Abstract
All-trans retinoic acid (ATRA) plays key roles in neurogenesis mediated by retinoic acid receptors (RARs). RARs are important targets for the therapeutic regulation of neurogenesis but effective drug development depends on modelling-based strategies to design high-specificity ligands in combination with good biological assays to discriminate between target-specificity and off-target effects. Using neuronal differentiation as a model, the aim of this study was to test the hypothesis that responses across different temporal scales and assay platforms can be used as comparable measures of retinoid activity. In biological assays based on cell phenotype or behaviour, two structurally similar synthetic retinoids, differing in RAR affinity and specificity, retained their relative activities across different temporal scales. In contrast, assays based on the transcriptional activation of specific genes in their normal genomic context were less concordant with biological assays. Gene-induction assays for retinoid activity as modulators of neurogenesis require careful interpretation in the light of variation in ligand-receptor affinity, receptor expression and gene function. A better characterization of neuronal phenotypes and their regulation by retinoids is badly needed as a framework for understanding how to regulate neuronal development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Retinoic acid (RA) receptor signalling plays key roles in cell and tissue patterning, neurogenesis and homeostasis, both directly via nuclear retinoic acid receptors (RARs) and indirectly by interactions with other ligand-dependent signalling mechanisms via shared co-factors, receptor partners and ligand cross-talk [1, 2]. This signalling diversity underlies the potential of RA and related compounds as important drugs for medicinal use, ranging from cancer therapeutics to novel treatments for diseases associated with ageing and neuronal health. In normal cellular and tissue development, intracellular levels of the main biologically active RA isomer, all-trans RA (ATRA), are finely regulated by conversion from vitamin A (retinol), via cellular binding proteins which mediate the transfer of retinol to retinol dehydrogenases for conversion to ATRA. The transfer of ATRA to RARs is mediated by cellular retinoic acid binding proteins (CRABPs) to achieve transcriptional regulation [3, 4] as part of normal cellular homeostasis, and to drive cell and tissue patterning during embryogenesis and tissue differentiation [1, 4, 5].
RARs are encoded by the transcripts of three separate genes, RARA (RAR-α), RARB (RAR-β) and RARG (RAR-γ), and specificity in responses at a cellular level are driven by tissue- and stage-specific variation in gene expression. This is also coupled with variation of splicing patterns to generate N-terminal variants [6] facilitating combinatorial interactions with different transcriptional coregulators [7]. Temporal regulation of gene expression is facilitated by ATRA degradation mediated by specific cytochrome P450 enzymes which are themselves regulated by ATRA [8, 9]. One consequence of this finely tuned metabolism is that ATRA has a short lifetime when added to cells or used therapeutically in vivo [10, 11]. Thus, although the spatial and temporal regulation of ATRA synthesis and delivery to RARs provides exquisite control of ATRA-dependent gene expression, this also provides significant challenges for the development of drugs to regulate ATRA signalling for clinical benefit. The key requirements for such drugs will be stability, so that ligand concentrations can be maintained in relevant cells and tissues, and providing sufficient RAR specificity to target particular processes, tissues or cell types.
There has been considerable progress in designing retinoid-like molecules which are considerably more stable than ATRA in intra- and extra-cellular environments [12]. To be classed as a retinoid, compounds should produce cellular effects by specific interactions with the ligand binding domain of the RARs. Recent modelling studies have shown that, despite the high degree of sequence conservation, there are important differences between receptor types in the shape of the ligand binding pocket [13]; this has implications for the design of modified stable synthetic retinoids for targeting specific biological processes. Furthermore, it is not just necessary to ensure good fit of the synthetic retinoid into the ligand binding domain (LBD) of RARs but also to ensure that the ligand-LBD complex has sufficient structural integrity to ensure effective coactivator recruitment for transcriptional regulation [14, 15].
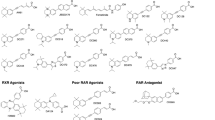
ATRA has an important role in neurogenesis [17], and we have developed the synthetic retinoids 4-(5,5,8,8-tetramethyl-5,6,7,8-tetrahydronaphthalen-2-ylethynyl)benzoic acid (para-isomer; EC23) and 3-(5,5,8,8-tetramethyl-5,6,7,8-tetrahydronaphthalen-2-ylethynyl)benzoic acid (meta-isomer; EC19) [18] as tools for studying neurogenesis in vitro (Fig. 1). These compounds are chemically and biologically more stable than ATRA; receptor binding and molecular docking studies show that EC23 binds to all three RARs in a manner similar to ATRA whereas EC19 has fewer interactions between key residues in the RAR-α and RAR-γ binding pockets while being a better fit to the larger binding pocket of RAR-β [13]. However, biological models are essential for assessing novel retinoids as potential clinical drugs or experimental tools and to discriminate between RAR-dependent activity and non-specific or downstream effects. In addition, ATRA may have distinct concentration-dependent effects in promoting alternative differentiation pathways [16] and for cellular homeostasis [17]. Temporal scale in biological models is of critical importance, and many assays of retinoid activity are carried out at long timescales which may make results hard to reconcile with known RAR specificity.
Chemical structures of ATRA and the synthetic analogues, EC23, EC19 and methyl esters. [9]
The aim of this study was to test the hypothesis that short- and long-term responses can be used as equivalent measures of retinoid activity and metabolic stability in assays of synthetic retinoids in neurogenesis models. Neurogenesis and gene expression at different temporal scales were compared in TERA2.cl.SP12 pluripotent stem cells and SH-SY5Y neuroblastoma cells in response to EC19 and EC23, and their methyl esters, using ATRA as the positive control. The methyl esters were included for some assays because although they show reduced RAR binding activity as a result of the absence of the key carboxylic acid-arginine residue interaction [13], esters may be relevant for biological studies if the parent compounds are released by endogenous esterase activity.
Material and Methods
Retinoid Solutions
Stock solutions of synthetic retinoids EC19 and EC23 and their methyl esters were prepared as reported earlier [18]; ATRA was from (Sigma-Aldrich, Poole, UK). All compounds were dissolved in DMSO (Sigma-Aldrich) to 10 mM. Aliquot stock solutions were stored at −20 °C in the dark.
Cell Culture
Human pluripotent TERA2.cl.SP12 embryonal carcinoma stem cells were cultured [19], under low-light conditions to minimize retinoid isomerisation, in Dulbecco’s modification of Eagle’s medium (DMEM; Sigma-Aldrich) supplemented with 10% FCS (Gibco), 2 mM L-glutamine and 100 units each of penicillin and streptomycin (Gibco). Cultures were passaged using acid-washed glass beads unless a single-cell suspension was required for counting, in which case, a 0.25% trypsin/EDTA (Lonza) solution was used. Human SH-SYSY neuroblastoma cells were cultured in DMEM F12/Ham (1:1) containing 2 mM L-glutamine, supplemented with 10% FCS at 37 °C with 5% CO2 in air [20]. Cell suspensions were obtained by treating adherent cells with 1 ml sterile PBS and incubation at 37 °C for 3–5 min. Culture media were replaced every 3–4 days.
Flow Cytometry
Flow cytometry was carried out on live TERA2.cl.SP12 cells, incubated at a density of 0.2 × 106 cells per 25-cm2 flask for 12–24 h before treatment with 10 μM retinoid for 7 days. Specific cell-surface primary antibodies were used: SSEA-3 (1:10), (University of Iowa Hybridoma Bank), TRA-1-60 (1:50), (Abcam) and neural cell marker A2B5 (1:40), (R&D Systems). Cell suspensions were centrifuged at 1000 rpm and resuspended in wash buffer (0.1% BSA in PBS) and added to 96-well plate for incubation with the primary antibodies, followed by several washes and then incubation with FITC-conjugated secondary antibody IgM (1:128) (Sigma-Aldrich). Labelled cells were analysed in a GuaveEasyCytePlus System (Millipore) flow cytometer and thresholds determining the numbers of positively expressing cells were set against the negative control antibody, P3X, a generous gift from Prof. P. Andrews, Sheffield University.
Gene Expression Analysis
Real-time quantitative PCR (qPCR) was carried out on cell lysates immediately after detachment with 0.25% trypsin–EDTA. Cells were seeded at a density of 1 × 106 cells per 25 cm2 flask (BD falcon) 12–24 h before retinoid treatment. Commercial RNA extraction (Qiagen) and reverse transcription (Applied Biosystems) kits were purchased and procedures used according to manufacturer instructions. Real-time qPCR was performed using the TaqMan® Universal PCR master Mix (Life technologies) and TaqMan® gene expression system (Applied Biosystems) based on probe sets to the specific genes to be analysed: RAR-β (Hs00233407_m1), CYP26A1-A1 (Hs00175627_m1), RAR-α (Hs00940448_g1), RAR-γ (Hs01559234_m1), PAX-6 (Hs01088112_m1), NeuroD1 (Hs01922995_s1). GADPH (Hs02758991_g1) and ACTB (Hs99999903_m1) were used as internal control genes for TERA2.cl.SP12 and SH-SY5Y cells, respectively.
Immunocytochemistry
TERA-2.cl.SP12 cells were seeded at 5000 cells per well on poly-d-lysine (25 μg/ml) coated cover slips 22 × 22 mm, high precision (170 ± 5 μm) in 6-well plates. At the end of the experiment, cells were fixed in 4% para-formaldehyde (PFA) in PBS for 30 min at room temperature and rinsed with PBS. For intracellular staining, cells were permeabilised with 1% Triton-X-100 (Sigma) in PBS for 10 min at room temperature. Non-specific labelling was blocked by incubation for 1 h at room temperature with 1% goat serum (Sigma) in PBS with 0.2% Tween-20 (Sigma). Primary antibodies were diluted in blocking solution and incubated with cells for 1 h at room temperature with a β-III tubulin antibody (TUJ-1) 1:200 (Affymetrix eBioscience) or CK-8 antibody 1:500 (Affymetrix eBioscience). After washing with PBS, cells were incubated for 1 h in the dark with anti-mouse FITC-conjugated secondary antibody IgM 1:128 (Sigma) for A2B5 staining or anti-mouse Alexafluor 488 IgG 1:600 (Invitrogen) for TUJ-1 and CK-8 staining. Hoechst 33,342 nuclear staining dye (Molecular Probes) was used at 1:1000 in blocking solution after the secondary antibody step. Fixed and stained cells were visualized using a Leica SP5CLSM FLIM FCCS confocal microscope.
X-gal Bioassay
Sil-15 cells (F9-RARE-lacZ cells) [21] were plated in a 0.1% gelatin-coated 96-well plate and grown to about 85–90% confluence in DMEM containing 10% foetal calf serum (Invitrogen/Gibco) and 0.8 mg/ml G418 sulphate (Sigma) for selection. Cells were washed twice with PBS, fixed with 100 μl per well of 1% glutaraldehyde for 15 min, washed again twice with PBS and β-galactosidase activity was visualized with 100 μl of a freshly prepared X-Gal developing solution (5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside) added to each well. Colour was read on an Emax microplate reader at 650 nm.
Neurite Outgrowth
SH-SY5Y cells were fixed and stained for TUJ-1 (1:1000; Sigma) after retinoid treatment for 5 days. For each neurite outgrowth experiment, 3 cover slips (in 3 wells) were used. The numbers of neurites were counted and traced for length measurement using a semi-automatic NeuronJ plugin for ImageJ software in each of 10 randomly-selected images for each cover slip. Average neurite length was calculated by dividing the total neurite length by the total number of neurites per image.
Results
Short-Term Responses to Retinoids: SH-SY5Y Cells
Retinoid-Induced Gene Expression
The induction of RAR-β or CYP26A1 transcription is well established as a marker of retinoid-response [22]; therefore, the expression of these genes was tested with respect to retinoid dose and time of treatment, in addition to investigating retinoid-induced changes in expression of RAR-α and RAR-γ. In SH-SY5Y cells, RAR-β expression in response to 10 μM ATRA or EC23 increased to a maximum at 6 h, with EC23 showing greater activity. In contrast, EC19 and the methyl ester showed no, or minimal, activity at 10 μM (Fig. 2a). With respect to the induction of CYP26A1, only ATRA had good activity, with a rapid and sustained induction from 2 to 10 h (Fig. 2c). In dose-response experiments from 1 to 0.001 μM, EC19 and EC23 had similar peak activities for induction of RAR-β at 0.1 μM which exceeded that of ATRA by about 30% (Fig. 2b). Conversely, for CYP26A1 induction, although the synthetic retinoids were not inactive, ATRA showed consistently higher levels of activity over the whole dose range compared to the synthetic retinoids; indeed, for the latter, EC19 had greater activity than EC23, except at 1 μM (Fig. 2d).
Short-term assay of retinoid properties in SH-SY5Y cells and Sil-15 reporter cells. a–f Real-time quantitative PCR (qPCR) analysis of mRNA expression for RAR-β (a time course, b dose-response), CYP26A1 (c time course, d dose-response), RAR-α (e time course) and RAR-γ (f time course) in SH-SY5Y cells treated with (left to right) ATRA, EC19, EC23 and methyl ester analogues. Dose-response experiments were performed with retinoid concentrations of 1, 0.1, 0.01 and 0.001 μM for 8 h, and time course experiments with 10 μM for up to 12 h. Quantification of target mRNA was relative to SH-SY5Y cells cultured with DMSO vehicle for the relevant time period and normalized to the internal reference gene (ACTB). Data represent mean ± SEM, n = 3. Neurite outgrowth by SH-SY5Y cells in response to control vehicle or EC19 or EC23 (10 μM each) is shown in the images (g–i), stained with the anti-beta III tubulin antibody TUJ-1 (scale bar 50 μm) (g control, h EC19, i EC23) and (j) bar graph for mean neurite length (pixels ± SEM, n = 3). There was a significant induction of neurite outgrowth in response to EC23 (ANOVA, P < 0.001) but not EC19, and no effect of dose at the concentrations used. The response by Sil-15 reporter cells to increasing concentrations of ATRA, EC19 or EC23 is shown in k where grey shading along the fitted curves defines the 95% confidence limits for the EC23 and EC19 data
As in some other cell types [23], ATRA induced RAR-α expression, but over a longer timescale than for RAR-β. In contrast to their induction of RAR-β, the synthetic retinoids were less effective at inducing RAR-α compared to ATRA; however, EC23 was at least twice as active as EC19 at the 12-h timepoint. Substantial induction of RAR-γ was only apparent after 12 h with ATRA and at this time EC19 and EC23 were only marginally more effective than their methyl esters (Fig. 2e, f).
Neurite Length
SH-SY5Y cells responded morphologically to retinoids with time-dependent increases in neurite length, with the greatest differential responses between the retinoids after 4 days or more [24]. At 0.01 and 10 μM, EC19 had no neurite-inducing capacity compared with the control vehicle (DMSO), unlike EC23, which had similar activity to ATRA at 0.01 μM (Fig. 2j); however, in contrast to ATRA which gave a dose-dependent increase in neurite length, the response to EC23 appeared saturated at 0.01 μM. Neurite extension is a lower-resolution technique over a more-extended timescale than the gene-expression assays, and there were clear differences between these assays in the relative activities of all retinoids at 10 μM (Fig. 2j).
X-Gal Reporter Bioassay Analysis
The relative transcriptional potency of the retinoids was also tested on Sil-15 reporter cells, which are F9 murine teratocarcinoma cells stably transfected with a LacZ gene under the control of a RA response element (RARE) promoter [21]. β-Galactosidase activity in these cells was assayed over retinoid concentration ranges of 10−6 M to 10−14 M over 24 h. EC23 was effective over the entire range, giving a 50% response between 10−11 to 10−12 M, compared to the 50% response of ATRA at 5 × 10−10 M. In contrast, EC19 showed very little activity (Fig. 2k).
Longer-Term Responses to Retinoids: TERA-2. cl.SP12 Cells
Cell Differentiation Markers
ATRA-induced differentiation of TERA2.cl.SP12 cells is characterized by the downregulation of the glycolipid antigen SSEA3 and the keratan sulphate-related antigen TRA-1-60, and the upregulation of the c-series ganglioside-specific antigen A2B5, characteristic of neuronal and glial cells [19]. TERA2.cl.SP12 cells were treated with 10 μM of each retinoid for 7 days and analysed for the expression of SSEA3, TRA-1-60 and A2B5 by flow cytometry. Control cultures, untreated or treated with the DMSO vehicle alone, showed high expression levels of SSEA-3 and TRA-1-60 with 60–70% of cells expressing these markers, but less than 20% of these cells expressing the neuronal differentiation marker A2B5 (Fig. 3a, b). After treatment with either 10 μM ATRA or EC23 for 7 days, there was a significant reduction in expression of SSEA-3 (20% of cells) and TRA-1-60 (50% of cells) and an induction of expression of A2B5, indicating a shift from a pluripotent state towards neuronal differentiation which was particularly marked with EC23; conversely, EC19 was less effective (Fig. 3a, b). The methyl ester of EC23 was less active than the parent compound, while the EC19 methyl ester was as active, or slightly more so, than EC19 (Fig. 3b).
Longer-term assay of retinoid responses using the cell differentiation model of TERA2.cl.SP12 stem cells treated with 10 μM ATRA, EC19, EC23 or their methyl esters after 7 days. Control TERA2.cl.SP12 cultures were treated with 1% DMSO vehicle. a Flow cytometry analysis showing 2D plots of gated cells expressing SSEA-3 and A2B5 after retinoid treatment. b Histograms of percentage positive cells for the stem cell-surface markers SSEA-3, TRA-1-60 and the early stage neuronal marker A2B5. Results are presented as ±SEM, n = 3; P values for comaprisons with control are: *P < 0.05; **P < 0.01; ***P < 0.001. c Photomicrographs of TERA2.cl.SP12 cells stained for the neuronal (A2B5 and TUJ-1) and epithelial (CK-8) proteins after exposure to 10 μM ATRA, EC19 or EC23 for 1, 2 and 3 weeks. Scale bar, 25 μm
Cell morphology and phenotypic fate within TERA2.cl.SP12 cell cultures were assessed by immunocytochemistry for neuronal markers (cell-surface A2B5; cytoplasmic βIII-tubulin, TUJ-1 antibody), and the epithelial marker cytokeratin 8 (CK-8). Control cultures (DMSO vehicle) showed low expression of all markers. After exposure to ATRA or EC23 for 7 days, expression of the neuronal markers substantially increased, in contrast to CK-8. After 14 and 21 days, βIII-tubulin expression increased further with the formation of more mature, differentiated neuronal cells where staining was localized to the cytoplasm and neuronal processes. In contrast, EC19 did not induce any substantial increase in expression of A2B5 or βIII-tubulin after 7 days, with low levels of βIII-tubulin-positive cells even after 21 days. However, the expression of CK-8 increased in EC19-treated cultures after 21 days, suggesting differentiation towards an epithelial phenotype (Fig. 3c).
Expression of Neuronal Lineage Marker Transcripts
The neuronal markers NeuroD1 and PAX-6 are predominantly expressed in the nervous system, particularly later in development [25, 26]. NeuroD1 was upregulated in TERA2.cl.SP12 cells showing a linear time-dependent increase in response to 10 μM ATRA or EC23; EC19 and the methyl esters of both synthetic retinoids had much lower activities (Fig. 4a). PAX6 was substantially upregulated only after 7 days of treatment and EC23 was 10 times more effective than ATRA with very little activity shown by EC19 (Fig. 4b).
NeuroD1 (a) and PAX6 (b) gene expression in TERA2.cl.SP12 stem cells treated with 10 μM of ATRA, EC19, EC23 or their methyl ester analogues for 3, 5 and 7 days. Quantification is relative to TERA2.cl.SP12 cells cultured with 0.1% DMSO vehicle for 7 days and all data (mean ± SEM, n = 3) were normalized to the internal reference gene (GADPH)
Discussion
With respect to the parent compounds, there were generally concordant responses, summarized by rank in Table 1, between long-term and short-term biological response assays (differentiation, neurite extension, reporter assays) where both EC23 and EC19 maintained their relative activities over different temporal scales. The methyl esters of EC23 and EC19 usually had low, or intermediate, activity. Although structural studies and ligand binding/activity assays suggest that these methyl esters may have direct activity as RAR ligands in their own right [13], esterase activity [27] releasing the free parent carboxylic acids may also contribute to biological activity. In contrast to biological response assays, assays of specific gene transcripts, whether as markers of neural differentiation status as with the NeuroD1 and PAX6 transcripts, or of short-term responses to retinoids such as the induction of expression of RARs or CYP26A1, were not directly comparable with the broader biological responses. This was particularly true for EC19 which, as predicted from structural studies, had activity in some gene-response assays but low activity in biological response assays. These results highlight two key factors in responses to retinoids: the dynamic mechanisms of individual gene regulation by retinoids and the retinoid mechanisms directing biological responses.
Individual retinoid-responsive genes, RAR-β and CYP26A1, responded differently with respect to temporal characteristics of activation and relative activities of the different retinoids at different doses. For CYP26A1 induction, ATRA showed the highest activity; unexpectedly, EC19 had greater CYP26A1 induction activity than EC23 at lower doses, but, in contrast to ATRA, both were ineffective at 10 μM. For RAR-β induction, both synthetic retinoids were equally effective at low doses and with greater activity than ATRA, whereas at higher doses, EC23 had the greatest activity with EC19 having much lower activity compared to ATRA.
Variability in behaviour between different short-term gene-response assays can be due to a combination of factors, particularly RARE context, RAR expression and specificity [28], and retinoid metabolism. Recent structural modelling studies have shown that the RAR-β LBD is better than other RAR LBDs at accommodating the geometrically differently substituted ring of EC19 with the carboxylic acid group in the meta-position [13]; the good activity of EC19 with respect to the induction of RAR-β suggests that this induction may be driven primarily by constitutive expression of RAR-β in these cells. This is in agreement with other studies [29] implying a dependence of RAR-β induction on RAR-β itself in SH-SY5Y cells. In the Sil-15 reporter cells, β-galactosidase activity is driven by an RAR-β RARE construct; the low activity of EC19 in this system compared to the induction of RAR-β transcripts in SH-SY5Y cells may result from differences in basal RAR-β expression between the two cell types, as this appears to be relatively lower in Sil-15 parental cells [30] compared to SH-SY5Y cells.
The genomic context of RAREs which drive marker genes, the availability of promoter-specific coregulators and retinoid-specific co-regulator interactions are also important considerations for the interpretation of retinoid activity assays. Transcriptional activation by ligand-bound RARs requires ligand-dependent conformational changes in the receptor to facilitate coactivator recruitment [31]; these may vary independently of ligand-LBD affinity such that retinoids with equivalent affinity for RAR LBDs may differ in their ability to facilitate coactivator recruitment [13]. Gene-specific induction mechanisms are also evidenced by the relatively poor activity of the synthetic retinoids on CYP26A1 induction compared to ATRA; in this respect, coregulators may be critical in driving specificity because CYP26A1 needs to respond to ATRA specifically to regulate ATRA levels. It is also possible that RARs may bind to other endogenous ligands, such as ATRA metabolites [32], and these should also be considered as potential drivers of CYP26A1 induction. Clearly, single gene assays have limited use as surrogate markers of the biological properties of synthetic retinoids.
Overall, these results imply that the assay based on Sil-15 cells, or on cells with an equivalent RARE-driven reporter, is a good short-term assay as it gave comparable results to longer-term, and more time-consuming, assays of biological responses for assessing the potency of novel synthetic retinoids. In the long-term assays, the relative induction of neurogenesis by different retinoids, as indicated by downregulation of markers of pluripotency and the upregulation of neuronal lineage markers, was comparable to the induction of RAR-α messenger RNA (mRNA) at a shorter timescale. This may imply a role for RAR-α as an initial step in the induction of neurogenesis, either via a transcriptional or non-transcriptional mechanism [33], and is supported by the different relative activities of EC19 in the induction of RAR-β and neurite extension in SH-SY5Y cells.
These studies also stress the importance of careful marker selection for retinoid assays. The transcription factor PAX6 is reported to promote neurogenesis [34], but in TERA2.cl.SP12 cells, PAX6 was upregulated substantially more in response to EC23 than ATRA, compared to the E-box transcription factor NeuroD1, implying that neuronal phenotypes may be driven to different extents by different retinoids. If neurogenesis is also regulated by the products of ATRA catabolism [9], then EC23 or comparable synthetic retinoids might have biological effects on neurogenesis that are qualitatively different to ATRA because of greater metabolic stability. As has been shown previously, EC19 does not induce neurogenesis of TERA2.cl.SP12 cells but induces an epithelial phenotype [18]; whether this is an RAR-driven process, perhaps mediated by different RAR specificities of EC19, is not clear. However, this could also result from non-specific effects such as arachidonic acid signalling as a consequence of high levels of lipophilic compounds impacting upon membrane lipids [20].
In summary, specific gene-induction assays for novel retinoids require careful interpretation as measures for their potential as modulators of neurogenesis. Changes in cell phenotype or behaviour over different temporal scales may, superficially, be simpler to interpret; nevertheless, a much better understanding and characterization of neuronal phenotypes and their regulation is badly needed to provide a framework for understanding the wider value of synthetic retinoids for regulating neuronal development.
References
Cunningham TJ, Duester G (2015) Mechanisms of retinoic acid signalling and its roles in organ and limb development. Nat Rev Mol Cell Biol 16:110–123. doi:10.1038/nrm3932
Janesick A, Wu SC, Blumberg B (2015) Retinoic acid signaling and neuronal differentiation. Cellular and molecular life sciences : CMLS 72:1559–1576. doi:10.1007/s00018-014-1815-9
Budhu AS, Noy N (2002) Direct channeling of retinoic acid between cellular retinoic acid-binding protein II and retinoic acid receptor sensitizes mammary carcinoma cells to retinoic acid-induced growth arrest. Mol Cell Biol 22:2632–2641
Rhinn M, Dolle P (2012) Retinoic acid signalling during development. Development 139:843–858. doi:10.1242/dev.065938
Duong V, Rochette-Egly C (2011) The molecular physiology of nuclear retinoic acid receptors. From health to disease Biochimica et Biophysica Acta 1812:1023–1031. doi:10.1016/j.bbadis.2010.10.007
Zelent A, Mendelsohn C, Kastner P, Krust A, Garnier JM, Ruffenach F, Leroy P, Chambon P (1991) Differentially expressed isoforms of the mouse retinoic acid receptor beta generated by usage of two promoters and alternative splicing. EMBO J 10:71–81
McKenna NJ, O'Malley BW (2002) Combinatorial control of gene expression by nuclear receptors and coregulators. Cell 108:465–474
Mahony S, Mazzoni EO, McCuine S, Young RA, Wichterle H, Gifford DK (2011) Ligand-dependent dynamics of retinoic acid receptor binding during early neurogenesis. Genome Biol 12:R2. doi:10.1186/gb-2011-12-1-r2
Sonneveld E, van den Brink CE, Tertoolen LG, van der Burg B, van der Saag PT (1999) Retinoic acid hydroxylase (CYP26) is a key enzyme in neuronal differentiation of embryonal carcinoma cells. Dev Biol 213:390–404. doi:10.1006/dbio.1999.9381
Adamson PC (1996) All-trans-retinoic acid pharmacology and its impact on the treatment of acute promyelocytic leukemia. Oncologist 1:305–314
Lansink M, van Bennekum AM, Blaner WS, Kooistra T (1997) Differences in metabolism and isomerization of all-trans-retinoic acid and 9-cis-retinoic acid between human endothelial cells and hepatocytes. Eur J Biochem 247:596–604
Clemens G, Flower KR, Gardner P, Henderson AP, Knowles JP, Marder TB, Whiting A, Przyborski S (2013) Design and biological evaluation of synthetic retinoids: probing length vs. stability vs. activity. Mol BioSyst 9:3124–3134. doi:10.1039/c3mb70273a
Haffez HR, Chisholm D, Valentine R, Pohl E, Redfern C, Whiting A (2017) The molecular basis of the interactions between synthetic retinoic acid analogues and the retinoic acid receptors. Medicinal Chemical Communications. doi:10.1039/C6MD00680A
Nettles KW, Greene GL (2005) Ligand control of coregulator recruitment to nuclear receptors. Annu Rev Physiol 67:309–333. doi:10.1146/annurev.physiol.66.032802.154710
Zechel C (2002) Synthetic retinoids dissociate coactivator binding from corepressor release. J Recept Signal Transduct Res 22:31–61. doi:10.1081/RRS-120014587
Okada Y, Shimazaki T, Sobue G, Okano H (2004) Retinoic-acid-concentration-dependent acquisition of neural cell identity during in vitro differentiation of mouse embryonic stem cells. Dev Biol 275:124–142. doi:10.1016/j.ydbio.2004.07.038
Jacobs S, Lie DC, DeCicco KL, Shi Y, DeLuca LM, Gage FH, Evans RM (2006) Retinoic acid is required early during adult neurogenesis in the dentate gyrus. Proc Natl Acad Sci U S A 103:3902–3907. doi:10.1073/pnas.0511294103
Christie VB, Barnard JH, Batsanov AS, Bridgens CE, Cartmell EB, Collings JC, Maltman DJ, Redfern CP et al (2008) Synthesis and evaluation of synthetic retinoid derivatives as inducers of stem cell differentiation. Organic & Biomolecular Chemistry 6:3497–3507
Przyborski SA, Christie VB, Hayman MW, Stewart R, Horrocks GM (2004) Human embryonal carcinoma stem cells: models of embryonic development in humans. Stem Cells Dev 13:400–408. doi:10.1089/scd.2004.13.400
Bell E, Ponthan F, Whitworth C, Westermann F, Thomas H, Redfern CPF (2013) Cell survival signalling through PPAR delta and arachidonic acid metabolites in neuroblastoma. PLos One 8 doi:10.1371/journal.pone.0068859
Wagner M, Han B, Jessell TM (1992) Regional differences in retinoid release from embryonic neural tissue detected by an in vitro reporter assay. Development 116:55–66
Armstrong JL, Taylor GA, Thomas HD, Boddy AV, Redfern CP, Veal GJ (2007) Molecular targeting of retinoic acid metabolism in neuroblastoma: the role of the CYP26 inhibitor R116010 in vitro and in vivo. Br J Cancer 96:1675–1683. doi:10.1038/sj.bjc.6603779
Halevy O, Lerman O (1993) Retinoic acid induces adult muscle cell differentiation mediated by the retinoic acid receptor-alpha. J Cell Physiol 154:566–572. doi:10.1002/jcp.1041540315
Lovat PE, Lowis SP, Pearson AD, Malcolm AJ, Redfern CP (1994) Concentration-dependent effects of 9-cis retinoic acid on neuroblastoma differentiation and proliferation in vitro. Neurosci Lett 182:29–32
Gaudilliere B, Konishi Y, de la Iglesia N, Yao G, Bonni A (2004) A CaMKII-NeuroD signaling pathway specifies dendritic morphogenesis. Neuron 41:229–241
Zhang J, Jiao J (2015) Molecular biomarkers for embryonic and adult neural stem cell and neurogenesis. Biomed Res Int 2015:727542. doi:10.1155/2015/727542
Wang CC, Hill DL (1977) Retinoid acid esterase activities: tissue and subcellular distribution in mice. Biochem Pharmacol 26:947–950
Rochette-Egly C, Germain P (2009) Dynamic and combinatorial control of gene expression by nuclear retinoic acid receptors (RARs). Nucl Recept Signal 7:e005. doi:10.1621/nrs.07005
Lindley D (2009) Retinoic acid receptor expression and activation and cellular responses to retinoic acid in neuroblastoma., Newcastle University,
Boylan JF, Lufkin T, Achkar CC, Taneja R, Chambon P, Gudas LJ (1995) Targeted disruption of retinoic acid receptor alpha (RAR alpha) and RAR gamma results in receptor-specific alterations in retinoic acid-mediated differentiation and retinoic acid metabolism. Mol Cell Biol 15:843–851
Bastien J, Rochette-Egly C (2004) Nuclear retinoid receptors and the transcription of retinoid-target genes. Gene 328:1–16. doi:10.1016/j.gene.2003.12.005
Pijnappel WW, Hendriks HF, Folkers GE, van den Brink CE, Dekker EJ, Edelenbosch C, van der Saag PT, Durston AJ (1993) The retinoid ligand 4-oxo-retinoic acid is a highly active modulator of positional specification. Nature 366:340–344. doi:10.1038/366340a0
Sarti F, Schroeder J, Aoto J, Chen L (2012) Conditional RARalpha knockout mice reveal acute requirement for retinoic acid and RARalpha in homeostatic plasticity. Front Mol Neurosci 5:16. doi:10.3389/fnmol.2012.00016
Kallur T, Gisler R, Lindvall O, Kokaia Z (2008) Pax6 promotes neurogenesis in human neural stem cells. Mol Cell Neurosci 38:616–628. doi:10.1016/j.mcn.2008.05.010
Acknowledgements
We would like to thank the Egyptian Council and Cultural Bureau, and Alzheimer Research UK Scotland for financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Haffez, H., Khatib, T., McCaffery, P. et al. Neurogenesis in Response to Synthetic Retinoids at Different Temporal Scales. Mol Neurobiol 55, 1942–1950 (2018). https://doi.org/10.1007/s12035-017-0440-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-017-0440-7