fiji/ImageJ (NIH).
Cell Counting
For EdU + cell number counting, we chose three regions to count the EdU + cell number, including SVZ, hippocampus, and olfactory bulb. For each region, we cut the brain from the same region and collected 20-µm cryosections of each brain region from three animals per group. In order to get the information in the whole region, we mounted, immunostained, and counted the EdU + cells in every sixth section of the brain region. To quantify the Td-tomato + cells in the targeted regions, we counted Td-tomato + cells from 3 to 4 animals per group in slices with the same region. We use the number of cells per field to measure the density of cells within SVZ, CC, SGZ, SCZ, and olfactory bulb (OB), with the field defined as the specific brain regions under 20 × magnification, and count the Td-Tomato + cell number in that specific region. To analyze the distribution of the newborn cells in dentate gyrus, we divided the granule cell layer into three layers equally, as shown in Fig. 7b, counted the Td-tomato–positive cells in each layer, and calculated the percentage of the cell number in each layer versus the whole Td-tomato + cell number in the dentate gyrus. To quantify the Td-tomato–positive cells that co-express Olig2, we counted Td-tomato–positive cells in three sections in each area per animal and quantified the double-labeled Td-tomato/Olig2–immunoreactive cells.
EdU Staining
EdU solution (10 mg/kg; Invitrogen, C10337) was injected intraperitoneally 24 h before tissue collection. Click-iT reaction cocktail (1 × Click-iT reaction buffer, CuSO4, Alexa Fluor azide, and 1 × reaction buffer additive) was prepared and applied to slides for 30 min and washed with PBS for 10 min three times, followed by first antibody incubation overnight. Then, the slides were incubated with second antibody for 1 h at room temperature and washed with PBS for 10 min, followed by DAPI staining.
Flow Cytometry
The brain region such as SVZ was freshly microdissected and homogenized. The homogenized tissue was filtered then mixed with Percol (the final concentration of Percol is 30%). After being centrifuged at 800 g for 20 min, the cells were harvested for the subsequent flow cytometry. Cells were first incubated with anti-mouse CD16/32 in PBS for 5 min to block the Fc receptor. Cells were then washed with PBS and incubated with near-IR (1:1000; BioLegend) for 30 min. Cells were washed with PBS and resuspended in the desired antibody mix in PBS and stained for 30 min at 4 °C. After washing with PBS, cells were fixed and permeabilized using the Foxp3 transcription factor staining buffer kit (Thermo Fisher) for intracellular and intranuclear staining and washed two times with Perm/Wash (0.01% sodium azide, 0.5% saponin, and 2% BSA in PBS). Cells were then resuspended in PBS and analyzed. Flow cytometry was performed using a FACS Symphony A5 (BD Biosciences) and analyzed with the FlowJo software. Cell sorting was performed using a FACSAria III. The following fluorochrome-conjugated monoclonal antibodies were used: Ki67 (clone SOLA15, 1:5000) and DCX (anti-rabbit, 1:100; Abcam). Dead cells and duplets were excluded for analysis using SSC-A/H, FSC-A/H, and a fixable viability kit (near-IR staining; BioLegend).
qRT-PCR
For qRT-PCR, cDNA synthesis was performed using a SuperScript III First-Strand Synthesis System (Invitrogen, 18,080–051). Quantitative PCR was performed using PerfeCTa SYBR Green FastMix Rox (Quanta Biosciences) in a qPCR system (Bio-Rad). β-Actin was used as the housekeeping gene for normalization. The following primers were used: Actb (β-actin), CCACACCCGCCACCAGTTCG/TACAGCCCGGGGAGCATCGT; Mycn, AGCGTTCAACTAGCAGACCAT/CGAAAGGGCAGATTGTGTGG; and Dcx, CGACCAAGACGCAAATGGAAC/CAATGACAGCGGCAGGTACA.
Experimental Design and Statistical Analysis
The experiments were conducted with 3–4 animals each group, the minimal number of animals to obtain results with statistical significance. For immunohistochemistry (IHC) studies, at least 3 sections from each animal were immuno-stained, and the average value of all the sections from the same animals was used to represent the value of that particular animal. For flow cytometry studies, one experiment was performed in one animal. For qRT-PCR, three duplicates were performed for each primer pair, and the average value of all three duplicates was used to represent the value of that particular animal. No data was excluded, and data are expressed as mean ± SEM. Statistical analysis was performed using GraphPad Prism, version 8.0. Unpaired two-tailed Student’s t tests were used for single comparisons between two groups. Other data were analyzed using one-way or two-way analysis of variance (ANOVA).
Results
Mycn Is Expressed in Adult Healthy Brain of Mice
To investigate if Mycn is expressed in adult brain, we studied young adult (8–12 weeks old) Mycn-eGFP reporter mice generated by GENSAT [17], in which eGFP is expressed under the control of bacterial artificial chromosome containing the promoter and the enhancers of mouse Mycn gene. We found that GFP was expressed in SVZ (Fig. 1a), SGZ (Fig. 1b), and olfactory bulb (OB) (Fig. 1c). We also found GFP ( +) cells in CC (Fig. 1a, arrowheads), as well as in SCZ (Fig. 1b, arrowheads), a thin region between the hippocampus and CC that contains neural stem cells [19]. We then dissected the SVZ and hippocampus from Mycn-eGFP reporter mice and sorted out the GFP ( +) cells from the SVZ, dentate gyrus, and SCZ with fluorescence-activated cell sorting (FACS). With qRT-PCR, we confirmed that Mycn gene was enriched in the GFP ( +) cells in SVZ (Supplementary Fig. 1a), dentate gyrus (Supplementary Fig. 1b), and SCZ (Supplementary Fig. 1c).
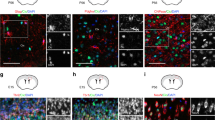
Mycn is expressed in specific sites in adult mouse brain. a–c IHC of GFP and DAPI in the subventricular zone (SVZ) and corpus callosum (CC) (a), subgranular zone (SGZ) and subcallosal zone (SCZ) (b), and olfactory bulb (OB) (c) of young adult Mycn-eGFP reporter mice. The arrowheads point to the GFP-positive cells in CC (a) and SCZ (b). d IHC of DAPI, DCX, and GFP in the SVZ of Mycn-eGFP reporter mice. The enlarged image shows the co-localization of GFP and DCX in SVZ. e–g IHC of DCX and GFP in the SGZ (e), SCZ (f), and olfactory bulb (g) of Mycn-eGFP reporter mice. The arrowheads point to the DCX and GFP double–positive cells (e, f), and the enlarged images show the GFP and DCX double–positive cells. h Quantification of d–g showing the percentage of DCX/GFP double–positive cells among GFP ( +) cells in the SVZ, SGZ, SCZ, and OB of young adult Mycn-eGFP reporter mice. i Quantification of d–g showing the percentage of DCX/GFP double–positive cells among DCX ( +) cells in the SVZ, SGZ, SCZ, and OB of young adult Mycn-eGFP reporter mice. j IHC of GFP and NeuN in the dentate gyrus of young adult Mycn-eGFP reporter mice. The arrow points to the GFP and NeuN double–positive cells, and the enlarged figure shows the NeuN and GFP double–positive cells. k Quantification of j showing the percentage of NeuN/GFP double–positive cells among GFP ( +) cells in the SGZ of Mycn-eGFP reporter mice. l IHC of GFP and Olig2 in the CC above SVZ of Mycn-eGFP reporter mice. The dashed line shows the boundary between SVZ and CC, and the arrows point to the GFP and Olig2 double–positive cells, with the enlarged figure showing the Olig2 and GFP double–positive cell. m Quantification of i showing the percentage of Olig2/GFP double–positive cells among GFP ( +) cells in the CC of young adult Mycn-eGFP reporter mice
Because Mycn is associated with neuroblastoma, we were particularly interested in finding out whether Mycn is expressed in the neuroblasts of adult healthy mice. Our IHC studies of brain sections of young adult Mycn-eGFP reporter mice with DCX, which is expressed in adult neuroblast and neuronal intermediate progenitor cells (nIPCs) [20], revealed the co-localization of DCX with GFP in SVZ (Fig. 1d), SGZ (Fig. 1e), SCZ (Fig. 1f), and OB (Fig. 1g). In fact, 72% of GFP ( +) cells in SVZ, 91% of GFP ( +) cells in SGZ, 49% of GFP ( +) cells in SCZ, and 96% of GFP ( +) cells in OB co-expressed DCX (Fig. 1h), whereas almost all DCX ( +) cells in these regions were GFP ( +) (Fig. 1i). We also found that 35% of the GFP ( +) cells in the dentate gyrus of young adult Mycn-eGFP reporter mice also expressed NeuN (Fig. 1j, k). On the other hand, 16% of GFP ( +) cells in CC co-expressed Olig2, a marker of oligodendrocyte lineage cells [21] (Fig. 1l, m). We did qRT-PCR from sorted GFP ( +) cells of young adult Mycn-eGFP reporter mice and found that GFP ( +) cells had significantly higher expression of Dcx gene than GFP ( −) cells in SVZ (Fig. 2a), dentate gyrus (Fig. 2b), and SCZ (Fig. 2c).
Mycn is enriched in Dcx-expressing cells. a–c In Mycn-eGFP reporter mice, qRT-PCR study shows that the GFP ( +) cells sorted by FACS from SVZ, dentate gyrus (DG), and SCZ had higher Dcx mRNA levels than GFP ( −) cells. Data are shown as mean ± SEM, and the analysis performed was unpaired two-tailed Student’s t test. d IHC of GFP, Td-Tomato, and DCX in SVZ and CC from a Dcx-creER/ + ; Td-Tomato/ + ; Mycn-eGFP mouse 1 day after hydroxytamoxifen treatment. In these mice, the cells expressing Dcx in the last 24 h were labeled with Td-Tomato. The dashed lines show the boundary between SVZ and CC. Virtually, all the Td-Tomato ( +) cells in SVZ and CC were GFP ( +). Most Td-Tomato ( +) cells in SVZ and some Td-Tomato ( +) cells in CC were also DCX ( +). Enlarged figures show the GFP/Td-Tomato double–positive cells and the GFP/DCX double–positive cells. Scale bar = 100 µm. e Quantification of d showing the percentage of DCX/Td-Tomato double–positive cells among Td-Tomato ( +) cells in the SVZ and CC of a Dcx-creER/ + ; Td-Tomato/ + ; Mycn-eGFP mouse 1 day after hydroxytamoxifen treatment. Data are shown as mean ± SEM, and the analysis performed was unpaired two-tailed Student’s t test
To further investigate Mycn expression in adult neuroblasts, we crossed a Mycn-eGFP mouse with a Dcx-creER mouse [22] and with a Td-Tomato reporter mouse [23]. In the young adult Dcx-creER/ + ; TdTomato/ + ; Mycn-eGFP/ + mice 1 day after hydroxytamoxifen treatment, the cells that expressed Dcx in the past 1 day were fate labeled by TdTomato (Fig. 2d). In these mice, we found both GFP ( +) and TdTomato ( +) cells in SVZ and CC, and virtually all TdTomato ( +) cells were GFP ( +) (Fig. 2d), suggesting that Mycn was expressed in cells with either current Dcx expression or recent history of Dcx expression in this region. Further IHC with DCX antibody showed that 67% of TdTomato ( +) cells in SVZ and 22% of TdTomato ( +) cells in CC were DCX ( +) and thus continued to express DCX (Fig. 2d, e).
Our IHC studies with young adult Mycn-eGFP reporter mice also showed that 6% of the GFP ( +) cells in SVZ, 17% of GFP ( +) cells in SGZ, and 56% of GFP ( +) cells in SCZ also expressed another neuroblast marker (Tbr2), encoded by gene Eomes [20] (Supplementary Fig. 1d–g). Moreover, our analysis of the published single-cell RNA-Seq (scRNA-Seq) dataset from perinatal, juvenile, and adult dentate gyrus [20] shows that Mycn is expressed in radial glia-like (RGL) cells, nIPCs, neuroblasts, and immature granule cells (GCs), as well as in some mature GCs (Supplementary Fig. 2a), consistent with our IHC study that some GFP ( +) cells in the dentate gyrus of Mycn-eGFP mice expressed the mature neuron marker NeuN (Fig. 1j, k). Interestingly, the expression pattern of Mycn is very similar to that of Dcx in dentate gyrus, which is also expressed in RGLs, nIPCs, neuroblasts, immature GCs, and some mature GCs (Supplementary Fig. 2b). In contrast, the neuroblast marker Eomes is expressed more specifically in nIPCs and neuroblasts (Supplementary Fig. 2c). On the other hand, our analysis of published scRNA-Seq dataset from oligodendrocyte lineage cells of juvenile and adult CNS [24] shows that Mycn is expressed in some oligodendrocyte precursor cells (OPCs), differentiation-committed oligodendrocyte precursors (COPs), and certain myelin-forming oligodendrocytes (MFOLs) and mature oligodendrocytes (MOLs) (Supplementary Fig. 3a). In comparison, Olig2 gene is expressed in every oligodendrocyte lineage cells (Supplementary Fig. 3b).
Mycn-Expressing Cells Can Proliferate
As one major character of adult neurogenesis and oligodendrogenesis is the proliferation of RGLs, nIPCs, neuroblasts, and OPCs [20, 21], we investigated if proliferation occurred in Mycn-expressing cells. Our IHC studies of brain sections from young adult Mycn-eGFP reporter mice with the proliferation marker Ki67 [25] showed that 4% of the GFP ( +) cells in SVZ, 28% of the GFP ( +) cells in CC, 3% of the GFP ( +) cells in SGZ, 13% of the GFP ( +) cells in SCZ, and 8% of the GFP ( +) cells in OB were Ki67 ( +) (Fig. 3a–e). To further confirm that Mycn-expressing cells do proliferate, we crossed the Mycn-eGFP mouse with the Mki67-creER mouse along with the Td-Tomato reporter mouse. In young adult Mki67-creER/ + ; TdTomato/ + ; Mycn-eGFP/ + mice, tamoxifen treatment for 5 days fate labeled the proliferating cells and the cells recently proliferated within last 5 days with Td-Tomato. We found that in this animal, 33% of GFP ( +) cells in SVZ, 38% of GFP ( +) cells in CC, 21% of GFP ( +) cells in SGZ, 45% of GFP ( +) cells in SCZ, and 34% of GFP ( +) cells in OB were TdTomato ( +) (Fig. 3f–j). All together, these results suggest that Mycn-expressing cells in adult healthy brain proliferate in multiple brain regions.
Mycn-expressing cells can proliferate. a–d IHC of GFP and Ki67 in the SVZ and CC (a), SGZ (b), SCZ (c), and OB (d) of a young adult Mycn-eGFP mouse. The dashed line in a shows the boundary between SVZ and CC. The arrows point to the GFP and Ki67 double–positive cells, with the enlarged figures showing the Ki67 and GFP double–positive cells. e The quantitative analysis of a–d shows the percentage of Ki67 ( +) cells in GFP ( +) cells in different brain regions of a Mycn-eGFP reporter mouse. f–i IHC of GFP and Td-Tomato in a Mki67-creER/ + ; Td-Tomato/ + ; Mycn-eGFP mouse in SVZ and CC (f), SGZ (g), SCZ (h), and OB (i) after tamoxifen treatment for 5 days. In these mice, the proliferating cells and the cells proliferated in the past 5 days were labeled with Td-Tomato. The dashed line in f shows the boundary between SVZ and CC, and the dashed lines in h show the boundary of SCZ. The arrows and arrowheads show the GFP and Td-Tomato double–positive cells, and the enlarged figures show the Td-Tomato and GFP double–positive cells. j The quantitative analysis of f–i shows the percentage of GFP and Td-Tomato double–positive cells among GFP ( +) cells in the SVZ, CC, SGZ, SCZ, and OB of a Mki67-creER/ + ; Td-Tomato/ + ; Mycn-eGFP mouse after 5-day tamoxifen treatment. Scale bar = 100 µm
Mycn Is Essential for Cell Proliferation in Adult Neurogenesis and Oligodendrogenesis
To investigate if Mycn contributes to cell proliferation in adult neurogenesis, we crossed Dcx-creER line with Mycn flox line [14] to generate Dcx-creER/ + ; Mycn fl/fl mice. We treated young adult Dcx-creER/ + ; Mycn fl/fl and control + / + ; Mycn fl/fl mice with daily tamoxifen for 5 days to delete Mycn from any cells that have expressed Dcx in the past 5 days in Dcx-creER/ + ; Mycn fl/fl mice. With flow cytometry to analyze DCX ( +) cells in SVZ (Supplementary Fig. 4a, b), we found that the DCX ( +) cells with Mycn deletion had significant less proliferating Ki67 ( +) cells compared to DCX ( +) cells from control + / + ; Mycn fl/fl littermates (Fig. 4a, b). We also crossed Mki67-creER line with Mycn flox line to generate Mki67-creER/ + ; Mycn fl/fl mice, and daily tamoxifen treatment of young adult Mki67-creER/ + ; Mycn fl/fl and control + / + ; Mycn fl/fl mice for 5 days will delete Mycn from any cells that have expressed Mki67 in the past 5 days in Mki67-creER/ + ; Mycn fl/fl mice. With flow cytometry (Supplementary Fig. 4c, d), we found that in SVZ, the deletion of Mycn from Mki67-expressing cells also resulted in significantly less proliferation of the DCX ( +) cells compared to that in control + / + ; Mycn fl/fl littermates (Fig. 4c, d). Of note, the proliferation rate of the total cell populations in SVZ also decreased in Mycn cKO animals compared with that in intact animals (Supplementary Fig. 5). These results suggest that Mycn contributes to the proliferation of DCX ( +) cells in SVZ in adult.
Mycn is required for the proliferation of DCX ( +) cells in SVZ. a Representative flow cytometry scatter plots to assess the proliferation of DCX ( +) cells in the SVZ of young adult Dcx-creER + / + ; Mycn fl/fl (control) versus Dcx-creER/ + ; Mycn fl/fl (cKO) mouse. b Quantification of a shows that the deletion of Mycn from Dcx-expressing cells significantly reduced the proliferation of DCX ( +) cells. c Representative flow cytometry scatter plots to assess the proliferation of DCX ( +) cells in the SVZ of young adult Mki67-creER + / + ; mycn fl/fl (control) versus Mki67-creER/ + ; Mycn fl/fl (cKO) mouse. d Quantification of c showing that the deletion of Mycn from Mki67-expressing cells significantly reduced the proliferation of DCX ( +) cells. Data are shown as mean ± SEM, and the analysis performed was unpaired two-tailed Student’s t test
To further study the functional role of Mycn in cell proliferation in brain regions of young healthy animals, we injected EdU, a nucleoside analog of thymidine that can be incorporated into DNA during DNA synthesis for cell proliferation [26], into young adult Mki67-creER/ + ; Mycn fl/fl and control + / + ; Mycn fl/fl mice after daily tamoxifen treatment for 5 days. Consistent with our flow cytometry result (Fig. 4c, d), our IHC with EdU showed that deleting Mycn from Mki67-expressing cells significantly reduced the total number of EdU ( +)-proliferating cells in SVZ compared to that in control + / + ; Mycn fl/fl mice (Fig. 5a, b). Remarkably, although many EdU ( +) cells were observed in SGZ, SCZ, and OB in control + / + ; Mycn fl/fl mice, almost no EdU ( +) cells were observed in these regions when Mycn was deleted from Mki67-expressing cells (Fig. 5c–f). These results suggest that Mycn is required for cell proliferation in the sites of adult neurogenesis (SVZ and SGZ) and oligodendrogenesis (SCZ).
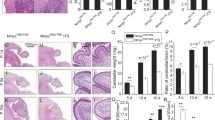
Mycn is required for cell proliferation in SVZ, SGZ, SCZ, and OB. IHC of EdU and DAPI in the SVZ (a), SGZ and SCZ (c), and OB (e) in young adult Mki67-creER + / + ; Mycn fl/fl (control) versus Mki67-creER/ + ; Mycn fl/fl (cKO) mouse. The dashed line shows the boundary between SVZ and CC (a). In c, the arrows point to the EdU-positive cells in SGZ, and the arrowheads point to the EdU-positive cells in SCZ. Scale bar = 100 µm. Quantification of a, c, and e shows that the deletion of Mycn from Mki67-expressing cells significantly reduced cell proliferation in SVZ (b), SGZ and SCZ (d), and OB (f). Data are shown as mean ± SEM, and the analysis performed was unpaired two-tailed Student’s t test
Mycn Inhibits the Maturation of Dcx-Expressing Cells in Adult Dentate Gyrus
To investigate if Mycn is involved in maturation of neuroblast in adult, we crossed Dcx-creER line with TdTomato reporter line and Mycn flox line. Treating young adult and Dcx-creER/ + ; TdTomato/ + ; Mycn fl/fl mice and control Dcx-creER/ + ; TdTomato/ + ; + / + mice with daily tamoxifen for 5 days fate labeled all the cells that have expressed Dcx in last 5 days with TdTomato, along with deletion of Mycn in TdTomato-labeled cells in Dcx-creER/ + ; TdTomato/ + ; Mycn fl/fl mice. We did IHC of dentate gyrus with DCX antibody 14 days after tamoxifen treatment and measured the percentage of TdTomato ( +) cells that are still DCX ( +). We found significantly less DCX ( +) cells in the TdTomato ( +) cells with Mycn deletion compared to the control TdTomato ( +) cells with intact Mycn (Fig. 6a, b), suggesting that TdTomato ( +) cells with Mycn deletion were more mature than TdTomato ( +) cells with intact Mycn. In addition, in the same mice, we observed that more TdTomato ( +) cells in the dentate gyrus of Mycn-deleted TdTomato ( +) cells expressed the mature neuron marker NeuN compared to that of control TdTomato ( +) cells with intact Mycn (Fig. 6c, d), further supporting that Mycn inhibits the maturation of Dcx-expressing cells.
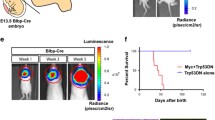
Mycn inhibits the maturation of Dcx-expressing cells in dentate gyrus. a IHC of DCX and Td-Tomato in the dentate gyrus of young adult Dcx-creER/ + ; Td-Tomato/ + ; + / + (control) versus Dcx-creER/ + ; Td-Tomato/ + ; mycn fl/fl (cKO) mice on day 14 after the 5-day tamoxifen treatment. In these mice, the cells that had expressed Dcx 14–19 days ago were fate mapped with Td-Tomato. b Quantification of a shows that the Dcx-creER/ + ; Td-Tomato/ + , mycn fl/fl (cKO) mice had significantly less DCX ( +) cells among the Td-Tomato ( +) cells compared to the control Dcx-creER/ + ; Td-Tomato/ + ; + / + mice. c IHC of NeuN and Td-Tomato in the dentate gyrus of young adult Dcx-creER/ + ; Td-Tomato/ + ; + / + (control) versus Dcx-creER/ + ; Td-Tomato/ + ; mycn fl/fl (cKO) mice on day 14 after the 5-day tamoxifen treatment. The enlarged figures show the NeuN and Td-Tomato double–positive cells. d Quantification of c shows that the Dcx-creER/ + ; Td-Tomato/ + ; mycn fl/fl (cKO) mice had significantly more NeuN ( +) cells among the Td-Tomato ( +) cells compared to the control Dcx-creER/ + ; Td-Tomato/ + ; + / + mice. Scale bar = 100 µm. Data are shown as mean ± SEM, and the analysis performed was unpaired two-tailed Student’s t test
Using the same type of animals, we also measured the distribution of TdTomato ( +) cells in the granule layer of dentate gyrus, because the newly generated neurons get more and more mature when they migrate from inner layers to outer layers [20, 27]. We found that 14 days after tamoxifen treatment of young adult and Dcx-creER/ + ; TdTomato/ + ; Mycn fl/fl mice and control Dcx-creER/ + ; TdTomato/ + ; + / + mice, more TdTomato ( +) cells with Mycn deletion migrated away from the inner layer to the middle layer of dentate gyrus compared to the control TdTomato ( +) cells with intact Mycn (Fig. 7). Taken together, these results indicate that Mycn inhibits the maturation of neuroblasts in adult dentate gyrus.
Mycn inhibits the migration of Dcx-expressed cells in dentate gyrus. a The distribution of Td-Tomato + cells in the dentate gyrus of young adult Dcx-creER/ + ; Td-Tomato/ + ; + / + (control) versus Dcx-creER/ + ; Td-Tomato/ + ; mycn fl/fl (cKO) mice on day 14 after the 5-day tamoxifen treatment. In these mice, the cells that had expressed Dcx 14–19 days ago were fate mapped with Td-Tomato. b A model for dividing the dental gyrus granular layer into 4 areas. c Quantitative analysis of a shows the distribution of Td-Tomato ( +) cells in different dentate gyrus granular layers. Significantly more Td-Tomato ( +) cells had migrated from domain 1 into domain 2 in the Dcx-creER/ + ; Td-Tomato/ + ; mycn fl/fl (cKO) mice compared to the control Dcx-creER/ + ; Td-Tomato/ + ; + / + mice. Data are shown as mean ± SEM, and the analysis performed was two-way ANOVA
Mycn Inhibits the Survival of Newly Generated Cells in Sites of Adult Neurogenesis and Oligodendrogenesis
To monitor the fate of proliferated cells in the sites of adult neurogenesis and oligodendrogenesis, we crossed Mki67-creER line with TdTomato reporter line and Mycn flox line. Treating young adult Mki67-creER/ + ; TdTomato/ + ; Mycn fl/fl mice and control Mki67-creER/ + ; TdTomato/ + ; + / + mice with daily tamoxifen for 5 days fate labeled all the cells that have expressed Mki67 in last 5 days with TdTomato, along with deletion of Mycn from TdTomato-labeled cells in Mki67-creER/ + ; TdTomato/ + ; Mycn fl/fl mice. Fourteen days after the last tamoxifen treatment in the control Mki67-creER/ + ; TdTomato/ + ; + / + mice, 94% of TdTomato ( +) cells in SVZ were DCX ( +) but none of them were NeuN ( +) (Supplementary Figs. 6a and 7), whereas in dentate gyrus, 65% of TdTomato ( +) cells were DCX ( +) and 72% of TdTomato ( +) cells were NeuN ( +) (Supplementary Figs. 6b and 7a, b), and in OB, 36% of TdTomato ( +) cells were DCX ( +) and 63% of TdTomato ( +) cells were NeuN ( +) (Supplementary Figs. 6c and 7a, b). These results suggest that most of the proliferating cells in SVZ, dentate gyrus, and OB develop into neuroblasts or mature neurons. On the other hand, 66% of TdTomato ( +) cells in SCZ and 83% of TdTomato ( +) cells in CC were Olig2 ( +) (Supplementary Figs. 6d, e and 7c), suggesting that the cells which proliferated 2 weeks ago mainly developed into oligodendrocyte lineage cells in SCZ and CC. Very little, if any, of the TdTomato ( +) cells in these regions were microglia or astrocytes (Supplementary Fig. 6).
We then compared the TdTomato ( +) cells between Mki67-creER/ + ; TdTomato/ + ; Mycn fl/fl mice and control Mki67-creER/ + ; TdTomato/ + ; + / + mice 14 days after the last tamoxifen treatment and found that animals with Mycn deletion in Mki67-expressing cells had significantly more TdTomato ( +) cells than the control animals in SVZ and CC (Fig. 8a–c), SGZ and SCZ (Fig. 8d–f), and OB (Fig. 8g, h), suggesting that Mycn inhibits the survival of newly generated cells in these regions.
Mycn inhibits the survival of newly generated cells in young adult brain. Fate mapping of newly generated Td-Tomato ( +) cells in the SVZ and CC (a), dentate gyrus and SCZ (d), and OB (g) of young adult Mki67-creER/ + ; Td-Tomato/ + ; + / + (control) versus Mki67-creER/ + ; Td-Tomato/ + ; Mycn fl/fl (cKO) mouse 14 days after the 5-day tamoxifen treatment. In these mice, the cells that had expressed Mki67 14–19 days ago were fate mapped with Td-Tomato. The dashed lines in a show the boundary between SVZ and CC. Scale bar = 100 µm. Quantification of a, d, and g shows that the deletion of Mycn from Mki67-expressing cells significantly enhanced the number of newly generated cells in SVZ (b), CC (c), dentate gyrus (e), SCZ (f), and OB (h). Data are shown as mean ± SEM, and the analysis performed was unpaired two-tailed Student’s t test
Discussion
Human MYCN is an oncogene involved in many tumors, including neuroblastoma, which is characterized as inappropriate proliferation of undifferentiated neuronal progenitor cells [5]. The copy numbers of MYCN are closely associated with the aggressiveness of neuroblastoma [28], with increased expression of MYCN that increases the proliferative potential of neuroblastoma cells [29]. Inhibition of MYCN function suppresses the proliferation and enhances the neuronal differentiation of MYCN-amplified neuroblastoma cells [30], and with different conditions, MYCN can either promote or inhibit apoptosis of neuroblastoma cells [5]. In addition to tumorigenesis, MYCN is also crucial for embryonic development, as it is expressed in human fetal brain [9], and heterozygous MYCN mutation causes microcephaly and learning disability in human [10]. Similarly, mouse Mycn is also expressed in embryonic central and peripheral nervous systems, and homozygous Mycn mutant mouse dies between 10.5 and 12.5 days of gestation [12, 13]. Deletion of Mycn from embryonic neuronal stem cells severely reduces the proliferation of embryonic neuronal progenitor cells and dramatically increases neuronal differentiation, with limited effect on apoptosis of embryonic neuronal progenitor cells [14]. On the other hand, overexpression of Mycn in mouse multipotent sympathoadrenal progenitors promotes proliferation, apoptosis, and neural lineage commitment [31]. In this study, we demonstrated that Mycn plays a crucial role for adult neurogenesis and oligodendrogenesis in mice.
Mycn Is Expressed in Adult Brain
With Mycn-EGFP reporter mice, we found that Mycn is expressed in SVZ (Fig. 1a) and SGZ (Fig. 1b), two major sites of adult neurogenesis, as well as in OB (Fig. 1c), CC (Fig. 1a), and SCZ (Fig. 1b). Our further analyses revealed that many Mycn-EGFP cells co-expressed the neuroblast marker DCX in SVZ, SGZ, SCZ, and OB (Fig. 1d–g), as well as the proliferation marker Ki67 in SVZ, SGZ, OB, SCZ, and CC (Fig. 3a–e). Some Mycn-EGFP cells were also found in the granular layer of dentate gyrus, and these cells were also NeuN ( +) (Fig. 1j), suggesting that Mycn was expressed in certain hippocampal neurons, like those newly differentiated from neuroblasts. In addition, some Mycn-EGFP cells in CC also co-expressed the oligodendrocyte marker Olig2 (Fig. 1l, m), indicating that Mycn was expressed in oligodendrocyte lineage cells. Indeed, analysis of the published scRNA-Seq result from adult mouse dentate gyrus [20] showed that Mycn is mainly expressed in nIPCs, neuroblasts, and immature GCs and in small portions of the mature GDs (Supplementary Fig. 2a), similar to the expression pattern of Dcx (Supplementary Fig. 2b). Interestingly, although almost all DCX ( +) cells in these regions co-expressed GFP in Mycn-EGFP reporter mice (Fig. 1j), not all GFP ( +) cells were DCX ( +) (Fig. 1i). As the DCX ( +) cells were detected by DCX IHC in this study, some cells with a relatively low level of Dcx expression might not be detected by DCX IHC.
Mycn Is Required for the Proliferation of Adult nIPCs and Neuroblasts
By deletion of Mycn from adult Dcx-expressing cells or in adult Mki67-expressing proliferating cells, we demonstrated that Mycn is required for the proliferation of nIPCs and neuroblasts in SVZ, SGZ, and OB. Unlike in SVZ where the proliferation is mildly reduced in Mycn cKO animals (Fig. 5b), in SGZ and OB, the proliferation is almost completely prevented in Mycn cKO animals (Fig. 5d, f), suggesting that Mycn is absolutely required for the proliferation of nIPCs and neuroblasts in SGZ and OB, whereas the proliferation in SVZ has Mycn-independent mechanism. The effect of Mycn on the proliferation of adult nIPCs and neuroblasts is likely mediated by the axis of cyclin-dependent kinase (CDK)-retinoblastoma protein (RB)-E2F transcription factor family, a core transcriptional machinery that determines if a cell exits the G1 phase to enter the S phase of cell cycle [32], because CDK6, RB, and E2F1 are required for the proliferation of adult neuronal progenitors and immature newborn dentate granule cell neurons [33,34,35], and knockdown of MYCN resulted in downregulation of CDK6 and E2F1 in neuroblastoma cells [36]. Deletion of Mycn from embryonic neuronal progenitor cells or knockdown of MYCN from neuroblastoma cells also upregulates CDK inhibitor p27 [14, 36], which is known to modulate CDK6 kinase activity in adult hippocampus to control the expansion of neuronal progenitor cells [34].
Mycn Inhibits the Maturation of Adult Neuroblasts and the Migration of Newly Generated Neurons
With the fate mapping of Dcx-expressing cells in dentate gyrus, we found that the Dcx-expressing cells with Mycn deletion matured faster than the control Dcx-expressing cells (Fig. 6), and the newly generated granule neurons with Mycn deletion migrated further than the control neurons (Fig. 7), suggesting that Mycn inhibits the maturation of adult neuroblasts and the newly generated neurons. Similarly, knockdown MYCN promotes neuronal differentiation in neuroblastoma [37], and retinoic acid, which induces morphological differentiation of neuroblastoma cells [38], inhibits the expression of MYCN in these tumor cells [39].
Mycn Inhibits the Survival of Adult Neuronal Progenitor Cells
With the fate mapping of Mki67-expressing proliferated cells in SVZ and CC (Fig. 8a–c), SGZ and SCZ (Fig. 8d–f), and OB (Fig. 8g, h) 14 days after they had expressed Mki67, we found that Mycn deletion from Mki67-expressing cells increased the number of the proliferated cells. The phenomenon of increased number of the proliferated cells was not the result of more cells being proliferated, because Mki67-expressing cells with Mycn deletion proliferated significantly less (Fig. 5). These results suggest that Mycn inhibits the survival of adult neuronal progenitor cells. The mechanism on how Mycn regulates the survival of adult neuronal progenitor cells needs to be further studied.
Mycn Is Required for the Genesis of Adult Oligodendrocytes
In Mycn-EGFP reporter mice, we found GFP ( +) cells in CC (Fig. 1a) and SCZ (Fig. 1b), a known adult oligodendrogenesis site between hippocampus and CC [19]. Our fate-mapping study of Dcx-expressing cells revealed that these GFP ( +) cells in CC originate from the Dcx-expressing cells in SVZ (Fig. 2d), and our IHC study showed that some of the GFP ( +) cells in SCZ co-expressed DCX (Fig. 1f). Some of these GFP ( +) cells in CC and SCZ also co-expressed Ki67 (Fig. 3a, c), suggesting that they were proliferating cells, and our fate-mapping studies showed that the majority of Mki67-expressing cells in these regions developed into Olig2 ( +) oligodendrocytes (Supplementary Figs. 6d, e and 7c). In fact, some of the GFP ( +) cells in CC were Olig2 ( +) (Fig. 1l). These results suggest that the Mycn-expressing cells in these regions developed into oligodendrocyte lineage cells. Indeed, analysis of published scRNA-Seq from cells of oligodendrocyte lineage [24] showed that Mycn is expressed in OPCs, CDPs, and certain MFOLs and MOLs (Supplementary Fig. 3a).
We found that deletion of Mycn from Mki67-expressing cells blocked cell proliferation in SCZ (Fig. 5c, d), suggesting that Mycn is required for the cell proliferation in this region. In addition, with fate mapping of proliferated cells, we found significantly more survived cells that have recently proliferated in CC and SCZ of Mycn cKO animals (Fig. 8c, f). Thus, like in adult neurogenesis, Mycn promotes cell proliferation and inhibits cell survival in adult oligodendrogenesis sites. Interestingly, CDK4, rather than CDK6, is essential for the proliferation of adult oligodendrocyte progenitor cells [40], and knockdown of CDK4 in neuroblastoma cells reduces their proliferation [41]. In addition, both E2F1 [42] and p27 [43] contribute to the regulation of oligodendrocyte progenitor proliferation. Thus, it would be interesting to investigate the possible interaction between Mycn and CBK-BR-E2F pathway in adult oligodendrogenesis.
Data Availability
No data or material needs to be submitted.
Code Availability
No code needs to be submitted.
References
Schwab M, Alitalo K, Klempnauer KH et al (1983) Amplified DNA with limited homology to myc cellular oncogene is shared by human neuroblastoma cell lines and a neuroblastoma tumour. Nature 305(5931):245–248
Wright JH (1910) Neurocytoma or neuroblastoma, a kind of tumor not generally recognized. J Exp Med 12(4):556–561
Brodeur GM, Seeger RC, Schwab M et al (1984) Amplification of N-myc in untreated human neuroblastomas correlates with advanced disease stage. Science 224(4653):1121–1124
Weiss WA, Aldape K, Mohapatra G et al (1997) Targeted expression of MYCN causes neuroblastoma in transgenic mice. EMBO J 16(11):2985–2995
Ruiz-Perez MV, Henley AB, and Arsenian-Henriksson M (2017) The MYCN protein in health and disease. Genes (Basel) 8(4)
Cancer Genome Atlas Research, N (2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455(7216):1061–1068
Ghasemi DR, Sill M, Okonechnikov K et al (2019) MYCN amplification drives an aggressive form of spinal ependymoma. Acta Neuropathol 138(6):1075–1089
Kohl NE, Legouy E, DePinho RA et al (1986) Human N-myc is closely related in organization and nucleotide sequence to c-myc. Nature 319(6048):73–77
Grady EF, Schwab M, Rosenau W (1987) Expression of N-myc and c-src during the development of fetal human brain. Cancer Res 47(11):2931–2936
van Bokhoven H, Celli J, van Reeuwijk J et al (2005) MYCN haploinsufficiency is associated with reduced brain size and intestinal atresias in Feingold syndrome. Nat Genet 37(5):465–467
Zimmerman KA, Yancopoulos GD, Collum RG et al (1986) Differential expression of myc family genes during murine development. Nature 319(6056):780–783
Stanton BR, Perkins AS, Tessarollo L et al (1992) Loss of N-myc function results in embryonic lethality and failure of the epithelial component of the embryo to develop. Genes Dev 6(12A):2235–2247
Charron J, Malynn BA, Fisher P et al (1992) Embryonic lethality in mice homozygous for a targeted disruption of the N-myc gene. Genes Dev 6(12A):2248–2257
Knoepfler PS, Cheng PF, Eisenman RN (2002) N-myc is essential during neurogenesis for the rapid expansion of progenitor cell populations and the inhibition of neuronal differentiation. Genes Dev 16(20):2699–2712
Kramer M, Ribeiro D, Arsenian-Henriksson M et al (2016) Proliferation and survival of embryonic sympathetic neuroblasts by MYCN and activated ALK signaling. J Neurosci 36(40):10425–10439
Blelloch R, Venere M, Yen J et al (2007) Generation of induced pluripotent stem cells in the absence of drug selection. Cell Stem Cell 1(3):245–247
Gong S, Zheng C, Doughty ML et al (2003) A gene expression atlas of the central nervous system based on bacterial artificial chromosomes. Nature 425(6961):917–925
Bond AM, Ming GL, Song H (2015) Adult mammalian neural stem cells and neurogenesis: five decades later. Cell Stem Cell 17(4):385–395
Seri B, Herrera DG, Gritti A et al (2006) Composition and organization of the SCZ: a large germinal layer containing neural stem cells in the adult mammalian brain. Cereb Cortex 16(Suppl 1):i103-111
Hochgerner H, Zeisel A, Lonnerberg P et al (2018) Conserved properties of dentate gyrus neurogenesis across postnatal development revealed by single-cell RNA sequencing. Nat Neurosci 21(2):290–299
Elbaz B, Popko B (2019) Molecular control of oligodendrocyte development. Trends Neurosci 42(4):263–277
Zhang J, Giesert F, Kloos K et al (2010) A powerful transgenic tool for fate mapping and functional analysis of newly generated neurons. BMC Neurosci 11:158
Shaner NC, Campbell RE, Steinbach PA et al (2004) Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat Biotechnol 22(12): 1567–1572
Marques S, Zeisel A, Codeluppi S et al (2016) Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science 352(6291):1326–1329
Tobin MK, Musaraca K, Disouky A et al (2019) Human hippocampal neurogenesis persists in aged adults and Alzheimer’s disease patients. Cell Stem Cell 24(6):974-982.e3
Salic A, Mitchison TJ (2008) A chemical method for fast and sensitive detection of DNA synthesis in vivo. Proc Natl Acad Sci U S A 105(7):2415–2420
Duan X, Chang JH, Ge S et al (2007) Disrupted-in-schizophrenia 1 regulates integration of newly generated neurons in the adult brain. Cell 130(6):1146–1158
Seeger RC, Brodeur GM, Sather H et al (1985) Association of multiple copies of the N-myc oncogene with rapid progression of neuroblastomas. N Engl J Med 313(18):1111–1116
Schweigerer L, Breit S, Wenzel A et al (1990) Augmented MYCN expression advances the malignant phenotype of human neuroblastoma cells: evidence for induction of autocrine growth factor activity. Cancer Res 50(14):4411–4416
Zirath H, Frenzel A, Oliynyk G et al (2013) MYC inhibition induces metabolic changes leading to accumulation of lipid droplets in tumor cells. Proc Natl Acad Sci U S A 110(25):10258–10263
Mobley BC, Kwon M, Kraemer BR et al (2015) Expression of MYCN in multipotent sympathoadrenal progenitors induces proliferation and neural differentiation, but is not sufficient for tumorigenesis. PLoS One 10(7): e0133897
Kent LN, Leone G (2019) The broken cycle: E2F dysfunction in cancer. Nat Rev Cancer 19(6):326–338
Cooper-Kuhn CM, Vroemen M, Brown J et al (2002) Impaired adult neurogenesis in mice lacking the transcription factor E2F1. Mol Cell Neurosci 21(2):312–323
Caron N, Genin EC, Marlier Q et al (2018) Proliferation of hippocampal progenitors relies on p27-dependent regulation of Cdk6 kinase activity. Cell Mol Life Sci 75(20):3817–3827
Fong BC, and Slack RS (2017) RB: an essential player in adult neurogenesis. Neurogenesis (Austin) 4(1): e1270382
Woo CW, Tan F, Cassano H et al (2008) Use of RNA interference to elucidate the effect of MYCN on cell cycle in neuroblastoma. Pediatr Blood Cancer 50(2):208–212
Negroni A, Scarpa S, Romeo A et al (1991) Decrease of proliferation rate and induction of differentiation by a MYCN antisense DNA oligomer in a human neuroblastoma cell line. Cell Growth Differ 2(10):511–518
Sidell N (1982) Retinoic acid-induced growth inhibition and morphologic differentiation of human neuroblastoma cells in vitro. J Natl Cancer Inst 68(4):589–596
Thiele CJ, Reynolds CP, Israel MA (1985) Decreased expression of N-myc precedes retinoic acid-induced morphological differentiation of human neuroblastoma. Nature 313(6001):404–406
Nobs L, Nestel S, Kulik A et al (2013) Cyclin D1 is required for proliferation of Olig2-expressing progenitor cells in the injured cerebral cortex. Glia 61(9):1443–1455
Molenaar JJ, Ebus ME, Koster J et al (2008) Cyclin D1 and CDK4 activity contribute to the undifferentiated phenotype in neuroblastoma. Cancer Res 68(8):2599–2609
Magri L, Swiss VA, Jablonska B et al (2014) E2F1 coregulates cell cycle genes and chromatin components during the transition of oligodendrocyte progenitors from proliferation to differentiation. J Neurosci 34(4):1481–1493
Jablonska B, Scafidi J, Aguirre A et al (2012) Oligodendrocyte regeneration after neonatal hypoxia requires FoxO1-mediated p27Kip1 expression. J Neurosci 32(42):14775–14793
Funding
Z.G. is supported by the National Institute of Neurological Disorder and Stroke R01NS121144.
Author information
Authors and Affiliations
Contributions
J.C. performed and analyzed the data of the experiments and wrote the manuscript. Z.G. designed the experiments, wrote the manuscript, and supervised all aspects of the study.
Corresponding author
Ethics declarations
Ethics approval and Consent to Participate
All animal procedures were performed according to protocols approved by Institutional Animal Care and Use of Laboratory Animals. Consent to participate is not needed.
Consent for Publication
All the data is suitable for publication.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, J., Guan, Z. Function of Oncogene Mycn in Adult Neurogenesis and Oligodendrogenesis. Mol Neurobiol 59, 77–92 (2022). https://doi.org/10.1007/s12035-021-02584-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-021-02584-7