Abstract
The PICALM rs541458 T allele has been recognized as a risk factor for late-onset Alzheimer’s disease, and age might modulate the effects that genetic factors have on cognitive functions and brain. Thus, the current study intended to examine whether the effects of rs541458 on cognitive functions, brain structure, and function were modulated by age in non-demented Chinese elderly. We enrolled 638 subjects aged 50 to 82 years and evaluated their cognitive functions through a series of neuropsychological tests. Seventy-eight of these participants also received T1-weighted structural and resting state functional magnetic resonance imaging. Dividing subjects into groups <65 and ≥65 years old, results of neuropsychological tests showed that interactive effects of rs541458 × age existed with regard to executive function and processing speed after controlling for gender, years of education and APOE ε4 status. In addition, the effects of rs541458 on resting state functional connectivity of left superior parietal gyrus within left frontal-parietal network and on gray matter volume of left middle temporal gyrus were modulated by age. Furthermore, reduction of functional connectivity of left superior parietal gyrus was closely related with better executive function in the T allele carriers <65 years old. Further, greater volume of left middle temporal gyrus was significantly related to better executive function in both CC genotype <65 years old and CC genotype ≥65 years old groups, separately. Pending further confirmation from additional studies, our results support the hypothesis that the modulation of age, with respect to the rs541458, has interactional effects on cognitive performance, brain function, and structural measurements.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alzheimer’s disease (AD) is the most common type of dementia. Due to the sharp increase in the number of patients [1] and the high mortality rate [2], AD is currently an important global health problem. Among many risk factors, increased age has always been a pivotal risk factor for late-onset AD (LOAD) [3]. In fact, the prevalence of AD increases with age [4], almost doubling every 5 years after the age of 60 [5].
In addition to age, several well-recognized genetic factors augment the complexity of LOAD [6]. For instance, the apolipoprotein E (APOE) ε4 allele is the best known genetic LOAD risk factor, which is associated with increased incidence and a decreased age of onset of LOAD [7]. It has been suggested that the ε4 allele would likely account for approximately 50% or more of the LOAD cases in the USA [8, 9]. Large-scale genome-wide association studies (GWAS) have been carried out to locate additional susceptible loci to more fully understand the genetic etiology of LOAD. Among these loci is the one encoding the -binding clathrin assembly protein (PICALM) [10]. The 112 kb PICALM gene is located on chromosome 11q14. In the central nervous system, PICALM has primarily been identified in neurons, astrocytes, and oligodendrocytes [11]. Evidence has shown that PICALM plays a major role in clathrin-mediated endocytosis [12], in which it recruits clathrin and adaptor protein 2 to the plasma membrane and transports target proteins to be processed in lysosomes or endosomes [12, 13]. Thus, PICALM would be further associated with amyloid precursor protein (APP) metabolism [14, 15], which is hypothesized to play a key role in AD pathogenesis.
The SNP rs541458 is located 8 kb upstream of the PICALM gene [16], with C as the minor allele and T as the major allele. An early GWAS study of Europeans found the rs541458 polymorphism to be a susceptible site for LOAD [17]. Subsequently, Lambert et al. verified the risk of the T allele of this SNP in a meta-analysis by examining three different European populations [18]. Other studies of Caucasians also reported the association between the rs541458 T allele and the LOAD risk [19, 20]. However, the LOAD risk associated with the rs541458 polymorphism was not duplicated in samples from the southern China region [21]. These results suggest that the effects of the rs541458 polymorphism might not be consistent among different ethnic groups.
Because of the crucial role of the brain in aging biology [22], many groups have investigated the patterns of brain aging with respect to brain structure and function [23, 24]. Additionally, the effects of a specific gene such as PICALM on the brain are not static but are dynamic and aging-related [25], indicating that age would modulate the effects of genetic factors in the brain. A previous study by Sweet et al. showed that the rs541458 T allele was associated with an earlier age at the midpoint of general cognitive function decline [26]. However, few studies have referred to the regulating effects of age on the rs541458 SNP based on brain measures. Taking 65 years of age or older as the onset age of LOAD [27], we divided the elderly into young (<65 years old) and old (≥65 years old) groups to assess the age modulation effects.
Indeed, cognitive deficits are often thought to be associated with brain functional and/or structural impairments [28, 29], which can be measured by using volumetric or functional magnetic resonance imaging (MRI) neuroimaging techniques. For example, resting state blood-oxygen-level dependent (BOLD) MRI allowed the delineation of the human neural functional architecture into multinetworks, such as the default mode network (DMN) and frontal-parietal network (FPN) [30]. Some studies have highlighted a number of functional networks showing significant age-related changes, including in the DMN and left FPN [31, 32]. The APOE disrupted the brain network connectivity, including the connectivity of both the posterior [33, 34] and anterior [35] DMNs as well as the left FPN [36]. Additionally, the risk allele (i.e., G allele) of the PICALM rs3851179 polymorphism was associated with weaker negative functional connectivity between the left hippocampus and the right precuneus and between the right hippocampus and the left superior frontal gyrus [37]. By using T1-weighted volumetric MRI to assess brain structures, a previous study suggested that atrophy was primarily located in the temporal and parietal regions in normal aging [38]. A recent study demonstrated that rs3851179 had a nominally significant main effect on hippocampal volume in healthy, young subjects [37], although negative findings have been reported in the elderly [39]. This may suggest that age would modulate the association between heredity and the human brain.
For the rs541458 SNP, we feel that relatively little is known about its effects on the resting state functional connectivity and whole brain gray matter volumes as well as neuropsychological performance (see the study by Sweet et al. [26]) and, more importantly, whether age could modulate such effects. Although it is weak, the existing evidence for PICALM itself and for genetic risk factors motivated us to make the following hypothesis: the effects of the PICALM rs541458 polymorphism on cognitive function, functional networks, and gray matter volumes are modulated by age. In the present study, we first examined such an age moderation effect on neuropsychological performance in a larger sample of non-demented Chinese elderly. Second, we investigated this modulated effect of age on functional connectivity and gray matter volumes in an imaging sub-cohort.
Materials and Methods
Participants
The subjects were from the Beijing Aging Brain Rejuvenation Initiative (BABRI) Study Group, which is an ongoing longitudinal study examining the brain and cognitive decline in an elderly, community-dwelling population [40]. All enrolled participants were Han Chinese. Participants were qualified for our study if they met the following criteria: right-handed and native Chinese speakers, no less than 50 years old, at least 6 years of education, no history of neurological or psychiatric disorders, and could provide a successful blood sample for the genotyping analysis. For this PICALM investigation, we only included individuals who were “clinically non-demented,” as determined by using the DSM IV, Petersen’s dementia criteria, and Clinical Dementia Rating (score = 0). Accordingly, a total of 638 subjects (aged 50–82 years) were included in the present study. To investigate the age modulation effects, the subjects were divided into young (<65 years old) and old (≥65 years old) groups. The demographic information for each group is presented in Table 1. The study was approved by the Institutional Review Board of the Beijing Normal University Imaging Center for Brain Research. Written informed consent was obtained from each participant.
Neuropsychological Testing
All the participants were subjected to a battery of neuropsychological tests that assessed several cognitive domains. As mentioned previously, the comprehensive neuropsychological battery comprised the following six cognition domains (the tests used to assess each domain are in parentheses): 1, general mental status (the Mini-Mental Status Examination—Chinese version (MMSE) [41]); 2, memory function (the Auditory Verbal Learning Test (AVLT) [42], the Rey-Osterrieth Complex Figure test (ROCF) (recall) [43], and the Digit Span test, which was a sub-test of the Wechsler Adult Intelligence Scale—Chinese revision); 3, spatial processing (ROCF-copy [43] and the Clock-Drawing Test (CDT) [44]); 4, language (the Category Verbal Fluency Test (CVFT) and the Boston Naming Test (BNT) [45]); 5, processing speed (the Trail Making Test (TMT) A [46] and the Symbol Digit Modalities Test (SDMT) [47]); and 6, executive function (the TMT-B [46] and the Stroop Color and Word Test C (SCWT) [48]).
Analysis of Genotyping
PICALM rs541458 was genotyped using TaqMan allele-specific assays on the 7900HT Fast Real-Time PCR System (Applied Biosystems, Foster City, CA, USA). Another two SNPs, rs429358 and rs7412, which collectively form the APOE ε2 (with the haplotype of rs429358-rs7412: T/T), APOE ε3 (G/T), and APOE ε4 alleles (G/G), were also genotyped. The sample success rate for all three SNPs were 100% (i.e., no failures across the participants to “call” the polymorphisms), and the reproducibility of all of the genotyping was 100% according to a duplicate analysis of 10% of the genotypes. Given the risk of harboring the T allele, we combined the CT and TT genotypes into T allele carriers. Thus, according to the rs541458 genotyping, all participants were divided into two groups: 151 CC and 487 T allele carriers.
MRI Data Acquisition
To investigate the interactive effects between age and PICALM genotypes on brain structure and function, we also acquired MRI data from a sub-cohort (n = 78) of the study participants using a SIEMENS TRIO 3T scanner in the Imaging Center for Brain Research, Beijing Normal University. The participants were placed in a supine position, with their head snugly fixed by straps and foam pads to minimize head movement. Resting state data were collected using a gradient echo EPI sequence [TE = 30 ms, TR = 2000 ms, flip angle = 90°, 33 slices, slice thickness = 4 mm, in-plane matrix = 64 × 64, field of view = 256 × 256 mm2]. During the single-run resting acquisition, the subjects were instructed to remain awake, relaxed, and with their eyes closed and to remain as motionless as possible. The resting acquisition lasted for 8 min, and 240 image volumes were obtained. T1-weighted structural images were acquired using 3D magnetization prepared rapid gradient echo (MP-RAGE) sequences [176 sagittal slices, TE = 3.44 ms, TR = 1900 ms, flip angle = 9°, slice thickness = 1 mm, acquisition matrix = 256 × 256, field of view = 256 × 256 mm2]. Table 2 provides further details of the imaging sub-sample.
Data Processing and Analysis
Resting state image preprocessing and analysis
For each participant, the first 10 volumes were discarded to allow the participants to adapt to the magnetic field. Functional data were preprocessed using SPM and DPARSF (http://rfmri.org/DPARSF), and the processing included slice timing, within-subject inter-scan realignment to correct for possible movement, spatial normalization to a standard brain template in the Montreal Neurological Institute (MNI) coordinate space, resampling to 3 × 3 × 3 mm3, and smoothing with an 8 mm full-width half-maximum Gaussian kernel. Three subjects were excluded because of unacceptable head movement (translation >3 mm or rotation >3°) during the fMRI scanning.
We then performed the independent component analysis (ICA) using the group ICA toolbox (GIFT version 2.0e; http://mialab.mrn.org/software/gift/). Twenty-five components were estimated for each subject. Three main stages were used when applying the ICA to all participants: (i) principal component analysis was performed for each subject for data reduction, (ii) application of the ICA algorithm, and (iii) back-reconstruction for each individual subject. After back-reconstruction, the mean spatial maps of each group at every time point were converted to z-scores for display. We focused on the DMN, left frontal-parietal network (FPN), and right FPN in the current study. The best-fit components for the three resting state networks were identified by visual inspection. For each network, a full factorial analysis of covariance (ANCOVA) (2 × 2) was conducted with PICALM rs541458 (T carriers versus CC genotype) and age (<65 years old versus ≥65 years old) as independent factors in SPM 8 (Statistical Parametric Mapping, www.fil.ion.ucl.ac.uk/spm) with gender, years of education, and APOE ε4 status included as covariates (p < 0.05, AlphaSim-corrected). Additionally, further analysis of the correlation of imaging measures with neuropsychological tests was performed on any significant clusters resulting from the voxel-wise comparisons. For each significant cluster, the connectivity values were extracted by averaging the intensities over all voxels within the cluster from every participant’s component map.
Structural Image Analysis
We used a voxel-based morphometry (VBM) analysis to compare the whole-brain gray matter volume of all subjects using the VBM8 software package (http://dbm.neuro.uni-jena.de/vbm/). The preprocessing of structural images used default parameters, except for the estimation using “ICBM space template—East Asian brains” and extended options using “thorough cleanup.” The images were bias-corrected, tissue classified, and normalized to Montreal Neurological Institute space using affine and non-linear transformations to compare the absolute amounts of tissue [49]. One subject with excessive head motion was identified and excluded. The images of the remaining 77 subjects were free of such problems. The modulated gray matter images were smoothed with a Gaussian kernel of 8 mm full width at half maximum (FWHM). We utilized the mean gray matter map (threshold = 0.3) of all subjects to obtain a group-based brain mask and used it for subsequent analysis. A full factorial ANCOVA (2 × 2) was calculated using SPM8 with PICALM rs541458 (T carriers versus CC genotype) and age (<65 years old versus ≥65 years old) as independent factors with gender, years of education, APOE ε4 status, and total intracranial volume included as covariates (p < 0.001, AlphaSim-corrected). The significant areas for the interaction of rs541458 × age in the VBM analysis were extracted as regions of interest (ROI) using REST v1.8 (http://www.restfmri.net).
Statistical Analysis
The Hardy-Weinberg test was completed using the PLINK software [50]. For the demographic factors of gender and years of education and the APOE ε4 status, the multivariable analysis of variance or Wald chi-square test was used to assess the PICALM rs541458 polymorphism and age effects. For the neuropsychological assessments, a multivariable analysis of covariance was conducted, with gender, years of education, and APOE ε4 status as covariates. In addition, for the functional connectivity, gray matter volume, and cognitive performance showing the significant interactive effects of rs541458 × age, we further calculated the correlation between them using Pearson partial correlation analyses, after controlling for the influences of gender, years of education, and APOE ε4 status in the four genotype and age groups, separately.
Results
Demographic and Neuropsychological Results
The rs541458 SNP did not show any deviations from Hardy-Weinberg equilibrium in all participants (P > 0.05). For all participants, the differences between the rs541458 groups and the rs541458 by age interactions were not significant for demographic factors or APOE ε4 status (Table 1). These results were the same in the imaging sub-sample (Table 2). All subsequent analyses were adjusted for gender, years of education, and APOE ε4 status.
For the neuropsychological tests, the interactions of rs541458 × age were significant for executive function (SCWT-C, P = 0.008, TMTb, P = 0.008, and TMTb-a, P = 0.029) and processing speed (TMTa, P = 0.028) in all participants. The effects of age, but not rs541458, were significant for all neuropsychological tests in all participants (Table 1). We found similar results in the imaging sub-sample, as the interactions of rs541458 × age were significant for executive function (SCWT C-B, P = 0.034, SCWT-C, P = 0.024, TMTb, P = 0.003, and TMTb-a, P = 0.011), processing speed (SDMT, P = 0.011), and general mental status (MMSE, P = 0.013). The age effects were significant for executive function (TMTb, P = 0.002, and TMTb-a, P = 0.001) and memory function (ROCF-delay, P = 0.038). The rs541458 effects were significant for general mental status (MMSE, P = 0.040) (Table 2, Fig. S1).
Interactive Effect of rs541458 × age on Resting State Networks
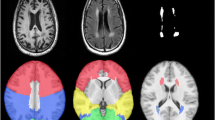
For the resting state networks, the DMN, left FPN, and right FPN were identified from the results of the group ICA (Fig. 1a). The full factorial analysis of covariance (ANCOVA) (2 × 2) revealed a significant interaction effect of rs541458 × age on the brain region connectivity of the left superior parietal gyrus (SPG.L) (x = −18 mm, y = −75 mm, z = 54 mm; voxel size = 73, P < 0.05, AlphaSim-corrected, Fig. 1b) within the left FPN. The simple effects of rs541458 and age were further analyzed (Fig. 1c). The functional connectivity of the SPG.L in the T allele carriers <65 years old group was significantly lower than that in the CC genotype <65 years old group (F = 8.56, P = 0.005). Further, the functional connectivity of the SPG.L in the CC genotype ≥65 years old group was significantly lower than that in the CC genotype <65 years old group (F = 9.35, P = 0.003).
a The spatial maps show the DMN, left FPN, and right FPN of the four genotype and age groups, separately. b The brain map represents voxel-wise interactive effect of rs541458 × age on the left FPN (SPG.L x = −18 mm, y = −75 mm, z = 54 mm; voxel size = 73, P < 0.05, AlphaSim-corrected). The x, y, z coordinates of the primary peak in MNI space. c The bar graph shows the ROI analysis on the significant regions from voxel-wise comparisons. Error bars denote the standard error of the mean. *Significant at P < 0.05. DMN default mode network, FPN frontal-parietal network, SPG.L left superior parietal gyrus, ROI region of interest
Interactive Effect of rs541458 × age on Gray Matter Volumes
We found a significant interaction effect of rs541458 × age on the gray matter volume of the left middle temporal gyrus (MTG.L) (x = −50 mm, y = −51 mm, z = 4 mm; voxel size = 3032, P < 0.001, AlphaSim-corrected, Fig. 2a). The simple effects of rs541458 and age were further analyzed (Fig. 2b). The gray matter volume of the MTG.L in the T allele carriers <65 years old group was significantly higher than that in the T allele carriers ≥65 years old group (F = 11.70, P = 0.001). And the gray matter volume of the MTG.L in the CC genotype ≥65 years old group was significantly higher than that in the T allele carriers ≥65 years old group (F = 8.69, P = 0.004).
a The brain map represents voxel-wise interactive effect of rs541458 × age on the gray matter volumes (MTG.L x = −50 mm, y = −51 mm, z = 4 mm; voxel size = 3032, P < 0.001, AlphaSim-corrected). The x, y, z coordinates of the primary peak in MNI space. b The bar graph shows the ROI analysis on the significant regions from voxel-wise comparisons. Error bars denote the standard error of the mean. *Significant at P < 0.05. MTG.L left middle temporal gyrus, ROI region of interest
Correlations Among Functional Connectivity, Gray Matter Volume, and Cognitive Performance
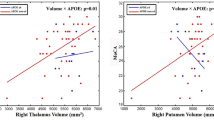
To assess whether the functional network and gray matter volume differences could explain the cognitive performance differences, we correlated the connectivity of the SPG.L within the left FPN and the gray matter volume of the MTG.L each with the cognitive test score, which also showed significant rs541458 × age interactive effects. For the functional network, the T allele carriers <65 years old group showed a significant correlation between TMTb-a and SPG.L connectivity (r = 0.460, P = 0.047) (Fig. 3). For the gray matter volume, the TMTb was negatively associated with MTG.L volume in the CC genotype <65 years old group (r = −0.588, P = 0.017), and SCWT was negatively associated with MTG.L volume in the CC genotype ≥65 years old group (SCWT C-B, r = −0.607, P = 0.028, SCWT-C, r = −0.635, P = 0.020) (Fig. 3). No significant correlations were found in the other groups or for the other cognitive performance measures. Furthermore, the CC genotype ≥65 years old group showed a negative correlation between SPG.L connectivity and MTG.L volume (r = −0.652, P = 0.022) (Fig. 3). Notably, there were no correlation results that survived (P < 0.05) after FDR correction for multiple comparisons.
Correlations between the functional connectivity of SPG.L, the gray matter volume of MTG.L, and the neuropsychological tests in the four genotype and age groups, separately. The significant correlation showed between the connectivity of SPG.L and the TMTb-a time in T allele carriers <65 years old group, between the volume of MTG.L and the TMTb time in CC genotype <65 years old group, between the volume of MTG.L and the SCWT C-B and SCWT-C time in the CC genotype ≥65 years old group, and between the connectivity of SPG.L and the volume of MTG.L in the CC genotype ≥65 years old group. SPG.L left superior parietal gyrus, MTG.L left middle temporal gyrus, TMT Trail Making Test, SCWT Stroop Color and Word Test
Discussion
The findings from the present study showed that age modulated the effects of PICALM on cognitive performance, which were mainly manifested in the executive function and processing speed, that is, the risk allele T was associated with better cognitive performance in the <65 years old group, but the opposite was found in the ≥65 years old group. The effects of PICALM on resting state functional networks were also modulated by age. We found that the risk allele T was associated with decreased functional connectivity of the SPG.L within the left FPN in the <65 years old group, whereas the T allele was associated with increased connectivity in the ≥65 years old group. Age also showed a modulatory effect on the association between PICALM and the gray matter volume of the MTG.L. In further correlation analyses, the functional connectivity of the SPG.L was positively correlated with TMTb-a time in the T allele carriers <65 years old group, which indicated that lower functional connectivity was linked to better executive function. Both CC genotype groups showed that an increased volume of the MTG.L was significantly related to better executive function.
First, we found that the age difference could affect the relations between the rs541458 T allele and both executive function and processing speed. With the exception observed in the study by Chen et al. [21] for Asian ethnic groups, several previous studies reported the T allele of rs541458 as a risk factor for LOAD in westerners [18–20]. However, few studies have directly investigated the association between rs541458 and cognitive functions, and none have been conducted on the rs541458 and age interaction. Indirectly, limited evidence has demonstrated that the rs541458 T allele would accelerate the decreases of general cognitive functions, thus suggesting the influence of age on the association between rs541458 and cognitive functions [26]. Implicitly, this was similar to our results, which indicated that age would modulate the effects of rs541458 on cognition. Interestingly, the risk G allele of another PICALM polymorphism, rs3851179, was associated with general cognition impairments in >70-year-old Parkinson’s disease patients but not in ≤70-year-old patients [51]. Further, the protective A allele of rs3851179 was associated with better cognitive composite scores only in men of the oldest elderly group [52]. These results together suggested that when the PICALM polymorphism is acting on cognition, other factors such as age would play regulatory effects. To further investigate the PICALM by age interaction, we further evaluated both the brain functional networks and its structure.
Ample data suggest that the FPN is sensitive to damage in mild cognitive impairment (MCI), which is a precursor phase of AD [53, 54]. A previous study of the APOE genotype found an intrinsic effect of APOE on the functional architecture of FPN [36]. In our study, the association between rs541458 and the functional connectivity of the left FPN was affected by age, and the pattern was contrary to that found for cognitive functions. One possible explanation for this difference is a compensation mechanism of functional networks. It is noteworthy that the FPN is essential for higher cognitive behaviors and that the decreases of effective connectivity in frontal-parietal circuits made for impairments in top-down attention control [55]. Recent studies have revealed the PICALM rs3851179 and clusterin/apolipoprotein J protein gene (CLU) interactions on functional connectivity, verified both in Caucasian and Chinese subjects [37, 56]. Nevertheless, the effects of rs541458 on functional networks and the role of age are still poorly understood. Pooling those previous studies and our current results would suggest, to some extent, that the effects of PICALM on brain functions are influenced not only by other genetic factors but also by age. Further, Wang et al. reported a decreased functional connectivity between the superior parietal cortex and thalamus in MCI patients [57], which may suggest the sensitivity of connectivity in the SPG for cognitive impairments.
Consistent with our brain connection findings, we also found that the effects of rs541458 on gray matter volume of the MTG.L were modulated by age. The MTG was among the first regions to show structural changes in amnestic MCI [58] and AD patients [59]. Compared with non-converters, MCI converters showed greater gray matter losses in the MTG area [60]. This may result from elevated tau pathology and neuronal loss [61]. In an elderly European group, the risk G allele of rs3851179 was related to a smaller gray matter volume of the inferior frontal gyrus compared to the protective A allele in APOE ε4 allele carriers, but this was not observed in non-carriers [39]. Similarly, CLU could affect the association between rs3851179 and hippocampal volume; that is, reversed patterns were shown in the CLU risk and protective groups [56]. This was consistent with our results, which showed that the risk allele T was associated with a higher gray matter volume of the MTG.L in the <65 years old group and with a lower volume in the ≥65 years old group.
Interestingly, the interactive effects of rs541458 × age on resting state functional networks and gray matter volume were both located on the left hemisphere, probably suggesting that lateralization were associated with aging and dementia. A recent research showed that age is strongly related to lateralization in multiple regions within the frontal network, attentional network, sensorimotor network, and visual network [62]. Early research has suggested that some neurodegenerative disease like dementia exhibited injury primarily in the left rather than the right hemisphere, and hypometabolism is more susceptible to neurodegeneration in the left hemisphere [63]. Some studies also demonstrated that brain asymmetry can be observed at biochemical level like hippocampal nitric oxide mediator system [64].
A growing body of research has demonstrated that particular cognitive impairments are closely related to the disconnection of brain networks. This has been verified in several neurodegenerative diseases, including AD [65] and frontotemporal dementia [66]. TMT is a frequently used neuropsychological test for executive function [46]. The executive function depended on disrupted regions to collaborate, mainly including the left SPG and lateral prefrontal cortex [67]. In MCI patients, the white matter hyperintensities in the FPN were associated with decreased executive function [68]. Our study found that the functional connectivity of the SPG.L was positively related with TMTb-a time scores in T allele carriers <65 years old, which probably suggests that better executive function performance did not need higher connectivity for support. Similarly, recent results have shown that increased connectivity between the left parietal and middle temporal cortices was associated with decreased global cognitive status in cognitively normal elderly [69]. The connectivity between the left inferior parietal and medial prefrontal cortex was related to episodic memory performance, which has also been found in cognitively normal older adults [70]. Furthermore, we found that the increased volume of the MTG.L was related to better executive function in both CC genotype groups. In the APOE ε4 carriers, the apparent diffusion coefficient of the temporal lobe could predict executive function [71]. In probable AD patients, decreased metabolism in the temporal cortex was correlated with poor performance on executive functions [72]. Additionally, a previous study indicated strong correlations between medial temporal lobe atrophy and executive functions in non-demented elderly [73]. As corollaries, some brain function- and structure-related measures, such as the functional connectivity of resting state networks and gray matter volumes, could be biomarkers for predicting variants in cognitive functions.
As mentioned above, abnormal PICALM expression would disturb APP metabolism [74] due to the crucial role of PICALM in clathrin-mediated endocytosis [12]. PICALM would promote the transportation of APP-cleaved C-terminal fragment (APP-CTF) from the plasma membrane intracellularly to allow fusion with autophagosomes and endosomes, thus increasing the degradation of APP-CTF and decreasing the production of amyloid-β (Aβ) [75]. Meanwhile, the PICALM rs541458 T allele is associated with decreased cerebrospinal fluid Aβ42 [76]. Considering its high expression in endothelial cells [77], this finding may imply that PICALM plays a major role in removing Aβ from the brain through the blood-brain barrier, supported by a close relationship between high levels of PICALM and increased Aβ clearance [78]. Moreover, the expression of PICALM was reduced and co-localized with hyperphosphorylated tau in AD patients [79], possibly indicating a role of PICALM in tauopathy [80]. Thus, one might speculate that an abnormal level of PICALM would contribute to the dysfunction of endocytosis and a series of relevant pathological changes related to AD. However, the exact mechanisms by which rs541458 contributes to AD etiology are yet to be confirmed in animal and human studies.
The present study had some limitations. First, the effects of different social class or level of educational achievement should be noted that they may have different effects on cognitive function and brain. Second, it is very important to validate the present results by other Chinese cohort and some longitudinal studies. Third, the significant correlations between neuroimaging measurements and cognitive performances reported in the present study should be regarded as exploratory in nature due to no correlation survived (P < 0.05) after FDR correction for multiple comparisons. Thus, they need to be confirmed in future additional studies. Overall, the findings of this study should be interpreted with these limitations in mind.
In summary, the present study suggested the modulation of age on the association of the PICALM rs541458 polymorphism with executive function, on its association with the left FPN and the MTG.L volume, and on the associations between neuropsychological performance and brain connection/structure in non-demented Chinese elderly. This finding highlights the importance of combining age and genetic polymorphisms when examining candidate genes that affect cognitive function. Further studies with a larger sample size and longitudinal design are needed to confirm our results.
References
Ferri CP, Prince M, Brayne C, Brodaty H, Fratiglioni L, Ganguli M, Hall K, Hasegawa K et al (2005) Global prevalence of dementia: a Delphi consensus study. Lancet 366(9503):2112–2117. doi:10.1016/S0140-6736(05)67889-0
Weuve J, Hebert LE, Scherr PA, Evans DA (2014) Deaths in the United States among persons with Alzheimer’s disease (2010-2050). Alzheimers Dement 10(2):e40–e46. doi:10.1016/j.jalz.2014.01.004
Kukull WA, Higdon R, Bowen JD, McCormick WC, Teri L, Schellenberg GD, van Belle G, Jolley L et al (2002) Dementia and Alzheimer disease incidence: a prospective cohort study. Arch Neurol 59(11):1737–1746
Niu H, Alvarez-Alvarez I, Guillen-Grima F, Aguinaga-Ontoso I (2016) Prevalence and incidence of Alzheimer’s disease in Europe: a meta-analysis. Neurologia 26(16):30003–30002
Fratiglioni L, Launer LJ, Andersen K, Breteler MM, Copeland JR, Dartigues JF, Lobo A, Martinez-Lage J et al (2000) Incidence of dementia and major subtypes in Europe: a collaborative study of population-based cohorts. Neurologic Diseases in the Elderly Research Group. Neurology 54(11 Suppl 5):S10–S15
Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, DeStafano AL, Bis JC et al (2013) Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet 45(12):1452–1458
Corder EH, Saunders AM, Strittmatter WJ, Schmechel DE, Gaskell PC, Small GW, Roses AD, Haines JL et al (1993) Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer’s disease in late onset families. Science 261(5123):921–923
Ashford JW, Mortimer JA (2002) Non-familial Alzheimer’s disease is mainly due to genetic factors. J Alzheimers Dis 4(3):169–177
Mayeux R, Saunders AM, Shea S, Mirra S, Evans D, Roses AD, Hyman BT, Crain B et al (1998) Utility of the apolipoprotein E genotype in the diagnosis of Alzheimer’s disease. Alzheimer’s disease centers consortium on apolipoprotein E and Alzheimer’s disease. N Engl J Med 338(8):506–511. doi:10.1056/NEJM199802193380804
Harold D, Abraham R, Hollingworth P, Sims R, Gerrish A, Hamshere ML, Pahwa JS, Moskvina V et al (2009) Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease. Nat Genet 41(10):1088–1093
Yao PJ, Zhang P, Mattson MP, Furukawa K (2003) Heterogeneity of endocytic proteins: distribution of clathrin adaptor proteins in neurons and glia. Neuroscience 121(1):25–37
Ford MG, Pearse BM, Higgins MK, Vallis Y, Owen DJ, Gibson A, Hopkins CR, Evans PR et al (2001) Simultaneous binding of PtdIns(4,5)P2 and clathrin by AP180 in the nucleation of clathrin lattices on membranes. Science 291(5506):1051–1055. doi:10.1126/science.291.5506.1051
Marsh M, McMahon HT (1999) The structural era of endocytosis. Science 285(5425):215–220
Cirrito JR, Kang JE, Lee J, Stewart FR, Verges DK, Silverio LM, Bu G, Mennerick S et al (2008) Endocytosis is required for synaptic activity-dependent release of amyloid-beta in vivo. Neuron 58(1):42–51. doi:10.1016/j.neuron.2008.02.003
Kyriazis GA, Wei Z, Vandermey M, Jo DG, Xin O, Mattson MP, Chan SL (2008) Numb endocytic adapter proteins regulate the transport and processing of the amyloid precursor protein in an isoform-dependent manner: implications for Alzheimer disease pathogenesis. J Biol Chem 283(37):25492–25502. doi:10.1074/jbc.M802072200
Holton P, Ryten M, Nalls M, Trabzuni D, Weale ME, Hernandez D, Crehan H, Gibbs JR et al (2013) Initial assessment of the pathogenic mechanisms of the recently identified Alzheimer risk loci. Ann Hum Genet 77(2):85–105. doi:10.1111/ahg.12000
Lambert JC, Heath S, Even G, Campion D, Sleegers K, Hiltunen M, Combarros O, Zelenika D et al (2009) Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer’s disease. Nat Genet 41(10):1094–1099
Lambert JC, Zelenika D, Hiltunen M, Chouraki V, Combarros O, Bullido MJ, Tognoni G, Fievet N et al (2011) Evidence of the association of BIN1 and PICALM with the AD risk in contrasting European populations. Neurobiol Aging 32(4):8
Corneveaux JJ, Myers AJ, Allen AN, Pruzin JJ, Ramirez M, Engel A, Nalls MA, Chen K et al (2010) Association of CR1, CLU and PICALM with Alzheimer’s disease in a cohort of clinically characterized and neuropathologically verified individuals. Hum Mol Genet 19(16):3295–3301
Jun G, Naj AC, Beecham GW, Wang LS, Buros J, Gallins PJ, Buxbaum JD, Ertekin-Taner N et al (2010) Meta-analysis confirms CR1, CLU, and PICALM as alzheimer disease risk loci and reveals interactions with APOE genotypes. Arch Neurol 67(12):1473–1484
Chen LH, Kao PYP, Fan YH, Ho DTY, Chan CSY, Yik PY, Ha JCT, Chu LW et al (2012) Polymorphisms of CR1, CLU and PICALM confer susceptibility of Alzheimer’s disease in a southern Chinese population. Neurobiol Aging 33(1):210.e211–210.e217. doi:10.1016/j.neurobiolaging.2011.09.016
Bishop NA, Lu T, Yankner BA (2010) Neural mechanisms of ageing and cognitive decline. Nature 464(7288):529–535. doi:10.1038/nature08983
Fjell AM, Westlye LT, Grydeland H, Amlien I, Espeseth T, Reinvang I, Raz N, Dale AM et al (2014) Accelerating cortical thinning: unique to dementia or universal in aging? Cereb Cortex 24(4):919–934. doi:10.1093/cercor/bhs379
Kurth S, Majerus S, Bastin C, Collette F, Jaspar M, Bahri MA, Salmon E (2016) Effects of aging on task- and stimulus-related cerebral attention networks. Neurobiol Aging 44:85–95
Buckner RL (2004) Memory and executive function in aging and AD: multiple factors that cause decline and reserve factors that compensate. Neuron 44(1):195–208. doi:10.1016/j.neuron.2004.09.006
Sweet RA, Seltman H, Emanuel JE, Lopez OL, Becker JT, Bis JC, Weamer EA, DeMichele-Sweet MA et al (2012) Effect of Alzheimer’s disease risk genes on trajectories of cognitive function in the Cardiovascular Health Study. Am J Psychiatry 169(9):954–962
Koedam EL, Lauffer V, van der Vlies AE, van der Flier WM, Scheltens P, Pijnenburg YA (2010) Early-versus late-onset Alzheimer’s disease: more than age alone. J Alzheimers Dis 19(4):1401–1408
Ma C, Zhang Y, Li X, Zhang J, Chen K, Liang Y, Chen Y, Liu Z et al (2016) Is there a significant interaction effect between apolipoprotein E rs405509 T/T and epsilon4 genotypes on cognitive impairment and gray matter volume? Eur J Neurol. doi:10.1111/ene.13052
Wang P, Zhou B, Yao H, Zhan Y, Zhang Z, Cui Y, Xu K, Ma J et al (2015) Aberrant intra- and inter-network connectivity architectures in Alzheimer’s disease and mild cognitive impairment. Sci Rep 5(14824)
Damoiseaux JS, Rombouts SA, Barkhof F, Scheltens P, Stam CJ, Smith SM, Beckmann CF (2006) Consistent resting-state networks across healthy subjects. Proc Natl Acad Sci U S A 103(37):13848–13853. doi:10.1073/pnas.0601417103
Balsters JH, O’Connell RG, Galli A, Nolan H, Greco E, Kilcullen SM, Bokde AL, Lai R et al (2013) Changes in resting connectivity with age: a simultaneous electroencephalogram and functional magnetic resonance imaging investigation. Neurobiol Aging 34(9):2194–2207
Damoiseaux JS, Beckmann CF, Arigita EJ, Barkhof F, Scheltens P, Stam CJ, Smith SM, Rombouts SA (2008) Reduced resting-state brain activity in the “default network” in normal aging. Cereb Cortex 18(8):1856–1864. doi:10.1093/cercor/bhm207
Machulda MM, Jones DT, Vemuri P, McDade E, Avula R, Przybelski S, Boeve BF, Knopman DS et al (2011) Effect of APOE epsilon4 status on intrinsic network connectivity in cognitively normal elderly subjects. Arch Neurol 68(9):1131–1136. doi:10.1001/archneurol.2011.108
Sheline YI, Morris JC, Snyder AZ, Price JL, Yan Z, D’Angelo G, Liu C, Dixit S et al (2010) APOE4 allele disrupts resting state fMRI connectivity in the absence of amyloid plaques or decreased CSF Abeta42. J Neurosci 30(50):17035–17040. doi:10.1523/JNEUROSCI.3987-10.2010
Damoiseaux JS, Seeley WW, Zhou J, Shirer WR, Coppola G, Karydas A, Rosen HJ, Miller BL et al (2012) Gender modulates the APOE epsilon4 effect in healthy older adults: convergent evidence from functional brain connectivity and spinal fluid tau levels. J Neurosci 32(24):8254–8262. doi:10.1523/JNEUROSCI.0305-12.2012
Trachtenberg AJ, Filippini N, Ebmeier KP, Smith SM, Karpe F, Mackay CE (2012) The effects of APOE on the functional architecture of the resting brain. NeuroImage 59(1):565–572
Zhang P, Qin W, Wang D, Liu B, Zhang Y, Jiang T, Yu C (2015) Impacts of PICALM and CLU variants associated with Alzheimer’s disease on the functional connectivity of the hippocampus in healthy young adults. Brain Struct Funct 220(3):1463–1475. doi:10.1007/s00429-014-0738-4
Espeseth T, Westlye LT, Fjell AM, Walhovd KB, Rootwelt H, Reinvang I (2008) Accelerated age-related cortical thinning in healthy carriers of apolipoprotein E epsilon 4. Neurobiol Aging 29(3):329–340. doi:10.1016/j.neurobiolaging.2006.10.030
Morgen K, Ramirez A, Frolich L, Tost H, Plichta MM, Kolsch H, Rakebrandt F, Rienhoff O et al (2014) Genetic interaction of PICALM and APOE is associated with brain atrophy and cognitive impairment in Alzheimer’s disease. Alzheimers Dement 10(5 Suppl):S269–S276. doi:10.1016/j.jalz.2013.11.001
Li X, Ma C, Zhang J, Liang Y, Chen Y, Chen K, Wang J, Zhang Z et al (2013) Prevalence of and potential risk factors for mild cognitive impairment in community-dwelling residents of Beijing. J Am Geriatr Soc 61(12):2111–2119
Zhang M, Katzman R, Salmon D, Jin H, Cai G, Wang Z, Qu G, Grant I et al (1990) The prevalence of dementia and Alzheimer’s disease in Shanghai, China: impact of age, gender, and education. Ann Neurol 27(4):428–437
Rosenberg SJ, Ryan JJ, Prifitera A (1984) Rey auditory-verbal learning test performance of patients with and without memory impairment. J Clin Psychol 40(3):785–787
Rey A (1941) L-examen psychologique dans les cas d’encephalopathie traumatique. Arch Psychologie 1941(28):286–340
Rouleau I, Salmon DP, Butters N, Kennedy C, McGuire K (1992) Quantitative and qualitative analyses of clock drawings in Alzheimer’s and Huntington’s disease. Brain Cogn 18(1):70–87
Guo Q (2006) Boston Naming Test in Chinese elderly, patient with mild cognitive impairment and Alzheimer’s dementia. Chin Ment Health J 20(2):81
Reitan R (1958) The validity of the trail making test as an indicator of organic brain damage. Percept Mot Skills 8:271–276
Sheridan LK, Fitzgerald HE, Adams KM, Nigg JT, Martel MM, Puttler LI, Wong MM, Zucker RA (2006) Normative symbol digit modalities test performance in a community-based sample. Arch Clin Neuropsychol 21(1):23–28. doi:10.1016/j.acn.2005.07.003
Guo Q, Hong Z, Lv C, Zhou Y, Lu J, Ding D (2005) Application of Stroop color-word test on Chinese elderly patients with mild cognitive impairment and mild Alzheimer’s dementia. Chinese Journal of Neuromedicine 4(7):701–704
Good CD, Johnsrude IS, Ashburner J, Henson RN, Friston KJ, Frackowiak RS (2001) A voxel-based morphometric study of ageing in 465 normal adult human brains. NeuroImage 14(1 Pt 1):21–36. doi:10.1006/nimg.2001.0786
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81(3):559–575
Barrett MJ, Koeppel AF, Flanigan JL, Turner SD, Worrall BB (2016) Investigation of genetic variants associated with Alzheimer disease in Parkinson disease cognition. J Parkinsons Dis 6(1):119–124
Mengel-From J, Christensen K, McGue M, Christiansen L (2011) Genetic variations in the CLU and PICALM genes are associated with cognitive function in the oldest old. Neurobiol Aging 32(3):23
Qi Z, Wu X, Wang Z, Zhang N, Dong H, Yao L, Li K (2010) Impairment and compensation coexist in amnestic MCI default mode network. NeuroImage 50(1):48–55
Xie C, Bai F, Yu H, Shi Y, Yuan Y, Chen G, Li W, Zhang Z et al (2012) Abnormal insula functional network is associated with episodic memory decline in amnestic mild cognitive impairment. NeuroImage 63(1):320–327
Neufang S, Akhrif A, Riedl V, Forstl H, Kurz A, Zimmer C, Sorg C, Wohlschlager AM (2011) Disconnection of frontal and parietal areas contributes to impaired attention in very early Alzheimer’s disease. J Alzheimers Dis 25(2):309–321
Yang X, Li J, Liu B, Li Y, Jiang T (2016) Impact of PICALM and CLU on hippocampal degeneration. Hum Brain Mapp 37(7):2419–2430. doi:10.1002/hbm.23183
Wang Z, Jia X, Liang P, Qi Z, Yang Y, Zhou W, Li K (2012) Changes in thalamus connectivity in mild cognitive impairment: evidence from resting state fMRI. Eur J Radiol 81(2):277–285
Zhao ZL, Fan FM, Lu J, Li HJ, Jia LF, Han Y, Li KC (2015) Changes of gray matter volume and amplitude of low-frequency oscillations in amnestic MCI: an integrative multi-modal MRI study. Acta Radiol 56(5):614–621. doi:10.1177/0284185114533329
Busatto GF, Garrido GE, Almeida OP, Castro CC, Camargo CH, Cid CG, Buchpiguel CA, Furuie S et al (2003) A voxel-based morphometry study of temporal lobe gray matter reductions in Alzheimer’s disease. Neurobiol Aging 24(2):221–231
Chetelat G, Landeau B, Eustache F, Mezenge F, Viader F, de la Sayette V, Desgranges B, Baron JC (2005) Using voxel-based morphometry to map the structural changes associated with rapid conversion in MCI: a longitudinal MRI study. NeuroImage 27(4):934–946
Ishiki A, Okamura N, Furukawa K, Furumoto S, Harada R, Tomita N, Hiraoka K, Watanuki S et al (2015) Longitudinal assessment of tau pathology in patients with Alzheimer’s disease using [18F]THK-5117 positron emission tomography. PLoS One 10(10):e0140311. doi:10.1371/journal.pone.0140311
Agcaoglu O, Miller R, Mayer AR, Hugdahl K, Calhoun VD (2015) Lateralization of resting state networks and relationship to age and gender. NeuroImage 104:310–325. doi:10.1016/j.neuroimage.2014.09.001
Loewenstein DA, Barker WW, Chang JY, Apicella A, Yoshii F, Kothari P, Levin B, Duara R (1989) Predominant left hemisphere metabolic dysfunction in dementia. Arch Neurol 46(2):146–152
Kristofikova Z, Kozmikova I, Hovorkova P, Ricny J, Zach P, Majer E, Klaschka J, Ripova D (2008) Lateralization of hippocampal nitric oxide mediator system in people with Alzheimer disease, multi-infarct dementia and schizophrenia. Neurochem Int 53(5):118–125
Dai Z, Yan C, Li K, Wang Z, Wang J, Cao M, Lin Q, Shu N et al (2014) Identifying and mapping connectivity patterns of brain network hubs in Alzheimer’s disease. Cereb Cortex. doi:10.1093/cercor/bhu246
Whitwell JL, Josephs KA, Avula R, Tosakulwong N, Weigand SD, Senjem ML, Vemuri P, Jones DT et al (2011) Altered functional connectivity in asymptomatic MAPT subjects: a comparison to bvFTD. Neurology 77(9):866–874. doi:10.1212/WNL.0b013e31822c61f2
Collette F, Van der Linden M, Laureys S, Delfiore G, Degueldre C, Luxen A, Salmon E (2005) Exploring the unity and diversity of the neural substrates of executive functioning. Hum Brain Mapp 25(4):409–423
Jacobs HI, Visser PJ, Van Boxtel MP, Frisoni GB, Tsolaki M, Papapostolou P, Nobili F, Wahlund LO et al (2012) Association between white matter hyperintensities and executive decline in mild cognitive impairment is network dependent. Neurobiol Aging 33(1):23
Jiang Y, Huang H, Abner E, Broster LS, Jicha GA, Schmitt FA, Kryscio R, Andersen A et al (2016) Alzheimer’s biomarkers are correlated with brain connectivity in older adults differentially during resting and task states. Front Aging Neurosci 8(15)
He J, Carmichael O, Fletcher E, Singh B, Iosif AM, Martinez O, Reed B, Yonelinas A et al (2012) Influence of functional connectivity and structural MRI measures on episodic memory. Neurobiol Aging 33(11):2612–2620
Ryan L, Walther K, Bendlin BB, Lue LF, Walker DG, Glisky EL (2011) Age-related differences in white matter integrity and cognitive function are related to APOE status. NeuroImage 54(2):1565–1577
Woo BK, Harwood DG, Melrose RJ, Mandelkern MA, Campa OM, Walston A, Sultzer DL (2010) Executive deficits and regional brain metabolism in Alzheimer’s disease. Int J Geriatr Psychiatry 25(11):1150–1158
Oosterman JM, Vogels RL, van Harten B, Gouw AA, Scheltens P, Poggesi A, Weinstein HC, Scherder EJ (2008) The role of white matter hyperintensities and medial temporal lobe atrophy in age-related executive dysfunctioning. Brain Cogn 68(2):128–133
Xiao Q, Gil SC, Yan P, Wang Y, Han S, Gonzales E, Perez R, Cirrito JR et al (2012) Role of phosphatidylinositol clathrin assembly lymphoid-myeloid leukemia (PICALM) in intracellular amyloid precursor protein (APP) processing and amyloid plaque pathogenesis. J Biol Chem 287(25):21279–21289. doi:10.1074/jbc.M111.338376
Tian Y, Chang JC, Fan EY, Flajolet M, Greengard P (2013) Adaptor complex AP2/PICALM, through interaction with LC3, targets Alzheimer’s APP-CTF for terminal degradation via autophagy. Proc Natl Acad Sci U S A 110(42):17071–17076. doi:10.1073/pnas.1315110110
Schjeide BM, Schnack C, Lambert JC, Lill CM, Kirchheiner J, Tumani H, Otto M, Tanzi RE et al (2011) The role of clusterin, complement receptor 1, and phosphatidylinositol binding clathrin assembly protein in Alzheimer disease risk and cerebrospinal fluid biomarker levels. Arch Gen Psychiatry 68(2):207–213
Baig S, Joseph SA, Tayler H, Abraham R, Owen MJ, Williams J, Kehoe PG, Love S (2010) Distribution and expression of picalm in Alzheimer disease. J Neuropathol Exp Neurol 69(10):1071–1077. doi:10.1097/NEN.0b013e3181f52e01
Zhao Z, Sagare AP, Ma Q, Halliday MR, Kong P, Kisler K, Winkler EA, Ramanathan A et al (2015) Central role for PICALM in amyloid-beta blood-brain barrier transcytosis and clearance. Nat Neurosci 18(7):978–987
Ando K, Brion JP, Stygelbout V, Suain V, Authelet M, Dedecker R, Chanut A, Lacor P et al (2013) Clathrin adaptor CALM/PICALM is associated with neurofibrillary tangles and is cleaved in Alzheimer’s brains. Acta Neuropathol 125(6):861–878
Moreau K, Fleming A, Imarisio S, Lopez Ramirez A, Mercer JL, Jimenez-Sanchez M, Bento CF, Puri C, Zavodszky E, Siddiqi F, Lavau CP, Betton M, O’Kane CJ, Wechsler DS, Rubinsztein DC (2014) PICALM modulates autophagy activity and tau accumulation. Nat Commun 5 (4998). doi:10.1038/ncomms5998
Acknowledgements
We thank all the volunteers for their participation in the study and anonymous reviewers for their insightful comments and suggestions. This work was supported by the State Key Program of National Natural Science of China (grant number: 81430100), by Natural Science Foundation of China (grant number: 30873458 and 81173460), by National Science Fund for Distinguished Young Scholars (grant number: 81625025), and by Beijing Municipal Science & Technology Commission (grant number: Z161100000216135).
Author Contributions
Study concept and design: DP and ZZ. Acquisition, analysis, or interpretation of data: ZL, XD, JZ, XL, YC, CM, and KC. Drafting of the manuscript: ZL. Critical revision of the manuscript for important intellectual content: All authors. Obtained funding: ZZ. Study supervision: DP and ZZ. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The study was approved by the Institutional Review Board of the Beijing Normal University Imaging Center for Brain Research. Written informed consent was obtained from each participant.
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic supplementary material

Fig. S1
Bar graphs show significant interactions of rs541458×age in the neuropsychological assessments in the imaging sub-sample. Error bars denote the standard error of the mean. *Significant at P<0.05. MMSE, Mini-Mental State Examination; SDMT, Symbol Digit Modalities Test; TMT, Trail Making Test; SCWT, Stroop Color and Word Test. (GIF 77 kb)
High Resolution Image
(TIFF 15190 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Liu, Z., Dai, X., Zhang, J. et al. The Interactive Effects of Age and PICALM rs541458 Polymorphism on Cognitive Performance, Brain Structure, and Function in Non-demented Elderly. Mol Neurobiol 55, 1271–1283 (2018). https://doi.org/10.1007/s12035-016-0358-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-016-0358-5