Abstract
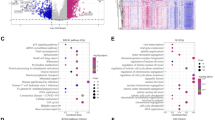
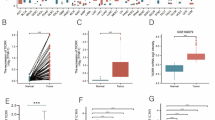
Hepatocellular carcinoma (HCC) is a common type of cancer that ranks first in cancer-associated death worldwide. Carbamoyl-phosphate synthetase 2, aspartate transcarbamylase, and dihydroorotase (CAD) are the key components of the pyrimidine pathway, which promotes cancer development. However, the function of CAD in HCC needs to be clarified. In this study, the clinical and transcriptome data of 424 TCGA-derived HCC cases were analyzed. The results demonstrated that high CAD expression was associated with poor prognosis in HCC patients. The effect of CAD on HCC was then investigated comprehensively using GO annotation analysis, KEGG enrichment analysis, Gene Set Enrichment Analysis (GSEA), and CIBERSORT algorithm. The results showed that CAD expression was correlated with immune checkpoint inhibitors and immune cell infiltration. In addition, low CAD levels in HCC patients predicted increased sensitivity to anti-CTLA4 and PD1, while HCC patients with high CAD expression exhibited high sensitivity to chemotherapeutic and molecular-targeted agents, including gemcitabine, paclitaxel, and sorafenib. Finally, the results from clinical sample suggested that CAD expression increased remarkably in HCC compared with non-cancerous tissues. Loss of function experiments demonstrated that CAD knockdown could significantly inhibit HCC cell growth and migration both in vitro and in vivo. Collectively, the results indicated that CAD is a potential oncogene during HCC metastasis and progression. Therefore, CAD is recommended as a candidate marker and target for HCC prediction and treatment.







Similar content being viewed by others
Data Availability
Please contact the corresponding author for all data requests.
References
Sung, H., Ferlay, J., Siegel, R. L., et al. (2021). global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians, 71(3), 209–249. https://doi.org/10.3322/caac.21660
Arnold, M., Abnet, C. C., Neale, R. E., et al. (2020). Global burden of 5 major types of gastrointestinal cancer. Gastroenterology, 159(1), 335.e15–349.e15. https://doi.org/10.1053/j.gastro.2020.02.068
Wang, K., Xiang, Y. J., Yu, H. M., et al. (2024). Adjuvant sintilimab in resected high-risk hepatocellular carcinoma: a randomized, controlled, phase 2 trial. Nature Medicine. Published online January 19, 2024. https://doi.org/10.1038/s41591-023-02786-7
Siegel, R. L., Miller, K. D., Fuchs, H. E., & Jemal, A. (2021). Cancer statistics, 2021. CA: A Cancer Journal for Clinicians, 71(1), 7–33. https://doi.org/10.3322/caac.21654
Chen, L., Zhang, C., Xue, R., et al. (2024). Deep whole-genome analysis of 494 hepatocellular carcinomas. Nature. Published online February 14, 2024. https://doi.org/10.1038/s41586-024-07054-3
Chen, W., Zheng, R., Baade, P. D., et al. (2016). Cancer statistics in China, 2015. CA: A Cancer Journal for Clinicians, 66(2), 115–132. https://doi.org/10.3322/caac.21338
Bruix, J., Reig, M., & Sherman, M. (2016). Evidence-based diagnosis, staging, and treatment of patients with hepatocellular carcinoma. Gastroenterology, 150(4), 835–853. https://doi.org/10.1053/j.gastro.2015.12.041
Vogel, A., Meyer, T., Sapisochin, G., Salem, R., & Saborowski, A. (2022). Hepatocellular carcinoma. The Lancet., 400(10360), 1345–1362. https://doi.org/10.1016/S0140-6736(22)01200-4
Arabi, F., Mansouri, V., & Ahmadbeigi, N. (2022). Gene therapy clinical trials, where do we go? An overview. Biomedicine & Pharmacotherapy, 153, 113324. https://doi.org/10.1016/j.biopha.2022.113324
Uddin, F., Rudin, C. M., & Sen, T. (2020). CRISPR gene therapy: Applications, limitations, and implications for the future. Frontiers in Oncology, 10, 1387. https://doi.org/10.3389/fonc.2020.01387
Bulaklak, K., & Gersbach, C. A. (2020). The once and future gene therapy. Nature Communications, 11(1), 5820. https://doi.org/10.1038/s41467-020-19505-2
Coleman, P. F., Suttle, D. P., & Stark, G. R. (1977). Purification from hamster cells of the multifunctional protein that initiates de novo synthesis of pyrimidine nucleotides. Journal of Biological Chemistry, 252(18), 6379–6385.
Lee, L., Kelly, R. E., Pastra-Landis, S. C., & Evans, D. R. (1985). Oligomeric structure of the multifunctional protein CAD that initiates pyrimidine biosynthesis in mammalian cells. Proceedings of the National Academy of Sciences U S A, 82(20), 6802–6806. https://doi.org/10.1073/pnas.82.20.6802
Sigoillot, F. D., Sigoillot, S. M., & Guy, H. I. (2004). Breakdown of the regulatory control of pyrimidine biosynthesis in human breast cancer cells. International Journal of Cancer, 109(4), 491–498. https://doi.org/10.1002/ijc.11717
Tlsty, T. D., Margolin, B. H., & Lum, K. (1989). Differences in the rates of gene amplification in nontumorigenic and tumorigenic cell lines as measured by Luria-Delbrück fluctuation analysis. Proc Natl Acad Sci U S A., 86(23), 9441–9445. https://doi.org/10.1073/pnas.86.23.9441
Khan, S., Abdelrahim, M., Samudio, I., & Safe, S. (2003). Estrogen receptor/Sp1 complexes are required for induction of cad gene expression by 17beta-estradiol in breast cancer cells. Endocrinology, 144(6), 2325–2335. https://doi.org/10.1210/en.2002-0149
Ridder, D. A., Schindeldecker, M., Weinmann, A., et al. (2021). Key enzymes in pyrimidine synthesis, CAD and CPS1, predict prognosis in hepatocellular carcinoma. Cancers (Basel)., 13(4), 744. https://doi.org/10.3390/cancers13040744
Wang, X., Yang, K., Wu, Q., et al. (2019). Targeting pyrimidine synthesis accentuates molecular therapy response in glioblastoma stem cells. Sci Transl Med., 11(504), eaau972. https://doi.org/10.1126/scitranslmed.aau4972
Lv, Y., Wang, X., Li, X., et al. (2020). Nucleotide de novo synthesis increases breast cancer stemness and metastasis via cGMP-PKG-MAPK signaling pathway. PLoS Biol, 18(11), e3000872. https://doi.org/10.1371/journal.pbio.3000872
Morin, A., Fritsch, L., Mathieu, J. R. R., et al. (2012). Identification of CAD as an androgen receptor interactant and an early marker of prostate tumor recurrence. The FASEB Journal, 26(1), 460–467. https://doi.org/10.1096/fj.11-191296
Xu, J. Y., Zhang, C., Wang, X., et al. (2020). Integrative proteomic characterization of human lung adenocarcinoma. Cell, 182(1), 245.e17–261.e17. https://doi.org/10.1016/j.cell.2020.05.043
Zhang ,Y., Liu, X., Liu, L., Li, J., Hu, Q., & Sun, R. (2020). Expression and prognostic significance of 6A-related genes in lung adenocarcinoma. Medical Science Monitor 26:e919644. https://doi.org/10.12659/MSM.919644
Liu, X. S., Zhou, L. M., Yuan, L. L., et al. (2021). NPM1 is a prognostic biomarker involved in immune infiltration of lung adenocarcinoma and associated with m6A modification and glycolysis. Frontiers in Immunology, 12, 724741. https://doi.org/10.3389/fimmu.2021.724741
Evans, D. R., & Guy, H. I. (2004). Mammalian pyrimidine biosynthesis: Fresh insights into an ancient pathway. Journal of Biological Chemistry, 279(32), 33035–33038. https://doi.org/10.1074/jbc.R400007200
Sigoillot, F. D., Kotsis, D. H., Masko, E. M., Bame, M., Evans, D. R., & Evans, H. I. G. (2007). Protein kinase C modulates the up-regulation of the pyrimidine biosynthetic complex, CAD, by MAP kinase. Frontiers in Bioscience, 12, 3892–3898. https://doi.org/10.2741/2358
Tu, H. F., Ko, C. J., Lee, C. T., et al. (2021). Afatinib exerts immunomodulatory effects by targeting the pyrimidine biosynthesis enzyme CAD. Cancer Research, 81(12), 3270–3282. https://doi.org/10.1158/0008-5472.CAN-20-3436
Del Caño-Ochoa, F., Moreno-Morcillo, M., & Ramón-Maiques, S. (2019). CAD, A multienzymatic protein at the head of de novo pyrimidine biosynthesis. SubCellular Biochemistry, 93, 505–538. https://doi.org/10.1007/978-3-030-28151-9_17
Novak, D. A., Carver, J. D., & Barness, L. A. (1994). Dietary nucleotides affect hepatic growth and composition in the weanling mouse. JPEN Journal of Parenteral and Enteral Nutrition, 18(1), 62–66. https://doi.org/10.1177/014860719401800162
Koch, J., Mayr, J. A., Alhaddad, B., et al. (2017). CAD mutations and uridine-responsive epileptic encephalopathy. Brain, 140(2), 279–286. https://doi.org/10.1093/brain/aww300
Yoshida, T., Stark, G. R., & Hoogenraad, J. (1974). Inhibition by N-(phosphonacetyl)-L-aspartate of aspartate transcarbamylase activity and drug-induced cell proliferation in mice. Journal of Biological Chemistry, 249(21), 6951–6955.
Clara, J. A., Monge, C., Yang, Y., & Takebe, N. (2020). Targeting signalling pathways and the immune microenvironment of cancer stem cells – A clinical update. Nature Reviews. Clinical Oncology, 17(4), 204–232. https://doi.org/10.1038/s41571-019-0293-2
Zanconato, F., Cordenonsi, M., & Piccolo, S. (2016). YAP/TAZ at the roots of cancer. Cancer Cell, 29(6), 783–803. https://doi.org/10.1016/j.ccell.2016.05.005
Wang, S., Zhang, Z., Qian, W., et al. (2018). Angiogenesis and vasculogenic mimicry are inhibited by 8-Br-cAMP through activation of the cAMP/PKA pathway in colorectal cancer. Oncotargets and Therapy, 11, 3765–3774. https://doi.org/10.2147/OTT.S164982
Pelletier, J., Thomas, G., & Volarević, S. (2018). Ribosome biogenesis in cancer: New players and therapeutic avenues. Nature Reviews Cancer, 18(1), 51–63. https://doi.org/10.1038/nrc.2017.104
de Visser, K. E., & Joyce, J. A. (2023). The evolving tumor microenvironment: From cancer initiation to metastatic outgrowth. Cancer Cell, 41(3), 374–403. https://doi.org/10.1016/j.ccell.2023.02.016
Yang, X., Yang, C., Zhang, S., et al. (2024). Precision treatment in advanced hepatocellular carcinoma. Cancer Cell, 42(2), 180–197. https://doi.org/10.1016/j.ccell.2024.01.007
Kim, H. D., Jung, S., Lim, H. Y., et al. (2024). Regorafenib plus nivolumab in unresectable hepatocellular carcinoma: the phase 2 RENOBATE trial. Nature Medicine. Published online February 19, 2024. https://doi.org/10.1038/s41591-024-02824-y
Funding
This study was supported by the National Natural Science Foundation of China (82072618), the National Key Research and Development Program of China (2022YFC2503700, 2022YFC2503705), and Shanghai Municipal Health Commission (2023ZZ02005).
Author information
Authors and Affiliations
Contributions
Conception and design: S-QC, JS, XW, J-KF, F-FM, and Y-CH; Administrative support: S-QC and JS; Provision of study materials: S-QC; Study validation and data curation: XW, J-KF, F-FM, Y-CH, Y-QZ, L-HL, QW, J-XS, and CL; Data analysis and interpretation: XW, J-KF, and F-FM; Manuscript writing: All authors; Final approval of manuscript: All authors.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethical Approval
The research protocol and acquirement of patients’ tumor samples were approved by the Institutional Ethics Committee of Eastern Hepatobiliary Surgery Hospital. All animal experiments were in agreement with the guidelines for the Care and Use of Laboratory Animals and were approved by the Ethics and Welfare Committee for Animal Research at the Yueyang Hospital of Integrated Traditional Chinese and Western Medicine.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, X., Feng, JK., Mao, FF. et al. Prognostic and Immunotherapeutic Predictive Value of CAD Gene in Hepatocellular Carcinoma: Integrated Bioinformatics and Experimental Analysis. Mol Biotechnol (2024). https://doi.org/10.1007/s12033-024-01125-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12033-024-01125-6