Abstract
Background
Deregulating cellular metabolism is one of the prominent hallmarks of malignancy, with a critical role in tumor survival and growth. However, the role of reprogramming aspartate metabolism in hepatocellular carcinoma (HCC) are largely unknown.
Methods
The multi-omics data of HCC patients were downloaded from public databases. Univariate and multivariate stepwise Cox regression were used to establish an aspartate metabolism-related gene signature (AMGS) in HCC. The Kaplan–Meier and receiver operating characteristic curve analyses were performed to evaluate the predictive ability for overall survival (OS) in HCC patients. Gene set enrichment analysis and immune infiltration analysis were operated to determine the potential mechanisms underlying the AMGS. Single-cell RNA sequencing (scRNA-seq) data of liver cancer stem cells were visualized by t-SNE algorithm. In vivo and in vitro experiments were implemented to investigate the biological function of CAD in HCC. In addition, a nomogram based on the AMGS and clinicopathologic characteristics was constructed by univariate and multivariate Cox regression analyses.
Results
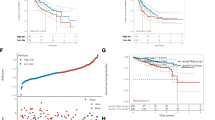
Patients in the high-AMGS subgroup exerted advanced tumor status and poor prognosis. Mechanistically, the high-AMGS subgroup patients had significantly enhanced proliferation and stemness-related pathways, increased infiltration of regulatory T cells and upregulated expression levels of suppressive immune checkpoints in the tumor immune microenvironment. Notably, scRNA-seq data revealed CAD, one of the aspartate metabolism-related gene, is significantly upregulated in liver cancer stem cells. Silencing CAD inhibited proliferative capacity and stemness properties of HCC cells in vitro and in vivo. Finally, a novel nomogram based on the AMGS showed an accurate prediction in HCC patients.
Conclusions
The AMGS represents a promising prognostic value for HCC patients, providing a perspective for finding novel biomarkers and therapeutic targets for HCC.










Similar content being viewed by others
Data availability
The datasets used in this study can be downloaded in the TCGA (https://gdc.cancer.gov/), ICGC (https://dcc.icgc.org/projects/), GEO (https://www.ncbi.nlm.nih.gov/gds/), CPTAC (https://proteomic.datacommons.cancer.gov/pdc/) and KEGG database (https://www.kegg.jp/).
Abbreviations
- HCC:
-
Hepatocellular carcinoma
- AMGS:
-
Aspartate metabolism-related gene signature
- TCGA:
-
The cancer genome atlas
- ICGC:
-
International cancer genome consortium
- GEO:
-
Gene expression omnibus
- scRNA-seq:
-
Single-cell sequencing
- CSCs:
-
Cancer stem cells
- TIME:
-
Tumor immune microenvironment
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Ca: A Cancer J Clinicians. 2021;71(3):209–49. https://doi.org/10.3322/caac.21660
Yang C, Zhang H, Zhang L, Zhu AX, Bernards R, Qin W, et al. Evolving therapeutic landscape of advanced hepatocellular carcinoma. Nat Rev Gastroenterol Hepatol. 2022. https://doi.org/10.1038/s41575-022-00704-9.
Llovet JM, Kelley RK, Villanueva A, Singal AG, Pikarsky E, Roayaie S, et al. Hepatocellular carcinoma. Nature reviews. Dis Primers. 2021;7(1):6. https://doi.org/10.1038/s41572-020-00240-3
Hanahan D. Hallmarks of cancer: new dimensions. Cancer Discov. 2022;12(1):31–46. https://doi.org/10.1158/2159-8290.CD-21-1059.
Martínez-Reyes I, Chandel NS. Cancer metabolism: looking forward. Nat Rev Cancer. 2021;21(10):669–80. https://doi.org/10.1038/s41568-021-00378-6.
Birsoy K, Wang T, Chen WW, Freinkman E, Abu-Remaileh M, Sabatini DM. An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell. 2015;162(3):540–51. https://doi.org/10.1016/j.cell.2015.07.016.
Sullivan LB, Gui DY, Hosios AM, Bush LN, Freinkman E, Vander Heiden MG. Supporting aspartate biosynthesis is an essential function of respiration in proliferating cells. Cell. 2015;162(3):552–63. https://doi.org/10.1016/j.cell.2015.07.017.
Choi B, Coloff JL. The diverse functions of non-essential amino acids in cancer. Cancers (Basel). 2019;11(5):675. https://doi.org/10.3390/cancers11050675.
De Falco P, Lazzarino G, Felice F, Desideri E, Castelli S, Salvatori I, et al. Hindering nat8l expression in hepatocellular carcinoma increases cytosolic aspartate delivery that fosters pentose phosphate pathway and purine biosynthesis promoting cell proliferation. Redox Biol. 2023;59:102585. https://doi.org/10.1016/j.redox.2022.102585.
Rabinovich S, Adler L, Yizhak K, Sarver A, Silberman A, Agron S, et al. Diversion of aspartate in ass1-deficient tumours fosters de novo pyrimidine synthesis. Nature. 2015;527(7578):379–83. https://doi.org/10.1038/nature15529.
Son J, Lyssiotis CA, Ying H, Wang X, Hua S, Ligorio M, et al. Glutamine supports pancreatic cancer growth through a kras-regulated metabolic pathway. Nature. 2013;496(7443):101–5. https://doi.org/10.1038/nature12040.
Helenius IT, Madala HR, Yeh JJ. An asp to strike out cancer? Therapeutic possibilities arising from aspartate’s emerging roles in cell proliferation and survival. Biomolecules. 2021;11(11):1666. https://doi.org/10.3390/biom11111666.
Mering CV. String: a database of predicted functional associations between proteins. Nucleic Acids Res. 2003;31(1):258–61. https://doi.org/10.1093/nar/gkg034.
Zheng H, Pomyen Y, Hernandez MO, Li C, Livak F, Tang W, et al. Single-cell analysis reveals cancer stem cell heterogeneity in hepatocellular carcinoma. Hepatology. 2018;68(1):127–40. https://doi.org/10.1002/hep.29778.
Wilkerson MD, Hayes DN. Consensusclusterplus: a class discovery tool with confidence assessments and item tracking. Bioinformatics. 2010;26(12):1572–3. https://doi.org/10.1093/bioinformatics/btq170.
Blanche P, Dartigues J, Jacqmin-Gadda H. Estimating and comparing time-dependent areas under receiver operating characteristic curves for censored event times with competing risks. Stat Med. 2013;32(30):5381–97. https://doi.org/10.1002/sim.5958.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. Limma powers differential expression analyses for rna-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7): e47. https://doi.org/10.1093/nar/gkv007.
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P. The molecular signatures database hallmark gene set collection. Cell Syst. 2015;1(6):417–25. https://doi.org/10.1016/j.cels.2015.12.004.
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12(5):453–7. https://doi.org/10.1038/nmeth.3337.
Butler A, Hoffman P, Smibert P, Papalexi E, Satija R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol. 2018;36(5):411–20. https://doi.org/10.1038/nbt.4096.
He X, Peng Y, He G, Ye H, Liu L, Zhou Q, et al. Increased co-expression of pd1 and tim3 is associated with poor prognosis and immune microenvironment heterogeneity in gallbladder cancer. J Transl Med. 2023;21(1):717. https://doi.org/10.1186/s12967-023-04589-3.
Zhou Q, Lin J, Yan Y, Meng S, Liao H, Chen R, et al. Inpp5f translocates into cytoplasm and interacts with asph to promote tumor growth in hepatocellular carcinoma. J Exp Clin Cancer Res. 2022;41(1). https://doi.org/10.1186/s13046-021-02216-x
Tauriello DVF, Sancho E, Batlle E. Overcoming tgfβ-mediated immune evasion in cancer. Nat Rev Cancer. 2022;22(1):25–44. https://doi.org/10.1038/s41568-021-00413-6.
Gorgoglione R, Impedovo V, Riley CL, Fratantonio D, Tiziani S, Palmieri L, et al. Glutamine-derived aspartate biosynthesis in cancer cells: role of mitochondrial transporters and new therapeutic perspectives. Cancers (Basel). 2022;14(1):245. https://doi.org/10.3390/cancers14010245.
Ridder DA, Schindeldecker M, Weinmann A, Berndt K, Urbansky L, Witzel HR, et al. Key enzymes in pyrimidine synthesis, cad and cps1, predict prognosis in hepatocellular carcinoma. Cancers (Basel). 2021;13(4):744. https://doi.org/10.3390/cancers13040744.
Li Y, Li B, Xu Y, Qian L, Xu T, Meng G, et al. Got2 silencing promotes reprogramming of glutamine metabolism and sensitizes hepatocellular carcinoma to glutaminase inhibitors. Cancer Res. 2022;82(18):3223–35. https://doi.org/10.1158/0008-5472.CAN-22-0042.
Long PM, Moffett JR, Namboodiri AMA, Viapiano MS, Lawler SE, Jaworski DM. N-acetylaspartate (naa) and n-acetylaspartylglutamate (naag) promote growth and inhibit differentiation of glioma stem-like cells. J Biol Chem. 2013;288(36):26188–200. https://doi.org/10.1074/jbc.M113.487553.
Sun C, Gu Y, Chen G, Du Y. Bioinformatics analysis of stromal molecular signatures associated with breast and prostate cancer. J Comput Biol. 2019;26(10):1130–9. https://doi.org/10.1089/cmb.2019.0045.
Clarke MF. Clinical and therapeutic implications of cancer stem cells. N Engl J Med. 2019;380(23):2237–45. https://doi.org/10.1056/NEJMra1804280.
Lytle NK, Barber AG, Reya T. Stem cell fate in cancer growth, progression and therapy resistance. Nat Rev Cancer. 2018;18(11):669–80. https://doi.org/10.1038/s41568-018-0056-x.
Lee TK, Guan X, Ma S. Cancer stem cells in hepatocellular carcinoma—from origin to clinical implications. Nat Rev Gastroenterol Hepatol. 2022;19(1):26–44. https://doi.org/10.1038/s41575-021-00508-3.
Loong JHC, Wong T, Tong M, Sharma R, Zhou L, Ng K, et al. Glucose deprivation–induced aberrant fut1-mediated fucosylation drives cancer stemness in hepatocellular carcinoma. J Clin Invest. 2021;131(11): e143377. https://doi.org/10.1172/JCI143377.
Mok EHK, Leung CON, Zhou L, Lei MML, Leung HW, Tong M, et al. Caspase-3–induced activation of srebp2 drives drug resistance via promotion of cholesterol biosynthesis in hepatocellular carcinoma. Cancer Res. 2022;82(17):3102–15. https://doi.org/10.1158/0008-5472.CAN-21-2934.
Wang X, Yang K, Wu Q, Kim LJY, Morton AR, Gimple RC, et al. Targeting pyrimidine synthesis accentuates molecular therapy response in glioblastoma stem cells. Sci Transl Med. 2019;11(504):eaau4972. https://doi.org/10.1126/scitranslmed.aau4972
Llovet JM, Castet F, Heikenwalder M, Maini MK, Mazzaferro V, Pinato DJ, et al. Immunotherapies for hepatocellular carcinoma. Nat Rev Clin Oncol. 2022;19(3):151–72. https://doi.org/10.1038/s41571-021-00573-2.
Yau T, Park J, Finn RS, Cheng A, Mathurin P, Edeline J, et al. Nivolumab versus sorafenib in advanced hepatocellular carcinoma (checkmate 459): a randomised, multicentre, open-label, phase 3 trial. Lancet Oncol. 2022;23(1):77–90. https://doi.org/10.1016/S1470-2045(21)00604-5.
Verset G, Borbath I, Karwal M, Verslype C, Van Vlierberghe H, Kardosh A, et al. Pembrolizumab monotherapy for previously untreated advanced hepatocellular carcinoma: data from the open-label, phase ii keynote-224 trial. Clin Cancer Res. 2022;28(12):2547–54. https://doi.org/10.1158/1078-0432.CCR-21-3807.
Donne R, Lujambio A. The liver cancer immune microenvironment: therapeutic implications for hepatocellular carcinoma. Hepatology. 2022:n/a–n/a. https://doi.org/10.1002/hep.32740
Li X, Wenes M, Romero P, Huang SC, Fendt S, Ho P. Navigating metabolic pathways to enhance antitumour immunity and immunotherapy. Nat Rev Clin Oncol. 2019;16(7):425–41. https://doi.org/10.1038/s41571-019-0203-7.
Zhou J, Ding T, Pan W, Zhu L, Li L, Zheng L. Increased intratumoral regulatory t cells are related to intratumoral macrophages and poor prognosis in hepatocellular carcinoma patients. Int J Cancer. 2009;125(7):1640–8. https://doi.org/10.1002/ijc.24556.
Kalathil S, Lugade AA, Miller A, Iyer R, Thanavala Y. Higher frequencies of garp+ctla-4+foxp3+ t regulatory cells and myeloid-derived suppressor cells in hepatocellular carcinoma patients are associated with impaired t-cell functionality. Cancer Res. 2013;73(8):2435–44. https://doi.org/10.1158/0008-5472.CAN-12-3381.
Chang H, Jung W, Kim A, Kim HK, Kim WB, Kim JH, et al. Expression and prognostic significance of programmed death protein 1 and programmed death ligand-1, and cytotoxic t lymphocyte-associated molecule-4 in hepatocellular carcinoma. APMIS. 2017;125(8):690–8. https://doi.org/10.1111/apm.12703.
Liu X, Li M, Wang X, Dang Z, Jiang Y, Wang X, et al. Pd-1+ tigit+ cd8+ t cells are associated with pathogenesis and progression of patients with hepatitis b virus-related hepatocellular carcinoma. Cancer Immunol Immunother. 2019;68(12):2041–54. https://doi.org/10.1007/s00262-019-02426-5.
Li H, Wu K, Tao K, Chen L, Zheng Q, Lu X, et al. Tim-3/galectin-9 signaling pathway mediates t-cell dysfunction and predicts poor prognosis in patients with hepatitis b virus-associated hepatocellular carcinoma. Hepatology. 2012;56(4):1342–51. https://doi.org/10.1002/hep.25777.
Han Y, Chen Z, Yang Y, Jiang Z, Gu Y, Liu Y, et al. Human cd14+ ctla-4+ regulatory dendritic cells suppress t-cell response by cytotoxic t-lymphocyte antigen-4-dependent il-10 and indoleamine-2,3-dioxygenase production in hepatocellular carcinoma. Hepatology. 2014;59(2):567–79. https://doi.org/10.1002/hep.26694.
Acknowledgements
We thank all the databases that provided public data downloads and the researchers who proposed the algorithms used in this study.
Funding
This research was funded by National Natural Science Foundation of China (No. 82073045, 82103090), Guangdong Basic and Applied Basic Research Foundation (No. 2021A1515012107, 2022A1515012391, 2020A1515010232), Beijing Xisike Clinical Oncology Research Foundation (Y-MSDPU2022-0826), Science and Technology Program of Guangzhou (No.202201020311), China Postdoctoral Science Foundation (2020TQ0384, 2021M703742), the Guangdong Science and Technology Department (2020B1212060018), the Key Laboratory of Malignant Tumour Gene Regulation and Target Therapy of Guangdong Higher Education Institutes, Sun Yat‐sen University (Grant KLB09001), and the Key Laboratory of Malignant Tumour Molecular Mechanism and Translational Medicine of Guangzhou Bureau of Science and Information Technology ([2013]163).
Author information
Authors and Affiliations
Contributions
Conceptualization: ZZ and ZX. Methodology: JS and YY. Software: KW. Data collection: JS. Manuscript preparation: HL, HL and CH. Manuscript editing: SM and ZZ. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
The study was approved by the Animal Ethics Committee of Sun Yat-sen University and conducted in accordance with institutional guidelines.
Informed consent
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shi, J., Wen, K., Mui, S. et al. Integrated analysis reveals an aspartate metabolism-related gene signature for predicting the overall survival in patients with hepatocellular carcinoma. Clin Transl Oncol (2024). https://doi.org/10.1007/s12094-024-03431-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12094-024-03431-6