Abstract
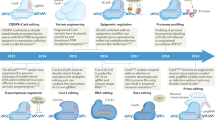
CRISPR genome editing technique has the potential to target cancer cells in a precise manner. The latest advancements have helped to address one of the prominent concerns about this strategy which is the off-target integrations observed with dsDNA and have resulted in more studies being carried out for potentially safer and more targeted gene therapy, so as to make it available for the clinical trials in order to effectively treat cancer. CRISPR screens offer great potential for the high throughput investigation of the gene functionality in various tumors. It extends its capability to identify the tumor growth essential genes, therapeutic resistant genes, and immunotherapeutic responses. CRISPR screens are mostly performed in in vitro models, but latest advancements focus on developing in vivo models to view cancer progression in animal models. It also allows the detection of factors responsible for tumorigenesis. In CRISPR screens key parameters are optimized in order to meet proficient gene targeting efficiencies. It also detects various molecular effectors required for gene regulation in different cancers, essential pathways which modulate cytotoxicity to immunotherapy in cancer cells, important genes which contribute to cancer cell survival in hypoxic states and modulate cancer long non-coding RNAs. The current review focuses on the recent developments in the therapeutic application of CRISPR technology for cancer therapy. Furthermore, the associated challenges and safety concerns along with the various strategies that can be implemented to overcome these drawbacks has been discussed.


Similar content being viewed by others
Abbreviations
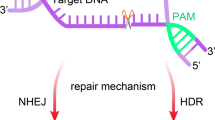
- NHEJ:
-
Non-homologous end joining
- HDR:
-
Homology directed repair
- ODN:
-
Oligodeoxynucleotides
- ZFNs:
-
Zinc-finger nucleases
- TALENs:
-
Transcription activator-like effector nucleases
- CSCs:
-
Cancer stem cells
- Oct2:
-
Organic cation transporter 2
- Sox2:
-
SRY-Box Transcription Factor 2
- STAT3:
-
Signal transducer and activator of transcription 3
- EMT:
-
Epithelial-mesenchymal transition
- ChIP:
-
Chromatin immunoprecipitation
- Csn2:
-
Casein Beta
- crRNA:
-
CRISPR RNAs
- tracrRNA:
-
Trans-activating crispr RNA
- KHOS:
-
Angiogenic and non-angiogenic human osteosarcoma cells
- U-2OS:
-
Human Bone Osteosarcoma Epithelial Cells
- EWSR1:
-
Ewing sarcoma breakpoint region 1
- FLI1:
-
Friend leukemia integration 1 transcription factor
- MCF-7:
-
Michigan Cancer Foundation-7
- CTCF:
-
CCCTC-binding factor
- LNCaP:
-
Lymph Node Carcinoma of the prostate
- KIF4A:
-
Kinesin Family Member 4A
- WDR62:
-
WD Repeat Domain 62
- AGS:
-
Adenocarcinoma gastric cell line.
- SGC:
-
Sebaceous gland carcinoma
- HEK293:
-
Human Embryonic Kidney 293 cells)
- LGALS2:
-
Galectin 2
- AOM/DSS:
-
Azoxymethane /Dextran Sodium Sulfate
- Pten:
-
Phosphatase and tensin homolog
- Usp-7:
-
Ubiquitin specific protease 7
- PTPM1:
-
Protein Tyrosine Phosphatase Mitochondrial 1
- NF-1:
-
Neurofibromin 1
- DUSP9:
-
Dual Specificity Phosphatase 9
- MAPK:
-
Mitogen-activated protein kinase
- FOXO3:
-
Forkhead box O3
- mTOR:
-
Mammalian target of rapamycin
- TGF-β:
-
Transforming growth factor-beta
- NRAS:
-
Neuroblastoma RAS viral oncogene homolog
- MEK:
-
Mitogen-activated extracellular signal-regulated kinase
- SOCS3:
-
Suppressor of cytokine signaling 3
- USP8:
-
Ubiquitin Specific Peptidase 8
- FANCA:
-
Fanconi anemia, complementation group A
- MGMT:
-
O6-Methylguanine-DNA-methyltransferase
- GDSC:
-
Genomics of Drug Sensitivity in Cancer
- CTRP:
-
Cancer Therapeutics Response Portal
- PI3k:
-
Phosphoinositide 3-kinase
- HER-2:
-
Human epidermal growth factor receptor 2
- LRP8:
-
Low-density lipoprotein receptor-related protein 8
- FGFR2:
-
Fibroblast growth factor receptor 2
- MUC1:
-
Mucin 1
References
Loftus, L. V., Amend, S. R., & Pienta, K. J. (2022). Interplay between cell death and cell proliferation reveals new strategies for cancer therapy. International journal of molecular sciences, 23(9), 4723.
Liu, Q., Wang, C., Zheng, Y., Zhao, Y., Wang, Y., Hao, J., Zhao, X., Yi, K., Shi, L., Kang, C., & Liu, Y. (2020). Virus-like nanoparticle as a co-delivery system to enhance efficacy of CRISPR/Cas9-based cancer immunotherapy. Biomaterials, 258, 120275.
Cong, L., Ran, F., Cox, D., Lin, S., Barretto, R., Habib, N., et al. (2013). Multiplex Genome Engineering Using CRISPR/Cas Systems. Science, 339(6121), 819–823.
Jamehdor, S., Zaboli, K. A., Naserian, S., Thekkiniath, J., Omidy, H. A., Teimoori, A., Johari, B., Taromchi, A. H., Sasano, Y., & Kaboli, S. (2020). An overview of applications of CRISPR-Cas technologies in biomedical engineering. Folia Histochemica et Cytobiologica, 58(3), 163–173.
Li, L., Hu, S., & Chen, X. (2018). Non-viral delivery systems for CRISPR/Cas9-based genome editing: Challenges and opportunities. Biomaterials, 171, 207–218.
Zhao, Z., Li, C., Tong, F., Deng, J., Huang, G., & Sang, Y. (2021). Review of applications of CRISPR-Cas9 gene-editing technology in cancer research. Biological Procedures Online, 23(1), 1–13.
Rad, S. M. A. H., Langroudi, L., Kouhkan, F., Yazdani, L., Koupaee, A. N., Asgharpour, S., Shojaei, Z., Bamdad, T., & Arefian, E. (2015). Transcription factor decoy: A pre-transcriptional approach for gene downregulation purpose in cancer. Tumor Biology, 36(7), 4871–4881.
Bigdelou, Z., Mortazavi, Y., Saltanatpour, Z., Asadi, Z., Kadivar, M., & Johari, B. (2020). Role of Oct4–Sox2 complex decoy oligodeoxynucleotides strategy on reverse epithelial to mesenchymal transition (EMT) induction in HT29-ShE encompassing enriched cancer stem-like cells. Molecular Biology Reports, 47(3), 1859–1869.
Johari, B., Rezaeejam, H., Moradi, M., Taghipour, Z., Saltanatpour, Z., Mortazavi, Y., & Nasehi, L. (2020). Increasing the colon cancer cells sensitivity toward radiation therapy via application of Oct4–Sox2 complex decoy oligodeoxynucleotides. Molecular Biology Reports, 47(9), 6793–6805.
Rahmati, M., Johari, B., Kadivar, M., Rismani, E., & Mortazavi, Y. (2020). Suppressing the metastatic properties of the breast cancer cells using STAT3 decoy oligodeoxynucleotides: A promising approach for eradication of cancer cells by differentiation therapy. Journal of Cellular Physiology, 235(6), 5429–5444.
Hille, F., & Charpentier, E. (2016). CRISPR-Cas: Biology, mechanisms and relevance. Philosophical Transactions of The Royal Society B: Biological Sciences, 371(1707), 20150496.
Costa, J. R., Bejcek, B. E., McGee, J. E., et al. (2017). Genome editing using engineered nucleases and their use in genomic screening. In S. Markossian, A. Grossman, K. Brimacombe, et al. (Eds.), Assay Guidance Manual. National Center for Biotechnology Information.
Barrangou, R., Fremaux, C., Deveau, H., Richards, M., Boyaval, P., Moineau, S., Romero, D. A., & Horvath, P. (2007). CRISPR provides acquired resistance against viruses in prokaryotes. Science, 315(5819), 1709–1712.
Makarova, K. S., Wolf, Y. I., Alkhnbashi, O. S., Costa, F., Shah, S. A., Saunders, S. J., Barrangou, R., Brouns, S. J., Charpentier, E., Haft, D. H., & Horvath, P. (2015). An updated evolutionary classification of CRISPR–Cas systems. Nature Reviews Microbiology, 13(11), 722–736.
Makarova, K. S., Haft, D. H., Barrangou, R., Brouns, S. J., Charpentier, E., Horvath, P., Moineau, S., Mojica, F. J., Wolf, Y. I., Yakunin, A. F., & Van Der Oost, J. (2011). Evolution and classification of the CRISPR–Cas systems. Nature Reviews Microbiology, 9(6), 467–477.
Babu, M., Beloglazova, N., Flick, R., Graham, C., Skarina, T., Nocek, B., Gagarinova, A., Pogoutse, O., Brown, G., Binkowski, A., & Phanse, S. (2011). A dual function of the CRISPR–Cas system in bacterial antivirus immunity and DNA repair. Molecular microbiology, 79(2), 484–502.
Wiedenheft, B., Zhou, K., Jinek, M., Coyle, S. M., Ma, W., & Doudna, J. A. (2009). Structural basis for DNase activity of a conserved protein implicated in CRISPR-mediated genome defense. Structure, 17(6), 904–912.
Beloglazova, N., Brown, G., Zimmerman, M. D., Proudfoot, M., Makarova, K. S., Kudritska, M., Kochinyan, S., Wang, S., Chruszcz, M., Minor, W., & Koonin, E. V. (2008). A novel family of sequence-specific endoribonucleases associated with the clustered regularly interspaced short palindromic repeats. Journal of Biological Chemistry, 283(29), 20361–20371.
Yosef, I., Goren, M. G., & Qimron, U. (2012). Proteins and DNA elements essential for the CRISPR adaptation process in Escherichia coli. Nucleic acids research, 40(12), 5569–5576.
Wei, Y., Chesne, M. T., Terns, R. M., & Terns, M. P. (2015). Sequences spanning the leader-repeat junction mediate CRISPR adaptation to phage in Streptococcus thermophilus. Nucleic acids research, 43(3), 1749–1758.
McGinn, J., & Marraffini, L. (2018). Molecular mechanisms of CRISPR–Cas spacer acquisition. Nature Reviews Microbiology, 17(1), 7–12.
Li, M., Wang, R., Zhao, D., & Xiang, H. (2014). Adaptation of the Haloarcula hispanica CRISPR-Cas system to a purified virus strictly requires a priming process. Nucleic acids research, 42(4), 2483–2492.
Vorontsova, D., Datsenko, K. A., Medvedeva, S., Bondy-Denomy, J., Savitskaya, E. E., Pougach, K., Logacheva, M., Wiedenheft, B., Davidson, A. R., Severinov, K., & Semenova, E. (2015). Foreign DNA acquisition by the IF CRISPR–Cas system requires all components of the interference machinery. Nucleic acids research, 43(22), 10848–10860.
Heler, R., Samai, P., Modell, J. W., Weiner, C., Goldberg, G. W., Bikard, D., & Marraffini, L. A. (2015). Cas9 specifies functional viral targets during CRISPR–Cas adaptation. Nature, 519(7542), 199–202.
Wei, Y., Terns, R. M., & Terns, M. P. (2015). Cas9 function and host genome sampling in Type II-A CRISPR–Cas adaptation. Genes & development, 29(4), 356–361.
Mojica, F., Díez-Villaseñor, C., García-Martínez, J., & Almendros, C. (2009). Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology, 155(3), 733–740.
Swarts, D. C., Mosterd, C., Van Passel, M. W., & Brouns, S. J. (2012). CRISPR interference directs strand specific spacer acquisition. PLoS ONE, 7(4), e35888.
Datsenko, K. A., Pougach, K., Tikhonov, A., Wanner, B. L., Severinov, K., & Semenova, E. (2012). Molecular memory of prior infections activates the CRISPR/Cas adaptive bacterial immunity system. Nature communications, 3(1), 1–7.
Richter, C., Dy, R. L., McKenzie, R. E., Watson, B. N., Taylor, C., Chang, J. T., McNeil, M. B., Staals, R. H., & Fineran, P. C. (2014). Priming in the Type IF CRISPR-Cas system triggers strand-independent spacer acquisition, bi-directionally from the primed protospacer. Nucleic acids research, 42(13), 8516–8526.
Redding, S., Sternberg, S. H., Marshall, M., Gibb, B., Bhat, P., Guegler, C. K., Wiedenheft, B., Doudna, J. A., & Greene, E. C. (2015). Surveillance and processing of foreign DNA by the Escherichia coli CRISPR-Cas system. Cell, 163(4), 854–865.
Carte, J., Wang, R., Li, H., Terns, R. M., & Terns, M. P. (2008). Cas6 is an endoribonuclease that generates guide RNAs for invader defense in prokaryotes. Genes & development, 22(24), 3489–3496.
Haurwitz, R. E., Jinek, M., Wiedenheft, B., Zhou, K., & Doudna, J. A. (2010). Sequence-and structure-specific RNA processing by a CRISPR endonuclease. Science, 329(5997), 1355–1358.
Garside, E. L., Schellenberg, M. J., Gesner, E. M., Bonanno, J. B., Sauder, J. M., Burley, S. K., Almo, S. C., Mehta, G., & MacMillan, A. M. (2012). Cas5d processes pre-crRNA and is a member of a larger family of CRISPR RNA endonucleases. RNA, 18(11), 2020–2028.
Deltcheva, E., Chylinski, K., Sharma, C. M., Gonzales, K., Chao, Y., Pirzada, Z. A., Eckert, M. R., Vogel, J., & Charpentier, E. (2011). CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature, 471(7340), 602–607.
Marraffini, L., & Sontheimer, E. (2010). CRISPR interference: RNA-directed adaptive immunity in bacteria and archaea. Nature Reviews Genetics, 11(3), 181–190.
Hale, C. R., Zhao, P., Olson, S., Duff, M. O., Graveley, B. R., Wells, L., Terns, R. M., & Terns, M. P. (2009). RNA-guided RNA cleavage by a CRISPR RNA-Cas protein complex. Cell, 139(5), 945–956.
Fonfara, I., Richter, H., Bratovič, M., Le Rhun, A., & Charpentier, E. (2016). The CRISPR-associated DNA-cleaving enzyme Cpf1 also processes precursor CRISPR RNA. Nature, 532(7600), 517–521.
Zetsche, B., Gootenberg, J. S., Abudayyeh, O. O., Slaymaker, I. M., Makarova, K. S., Essletzbichler, P., Volz, S. E., Joung, J., Van Der Oost, J., Regev, A., & Koonin, E. V. (2015). Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell, 163(3), 759–771.
Marraffini, L. A., & Sontheimer, E. J. (2010). Self-versus non-self-discrimination during CRISPR RNA-directed immunity. Nature, 463(7280), 568–571.
Brouns, S. J., Jore, M. M., Lundgren, M., Westra, E. R., Slijkhuis, R. J., Snijders, A. P., Dickman, M. J., Makarova, K. S., Koonin, E. V., & Van Der Oost, J. (2008). Small CRISPR RNAs guide antiviral defense in prokaryotes. Science, 321(5891), 960–964.
Westra, E. R., van Erp, P. B., Künne, T., Wong, S. P., Staals, R. H., Seegers, C. L., Bollen, S., Jore, M. M., Semenova, E., Severinov, K., & de Vos, W. M. (2012). CRISPR immunity relies on the consecutive binding and degradation of negatively supercoiled invader DNA by Cascade and Cas3. Molecular cell, 46(5), 595–605.
Jinek, M., Chylinski, K., Fonfara, I., Hauer, M., Doudna, J. A., & Charpentier, E. (2012). A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science, 337(6096), 816–821.
Chow, R., & Chen, S. (2018). Cancer CRISPR screens in vivo. Trends in Cancer, 4(5), 349–358.
Xue, H. Y., Ji, L. J., Gao, A. M., Liu, P., He, J. D., & Lu, X. J. (2016). CRISPR-Cas9 for medical genetic screens: Applications and future perspectives. Journal of Medical Genetics, 53, 91–97.
Schuster, A., Erasimus, H., Fritah, S., Nazarov, P. V., van Dyck, E., Niclou, S. P., & Golebiewska, A. (2019). RNAi/CRISPR screens: From a pool to a valid hit. Trends in Biotechnology, 37, 38–55.
He, C., Han, S., Chang, Y., Wu, M., Zhao, Y., Chen, C., & Chu, X. (2021). CRISPR screen in cancer: Status quo and future perspectives. American Journal of Cancer research, 11(4), 1031–1050.
Kuhn, M., Santinha, A., & Platt, R. (2021). Moving from in vitro to in vivo CRISPR screens. Gene And Genome Editing, 2, 100008.
Boutros, M., & Ahringer, J. (2008). The art and design of genetic screens: RNA interference. Nature Reviews Genetics, 9(7), 554–566.
Grimm, S. (2004). The art and design of genetic screens: Mammalian culture cells. Nature Reviews Genetics, 5(3), 179–189.
Shalem, O., Sanjana, N., & Zhang, F. (2015). High-throughput functional genomics using CRISPR–Cas9. Nature Reviews Genetics, 16(5), 299–311.
Sharma, G., Sharma, A., Bhattacharya, M., Lee, S., & Chakraborty, C. (2021). CRISPR-Cas9: A preclinical and clinical perspective for the treatment of human diseases. Molecular Therapy, 29(2), 571–586.
Feng, Y., Sassi, S., Shen, J., Yang, X., Gao, Y., Osaka, E., et al. (2014). Targeting Cdk11 in osteosarcoma cells using the CRISPR-cas9 system. Journal Of Orthopaedic Research, 33(2), 199–207.
Cervera, S. T., Rodríguez-Martín, C., Fernández-Tabanera, E., Melero-Fernández de Mera, R. M., Morin, M., Fernández-Peñalver, S., Iranzo-Martínez, M., Amhih-Cardenas, J., García-García, L., González-González, L., & Moreno-Pelayo, M. A. (2021). Therapeutic potential of EWSR1–FLI1 inactivation by CRISPR/Cas9 in ewing sarcoma. Cancers, 13(15), 3783.
DeSantis, C., Ma, J., Goding Sauer, A., Newman, L., & Jemal, A. (2017). Breast cancer statistics, 2017, racial disparity in mortality by state. CA: A Cancer Journal For Clinicians., 67(6), 439–448.
Wang, Y., Zhang, T., Kwiatkowski, N., Abraham, B. J., Lee, T. I., Xie, S., Yuzugullu, H., Von, T., Li, H., Lin, Z., & Stover, D. G. (2015). CDK7-dependent transcriptional addiction in triple-negative breast cancer. Cell, 163(1), 174–186.
Patel, H., Abduljabbar, R., Lai, C. F., Periyasamy, M., Harrod, A., Gemma, C., Steel, J. H., Patel, N., Busonero, C., Jerjees, D., & Remenyi, J. (2016). Expression of CDK7, Cyclin H, and MAT1 Is elevated in breast cancer and is prognostic in estrogen receptor-positive breast CancerCDK7 expression in breast cancer. Clinical Cancer Research, 22(23), 5929–5938.
Al-Mulhim, F., Alqosaibi, A. I., Al-Muhnna, A., Farid, K., Abdel-Ghany, S., Rizk, H., Prince, A. B., Isichei, A., & Sabit, H. (2021). CRISPR/Cas9-mediated activation of CDH1 suppresses metastasis of breast cancer in rats. Electronic Journal of Biotechnology, 53, 54–60.
Elman, J., Ni, T., Mengwasser, K., Jin, D., Wronski, A., Elledge, S., & Kuperwasser, C. (2019). Identification of FUBP1 as a long tail cancer driver and widespread regulator of tumor suppressor and oncogene alternative splicing. Cell Reports, 28(13), 3435-3449.e5.
Provance, O. K., Geanes, E. S., Lui, A. J., Roy, A., Holloran, S. M., Gunewardena, S., Hagan, C. R., Weir, S., & Lewis-Wambi, J. (2021). Disrupting interferon-alpha and NF-kappaB crosstalk suppresses IFITM1 expression attenuating triple-negative breast cancer progression. Cancer letters, 514, 12–29.
Emadi, F., Teo, T., Rahaman, M., & Wang, S. (2020). CDK12: A potential therapeutic target in cancer. Drug Discovery Today, 25(12), 2257–2267.
Reimers, M. A., Yip, S. M., Zhang, L., Cieslik, M., Dhawan, M., Montgomery, B., Wyatt, A. W., Chi, K. N., Small, E. J., Chinnaiyan, A. M., & Alva, A. S. (2020). Clinical outcomes in cyclin-dependent kinase 12 mutant advanced prostate cancer. European urology, 77(3), 333–341.
Lei, H., Wang, Z., Jiang, D., Liu, F., Liu, M., Lei, X., Yang, Y., He, B., Yan, M., Huang, H., & Liu, Q. (2021). CRISPR screening identifies CDK12 as a conservative vulnerability of prostate cancer. Cell Death & Disease, 12(8), 1–11.
Ahmed, M., Soares, F., Xia, J. H., Yang, Y., Li, J., Guo, H., Su, P., Tian, Y., Lee, H. J., Wang, M., & Akhtar, N. (2021). CRISPRi screens reveal a DNA methylation-mediated 3D genome dependent causal mechanism in prostate cancer. Nature communications, 12(1), 1–15.
Das, R., Sjöström, M., Shrestha, R., Yogodzinski, C., Egusa, E. A., Chesner, L. N., Chen, W. S., Chou, J., Dang, D. K., Swinderman, J. T., & Ge, A. (2021). An integrated functional and clinical genomics approach reveals genes driving aggressive metastatic prostate cancer. Nature Communications, 12(1), 1–12.
Zhang, Y., Peia, J., Shib, S., Guoc, X., Cuid, G., Lie, Y., Zhang, H., & Hu, W. (2019). CRISPR/Cas9-mediated knockout of the PDEF gene inhibits migration and invasion of human gastric cancer AGS cells. Biomedicine & Pharmacotherapy, 111, 76–85.
Haghighi, N., Doosti, A., & Kiani, J. (2021). Evaluation of CRISPR/Cas9 system effects on knocking out NEAT1 gene in AGS gastric cancer cell line with therapeutic perspective. Journal of Gastrointestinal Cancer. https://doi.org/10.1007/s12029-021-00669-z
Shi, M., Wang, C., Ji, J., Cai, Q., Zhao, Q., Xi, W., & Zhang, J. (2022). CRISPR/Cas9-mediated knockout of SGLT1 inhibits proliferation and alters metabolism of gastric cancer cells. Cellular Signalling, 90, 110192.
Su, S., Zou, Z., Chen, F., Ding, N., Du, J., Shao, J., Li, L., Fu, Y., Hu, B., Yang, Y., & Sha, H. (2017). CRISPR-Cas9-mediated disruption of PD-1 on human T cells for adoptive cellular therapies of EBV positive gastric cancer. Oncoimmunology, 6(1), e1249558.
Li, H., Zhao, L., Lau, Y., Zhang, C., & Han, R. (2020). Genome-wide CRISPR screen identifies LGALS2 as an oxidative stress-responsive gene with an inhibitory function on colon tumor growth. Oncogene, 40(1), 177–188.
Chen, Z., Wu, J., Liu, B., Zhang, G., Wang, Z., Zhang, L., Wang, K., Fan, Z., & Zhu, P. (2021). Identification of cis-HOX-HOXC10 axis as a therapeutic target for colorectal tumor-initiating cells without APC mutations. Cell Reports, 36(4), 109431.
Michels, E. B., Mosa, H. M., Streibl, I. B., Zhan, T., Menche, C., Ardat, A. K., Darvishi, T., Czlonka, E., Wagner, S., Winter, J., Medyouf, H., Boutros, M., & Farin, F. H. (2020). Pooled in vitro and in vivo CRISPR-Cas9 screening identifies tumor suppressors in human colon organoids. Cell Stem Cell, 26, 782–792.
Veo, B., Danis, E., Pierce, A., Wang, D., Fosmire, S., Sullivan, K. D., Joshi, M., Khanal, S., Dahl, N., Karam, S., & Serkova, N. (2021). Transcriptional control of DNA repair networks by CDK7 regulates sensitivity to radiation in MYC-driven medulloblastoma. Cell Reports, 35(4), 109013.
Zhan, M., Sun, X., Liu, J., Li, Y., Li, Y., He, X., Zhou, Z., & Lu, L. (2017). Usp7 promotes medulloblastoma cell survival and metastasis by activating Shh pathway. Biochemical and biophysical research communications, 484(2), 429–434.
Bao, M. H. R., Yang, C., Tse, A. P. W., Wei, L., Lee, D., Zhang, M. S., Goh, C. C., Chiu, D. K. C., Yuen, V. W. H., Law, C. T., & Chin, W. C. (2021). Genome-wide CRISPR-Cas9 knockout library screening identified PTPMT1 in cardiolipin synthesis is crucial to survival in hypoxia in liver cancer. Cell Reports, 34(4), 108676.
Lu, Y., Shen, H., Huang, W., He, S., Chen, J., Zhang, D., Shen, Y., & Sun, Y. (2021). Genome-scale CRISPR-Cas9 knockout screening in hepatocellular carcinoma with lenvatinib resistance. Cell death discovery, 7(1), 1–12.
Abraham, C. G., Ludwig, M. P., Andrysik, Z., Pandey, A., Joshi, M., Galbraith, M. D., Sullivan, K. D., & Espinosa, J. M. (2018). ΔNp63α suppresses TGFB2 expression and RHOA activity to drive cell proliferation in squamous cell carcinomas. Cell reports, 24(12), 3224–3236.
Takahashi, N., Cho, P., Selfors, L. M., Kuiken, H. J., Kaul, R., Fujiwara, T., Harris, I. S., Zhang, T., Gygi, S. P., & Brugge, J. S. (2020). 3D culture models with CRISPR screens reveal hyperactive NRF2 as a prerequisite for spheroid formation via regulation of proliferation and ferroptosis. Molecular cell, 80(5), 828–844.
Wohlhieter, C. A., Richards, A. L., Uddin, F., Hulton, C. H., Quintanal-Villalonga, À., Martin, A., de Stanchina, E., Bhanot, U., Asher, M., Shah, N. S., & Hayatt, O. (2020). Concurrent mutations in STK11 and KEAP1 promote ferroptosis protection and SCD1 dependence in lung cancer. Cell reports, 33(9), 108444.
Inturi, R., & Jemth, P. (2021). CRISPR/Cas9-based inactivation of human papillomavirus oncogenes E6 or E7 induces senescence in cervical cancer cells. Virology, 562, 92–102.
Li, Z., Wang, B., Gu, S., Jiang, P., Sahu, A., Chen, C., et al. (2020). CRISPR Screens Identify Essential Cell Growth Mediators in BRAF Inhibitor-resistant Melanoma. Genomics, Proteomics & Bioinformatics, 18(1), 26–40.
Nagler, A., Vredevoogd, D. W., Alon, M., Cheng, P. F., Trabish, S., Kalaora, S., Arafeh, R., Goldin, V., Levesque, M. P., Peeper, D. S., & Samuels, Y. (2020). A genome-wide CRISPR screen identifies FBXO42 involvement in resistance toward MEK inhibition in NRAS-mutant melanoma. Pigment cell & melanoma research, 33(2), 334–344.
Palit, S. A., van Dorp, J., Vis, D., Lieftink, C., Linder, S., Beijersbergen, R., Bergman, A. M., Zwart, W., & van der Heijden, M. S. (2021). A kinome-centered CRISPR-Cas9 screen identifies activated BRAF to modulate enzalutamide resistance with potential therapeutic implications in BRAF-mutated prostate cancer. Scientific reports, 11(1), 1–8.
Chen, J., Huang, Y., Tang, Z., Li, M., Ling, X., Liao, J., Zhou, X., Fang, S., Zhao, H., Zhong, W., & Yuan, X. (2021). Genome-Scale CRISPR-Cas9 transcriptional activation screening in metformin resistance related gene of prostate cancer. Frontiers in Cell and Developmental Biology, 8, 616332.
MacLeod, G., Bozek, D. A., Rajakulendran, N., Monteiro, V., Ahmadi, M., Steinhart, Z., Kushida, M. M., Yu, H., Coutinho, F. J., Cavalli, F. M., & Restall, I. (2019). Genome-wide CRISPR-Cas9 screens expose genetic vulnerabilities and mechanisms of temozolomide sensitivity in glioblastoma stem cells. Cell reports, 27(3), 971–986.
Oberlick, E. M., Rees, M. G., Seashore-Ludlow, B., Vazquez, F., Nelson, G. M., Dharia, N. V., Weir, B. A., Tsherniak, A., Ghandi, M., Krill-Burger, J. M., & Meyers, R. M. (2019). Small-molecule and CRISPR screening converge to reveal receptor tyrosine kinase dependencies in pediatric rhabdoid tumors. Cell reports, 28(9), 2331–2344.
Ayestaran, I., Galhoz, A., Spiegel, E., Sidders, B., Dry, J. R., Dondelinger, F., Bender, A., McDermott, U., Iorio, F., & Menden, M. P. (2020). Identification of intrinsic drug resistance and its biomarkers in High-throughput pharmacogenomic and CRISPR screens. Patterns, 1(5), 100065.
Hou, J., Cao, X., Cheng, Y., & Wang, X. (2020). Roles of TP53 gene in the development of resistance to PI3K inhibitor resistances in CRISPR-Cas9-edited lung adenocarcinoma cells. Cell Biology and Toxicology, 36(5), 481–492.
Shu, S., Wu, H. J., Jennifer, Y. G., Zeid, R., Harris, I. S., Jovanović, B., Murphy, K., Wang, B., Qiu, X., Endress, J. E., & Reyes, J. (2020). Synthetic lethal and resistance interactions with BET bromodomain inhibitors in triple-negative breast cancer. Molecular cell, 78(6), 1096–1113.
Merino, D., Whittle, J. R., Vaillant, F., Serrano, A., Gong, J. N., Giner, G., Maragno, A. L., Chanrion, M., Schneider, E., Pal, B., & Li, X. (2017). Synergistic action of the MCL-1 inhibitor S63845 with current therapies in preclinical models of triple-negative and HER2-amplified breast cancer. Science translational medicine, 9(401), 7049.
Cai, J., Chen, J., Wu, T., Cheng, Z., Tian, Y., Pu, C., Shi, W., Suo, X., Wu, X., & Zhang, K. (2020). Genome-scale CRISPR activation screening identifies a role of LRP8 in Sorafenib resistance in Hepatocellular carcinoma. Biochemical and Biophysical Research Communications, 526(4), 1170–1176.
Huang, S., Ma, Z., Zhou, Q., Wang, A., Gong, Y., Li, Z., Wang, S., Yan, Q., Wang, D., Hou, B., & Zhang, C. (2022). Genome-wide CRISPR/Cas9 library screening identified that dusp4 deficiency induces lenvatinib resistance in hepatocellular carcinoma. International Journal of Biological Sciences, 18(11), 4357–4371.
Bester, A. C., Lee, J. D., Chavez, A., Lee, Y. R., Nachmani, D., Vora, S., Victor, J., Sauvageau, M., Monteleone, E., Rinn, J. L., & Provero, P. (2018). An integrated genome-wide CRISPRa approach to functionalize lncRNAs in drug resistance. Cell, 173(3), 649–664.
Damnernsawad, A., Bottomly, D., Kurtz, S. E., Eide, C. A., McWeeney, S. K., Tyner, J. W., & Nechiporuk, T. (2022). A genome-wide CRISPR screen identifies regulators of MAPK and MTOR pathways that mediate resistance to sorafenib in acute myeloid leukemia. Haematologica, 107(1), 77.
Chen, J., Bell, J., Lau, B., Whittaker, T., Stapleton, D., & Ji, H. (2019). A functional CRISPR/Cas9 screen identifies kinases that modulate FGFR inhibitor response in gastric cancer. Oncogenesis, 8(5), 1–9.
Ning, G., Zhu, Q., Kang, W., Lee, H., Maher, L., Suh, Y. S., Michaud, M., Silva, M., Kwon, J. Y., Zhang, C., & Lee, C. (2021). A novel treatment strategy for lapatinib resistance in a subset of HER2-amplified gastric cancer. BMC Cancer, 21(1), 1–17.
Yuan, F., Sun, M., Liu, H., & Qian, F. (2020). Albumin-conjugated drug is irresistible by single gene mutation of endocytic system: Verification by genome-wide CRISPR-Cas9 loss-of-function screens. Journal Of Controlled Release, 323, 311–320.
Skripova, V., Serebriiskii, I., Abramova, Z., Astsaturov, I., & Kiyamova, R. (2017). CRISPR/Cas9 technique for identification of genes regulating oxaliplatin resistance of pancreatic cancer cell line. BioNanoScience, 7(1), 97–100.
Liu, T., Li, Z., Zhang, Q., Bernstein, K. D. A., Lozano-Calderon, S., Choy, E., Hornicek, F. J., & Duan, Z. (2016). Targeting ABCB1 (MDR1) in multi-drug resistant osteosarcoma cells using the CRISPR-Cas9 system to reverse drug resistance. Oncotarget, 7(50), 83502.
Xiao, Z., Wan, J., Nur, A. A., Dou, P., Mankin, H., Liu, T., & Ouyang, Z. (2018). Targeting CD44 by CRISPR-Cas9 in multi-drug resistant osteosarcoma cells. Cellular physiology and biochemistry, 51(4), 1879–1893.
Liu, D., Zhao, X., Tang, A., Xu, X., Liu, S., Zha, L., Ma, W., Zheng, J., & Shi, M. (2020). CRISPR screen in mechanism and target discovery for cancer immunotherapy. Biochimica et Biophysica Acta (BBA)-Reviews on Cancer, 1874(1), 188378.
Sharma, P., & Allison, J. P. (2015). The future of immune checkpoint therapy. Science, 348(6230), 56–61.
Shiota, M., Fujimoto, J., Semba, T., Satoh, H., Yamamoto, T., & Mori, S. (1994). Hyperphosphorylation of a novel 80 kDa protein-tyrosine kinase similar to Ltk in a human Ki-1 lymphoma cell line, AMS3. Oncogene, 9(6), 1567–1574.
Morris, S. W., Naeve, C., Mathew, P., James, P. L., Kirstein, M. N., Cui, X., & Witte, D. P. (1997). ALK, the chromosome 2 gene locus altered by the t (2; 5) in non-Hodgkin’s lymphoma, encodes a novel neural receptor tyrosine kinase that is highly related to leukocyte tyrosine kinase (LTK). Oncogene, 14(18), 2175–2188.
Zhang, J., Song, Z., Wang, H., Lang, L., Yang, Y., Xiao, W., Webster, D., Wei, W., Barta, S., Kadin, M., Staudt, L., Nakagawa, M., & Yang, Y. (2019). A novel model of controlling PD-L1 expression in ALK+ anaplastic large cell lymphoma revealed by CRISPR screening. Blood, 134(2), 171–185.
Majzner, R. G., & Mackall, C. L. (2018). Tumor antigen escape from CAR T-cell therapy. Cancer discovery, 8(10), 1219–1226.
Iorgulescu, J. B., Braun, D., Oliveira, G., Keskin, D. B., & Wu, C. J. (2018). Acquired mechanisms of immune escape in cancer following immunotherapy. Genome Med., 10(1), 87.
Brudno, J. N., Maric, I., Hartman, S. D., Rose, J. J., Wang, M., Lam, N., Stetler-Stevenson, M., Salem, D., Yuan, C., Pavletic, S., & Kanakry, J. A. (2018). T cells genetically modified to express an anti–B-cell maturation antigen chimeric antigen receptor cause remissions of poor-prognosis relapsed multiple myeloma. Journal of Clinical Oncology, 36(22), 2267.
Cohen, A. D., Garfall, A. L., Stadtmauer, E. A., Melenhorst, J. J., Lacey, S. F., Lancaster, E., Vogl, D. T., Weiss, B. M., Dengel, K., Nelson, A., & Plesa, G. (2019). B cell maturation antigen–specific CAR T cells are clinically active in multiple myeloma. The Journal of clinical investigation, 129(6), 2210–2221.
Ramkumar, P., Abarientos, A., Tian, R., Seyler, M., Leong, J., Chen, M., Choudhry, P., Hechler, T., Shah, N., Wong, S., Martin, T., Wolf, J., Roybal, K., Pahl, A., Taunton, J., Wiita, A., & Kampmann, M. (2020). CRISPR-based screens uncover determinants of immunotherapy response in multiple myeloma. Blood Advances, 4(13), 2899–2911. https://doi.org/10.1182/bloodadvances.2019001346
Dufva, O., Koski, J., Maliniemi, P., Ianevski, A., Klievink, J., Leitner, J., Pölönen, P., Hohtari, H., Saeed, K., Hannunen, T., & Ellonen, P. (2020). Integrated drug profiling and CRISPR screening identify essential pathways for CAR T-cell cytotoxicity. Blood, 135(9), 597–609.
Dong, M. B., Wang, G., Chow, R. D., Ye, L., Zhu, L., Dai, X., Park, J. J., Kim, H. R., Errami, Y., Guzman, C. D., & Zhou, X. (2019). Systematic immunotherapy target discovery using genome-scale in vivo CRISPR screens in CD8 T cells. Cell, 178(5), 1189–1204.
Siolas, D., Vucic, E., Kurz, E., Hajdu, C., & Bar-Sagi, D. (2021). Gain-of-function p53R172H mutation drives accumulation of neutrophils in pancreatic tumors, promoting resistance to immunotherapy. Cell Reports, 36(8), 109578.
Soares, F., Chen, B., Lee, J. B., Ahmed, M., Ly, D., Tin, E., Kang, H., Zeng, Y., Akhtar, N., Minden, M. D., & He, H. H. (2021). CRISPR screen identifies genes that sensitize AML cells to double-negative T-cell therapy. Blood, 137(16), 2171–2181.
Dersh, D., Phelan, J. D., Gumina, M. E., Wang, B., Arbuckle, J. H., Holly, J., Kishton, R. J., Markowitz, T. E., Seedhom, M. O., Fridlyand, N., & Wright, G. W. (2021). Genome-wide screens identify lineage-and tumor-specific genes modulating MHC-I-and MHC-II-restricted immunosurveillance of human lymphomas. Immunity, 54(1), 116–131.
Kim, M. Y., Yu, K. R., Kenderian, S. S., Ruella, M., Chen, S., Shin, T. H., Aljanahi, A. A., Schreeder, D., Klichinsky, M., Shestova, O., & Kozlowski, M. S. (2018). Genetic inactivation of CD33 in hematopoietic stem cells to enable CAR T cell immunotherapy for acute myeloid leukemia. Cell, 173(6), 1439–1453.
Lu, Y., Xue, J., Deng, T., Zhou, X., Yu, K., Deng, L., Huang, M., Yi, X., Liang, M., Wang, Y., & Shen, H. (2020). Safety and feasibility of CRISPR-edited T cells in patients with refractory non-small-cell lung cancer. Nature medicine, 26(5), 732–740.
Stadtmauer, E. A., Fraietta, J. A., Davis, M. M., Cohen, A. D., Weber, K. L., Lancaster, E., Mangan, P. A., Kulikovskaya, I., Gupta, M., Chen, F., & Tian, L. (2020). CRISPR-engineered T cells in patients with refractory cancer. Science, 367(6481), eaba7365.
Tian, X., Gu, T., Patel, S., Bode, A. M., Lee, M. H., & Dong, Z. (2019). CRISPR/Cas9–An evolving biological tool kit for cancer biology and oncology. NPJ precision oncology, 3(1), 1–8.
Hu, Z., Yu, L., Zhu, D., Ding, W., Wang, X., Zhang, C., Wang, L., Jiang, X., Shen, H., He, D., & Li, K. (2014). Disruption of HPV16-E7 by CRISPR/Cas system induces apoptosis and growth inhibition in HPV16 positive human cervical cancer cells. BioMed research international. https://doi.org/10.1155/2014/612823
Uddin, F., Rudin, C., & Sen, T. (2020). CRISPR gene therapy: applications, limitations, and implications for the future. Frontiers In Oncology, 10, 1387.
Sýkora, P. (2018). Chapter 11 germline gene therapy in the era of precise genome editing: how far should we go? In M. Soniewicka (Ed.), The ethics of reproductive genetics (pp. 157–171). Springer.
Liang, P., Xu, Y., Zhang, X., Ding, C., Huang, R., Zhang, Z., Lv, J., Xie, X., Chen, Y., Li, Y., & Sun, Y. (2015). CRISPR/Cas9-mediated gene editing in human tripronuclear zygotes. Protein & cell, 6(5), 363–372.
Robert, F., Barbeau, M., Éthier, S., Dostie, J., & Pelletier, J. (2015). Pharmacological inhibition of DNA-PK stimulates Cas9-mediated genome editing. Genome Medicine, 7(1), 1–11.
Brokowski, C., & Adli, M. (2019). CRISPR ethics: moral considerations for applications of a powerful tool. Journal Of Molecular Biology, 431(1), 88–101.
Tang, L., Zeng, Y., Du, H., Gong, M., Peng, J., Zhang, B., Lei, M., Zhao, F., Wang, W., Li, X., & Liu, J. (2017). CRISPR/Cas9-mediated gene editing in human zygotes using Cas9 protein. Molecular genetics and genomics, 292(3), 525–533.
Kaiser, J., & Normile, D. (2015). Embryo engineering study splits scientific community. Science, 348(6234), 486–487.
Janik, E., Niemcewicz, M., Ceremuga, M., Krzowski, L., Saluk-Bijak, J., & Bijak, M. (2020). Various aspects of a gene editing system—CRISPR–Cas9. International Journal Of Molecular Sciences, 21(24), 9604.
Khalaf, K., Janowicz, K., Dyszkiewicz-Konwińska, M., Hutchings, G., Dompe, C., Moncrieff, L., Jankowski, M., Machnik, M., Oleksiewicz, U., Kocherova, I., & Petitte, J. (2020). CRISPR/Cas9 in cancer immunotherapy: Animal models and human clinical trials. Genes, 11(8), 921.
He, S. (2020). The first human trial of CRISPR-based cell therapy clears safety concerns as new treatment for late-stage lung cancer. Signal Transduction and Targeted Therapy, 5(1), 1–2.
Lino, C., Harper, J., Carney, J., & Timlin, J. (2018). Delivering CRISPR: A review of the challenges and approaches. Drug Delivery, 25(1), 1234–1257.
Soussi, T., & Wiman, K. (2015). TP53: An oncogene in disguise. Cell Death & Differentiation, 22(8), 1239–1249.
Moon, S., Kim, D., Ko, J., & Kim, Y. (2019). Recent advances in the CRISPR genome editing tool set. Experimental & Molecular Medicine, 51(11), 1–11.
Han, H., Pang, J., & Soh, B. (2020). Mitigating off-target effects in CRISPR/Cas9-mediated in vivo gene editing. Journal Of Molecular Medicine, 98(5), 615–632.
Araki, M., & Ishii, T. (2016). Providing appropriate risk information on genome editing for patients. Trends In Biotechnology, 34(2), 86–90.
Lau, C., & Suh, Y. (2017). In vivo genome editing in animals using AAV-CRISPR system: applications to translational research of human disease. F1000Research, 6, 2153.
Walton, R., Christie, K., Whittaker, M., & Kleinstiver, B. (2020). Unconstrained genome targeting with near-PAMless engineered CRISPR-Cas9 variants. Science, 368(6488), 290–296.
O’Connell, M., Oakes, B., Sternberg, S., East-Seletsky, A., Kaplan, M., & Doudna, J. (2014). Programmable RNA recognition and cleavage by CRISPR/Cas9. Nature, 516(7530), 263–266.
Strutt, S., Torrez, R., Kaya, E., Negrete, O., & Doudna, J. (2018). RNA-dependent RNA targeting by CRISPR-Cas9. eLife. https://doi.org/10.7554/eLife.32724
Ihry, R. J., Worringer, K. A., Salick, M. R., Frias, E., Ho, D., Theriault, K., Kommineni, S., Chen, J., Sondey, M., Ye, C., & Randhawa, R. (2018). p53 inhibits CRISPR–Cas9 engineering in human pluripotent stem cells. Nature medicine, 24(7), 939–946.
Haapaniemi, E., Botla, S., Persson, J., Schmierer, B., & Taipale, J. (2018). CRISPR–Cas9 genome editing induces a p53-mediated DNA damage response. Nature Medicine, 24(7), 927–930.
Kosicki, M., Tomberg, K., & Bradley, A. (2018). Repair of double-strand breaks induced by CRISPR–Cas9 leads to large deletions and complex rearrangements. Nature Biotechnology, 36(8), 765–771.
Moses, C., Nugent, F., Waryah, C., Garcia-Bloj, B., Harvey, A., & Blancafort, P. (2019). Activating PTEN Tumor Suppressor Expression with the CRISPR/dCas9 System. Molecular Therapy - Nucleic Acids, 14, 287–300.
Xu, X., & Qi, L. (2019). A CRISPR–dCas toolbox for genetic engineering and synthetic biology. Journal Of Molecular Biology, 431(1), 34–47.
Charlesworth, C. T., Deshpande, P. S., Dever, D. P., Camarena, J., Lemgart, V. T., Cromer, M. K., Vakulskas, C. A., Collingwood, M. A., Zhang, L., Bode, N. M., & Behlke, M. A. (2019). Identification of preexisting adaptive immunity to Cas9 proteins in humans. Nature medicine, 25(2), 249–254.
Moreno, A. M., Palmer, N., Alemán, F., Chen, G., Pla, A., Jiang, N., Leong Chew, W., Law, M., & Mali, P. (2019). Immune-orthogonal orthologues of AAV capsids and of Cas9 circumvent the immune response to the administration of gene therapy. Nature biomedical engineering, 3(10), 806–816.
Selvakumar, S. C., Preethi, K. A., Ross, K., Tusubira, D., Khan, M. W. A., Mani, P., Rao, T. N., & Sekar, D., (2022). CRISPR/Cas9 and next generation sequencing in the personalized treatment of Cancer. Molecular Cancer, 21(1), 1–14.
Tang, H., & Shrager, J. B. (2016). CRISPR/Cas-mediated genome editing to treat EGFR-mutant lung cancer: A personalized molecular surgical therapy. EMBO Molecular Medicine, 8(2), 83–85.
Chen, Z. H., Yu, Y. P., Zuo, Z. H., Nelson, J. B., Michalopoulos, G. K., Monga, S., Liu, S., Tseng, G., & Luo, J. H. (2017). Targeting genomic rearrangements in tumor cells through Cas9-mediated insertion of a suicide gene. Nature biotechnology, 35(6), 543–550.
Liu, J. J., Orlova, N., Oakes, B. L., Ma, E., Spinner, H. B., Baney, K. L., Chuck, J., Tan, D., Knott, G. J., Harrington, L. B., & Al-Shayeb, B. (2019). CRISPR-CasX is an RNA-dominated enzyme active for human genome editing. Nature, 566(7743), 218.
Tsuchida, C. A., Zhang, S., Doost, M. S., Zhao, Y., Wang, J., O’Brien, E., Fang, H., Li, C. P., Li, D., Hai, Z. Y., & Chuck, J. (2022). Chimeric CRISPR-CasX enzymes and guide RNAs for improved genome editing activity. Molecular Cell, 82(6), 1199–1209.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Maity, S., Mukherjee, R. & Banerjee, S. Recent Advances and Therapeutic Strategies Using CRISPR Genome Editing Technique for the Treatment of Cancer. Mol Biotechnol 65, 206–226 (2023). https://doi.org/10.1007/s12033-022-00550-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-022-00550-9