Abstract
COVID-19, caused by SARS-CoV-2, is currently spreading around the world and causing many casualties. Antibodies against such emerging infectious diseases are one of the important tools for basic viral research and the development of diagnostic and therapeutic agents. CR3022 is a monoclonal antibody against the receptor binding domain (RBD) of the spike protein (S protein) of SARS-CoV found in SARS patients, but it was also shown to have strong affinity for that of SARS-CoV-2. In this study, we produced large amounts of three formats of CR3022 antibodies (scFv, Fab and IgG) with high purity using a silkworm-baculovirus expression vector system. Furthermore, SPR measurements showed that the affinity of those silkworm-produced IgG antibodies to S protein was almost the same as that produced in mammalian expression system. These results indicate that the silkworm-baculovirus expression system is an excellent expression system for emerging infectious diseases that require urgent demand for diagnostic agents and therapeutic agents.
Similar content being viewed by others
Introduction
In December 2019, coronavirus disease 2019 (COVID-19) caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) broke out in Wuhan, China and rapidly spread around the world [1]. In May 2020, the WHO recognized it as a pandemic, more than 106 million people infected and 2.3 million people dead as of February 10, 2020 [2]. In 2002 and 2012, two other human coronaviruses, SARS-CoV [3] and MERS-CoV [4], have spread wide area. These three coronaviruses belong to beta-coronaviruses, probably originated from bats, and have viral envelope with spike glycoprotein (S protein) [5,6,7]. The S protein is the major viral antigen and binds the host receptors and mediates virus-host membrane fusion when the virus invades the host cells [8, 9]. The structure and function of the SARS-CoV-2 S protein have been analyzed in detail by the previous studies [10] and revealed that its receptor binding domain (RBD) adopts receptor binding sensitive (open or up) or insensitive (close or down) conformations within S protein [11,12,13]. Both SARS-CoV-2 and SARS-CoV mainly use human ACE2 membrane protein as a receptor [9] and their amino acid sequences of RBD are highly conserved 73% [14].
The monoclonal antibody CR3022, a neutralizing antibody targeting the RBD domain of SARS-CoV, was isolated by screening using a phage display library constructed from blood of a convalescent SARS patient from Singapore [15, 16]. Interestingly, the previous studies have shown that CR3022 is capable of binding to the RBD of SARS-CoV-2 in which 24 of the CR3022 buried region (28 residues) are conserved between SARS-CoV and SARS-CoV-2 [17]. Recent report also demonstrated that CR3022 neutralizes SARS-CoV-2 through a neutralization mechanism that is not detectable in some assays [18], although other studies showed that CR3022 does not neutralize SARS-CoV-2 probably due to a single mutation, A384P, among the four unconserved residues [17, 19]. These studies suggest that CR3022 could be useful for therapeutic regent against COVID-19, especially in combination with other neutralizing antibodies. Structural and biochemical analyses revealed that the binding epitope of CR3022 does not overlap with that of a neutralizing antibody isolated from a SARS-CoV-2 patient, thus the multiple antibodies are able to bind to the S protein without competition [20, 21].
In therapeutic and serological applications of antibodies, in addition to IgG, which has high detection sensitivity and high affinity for antigens due to its avidity effect, scFv and Fab are used because of their simple structure, high productivity, and small molecular weight, which makes it easy to control chemical modification of proteins and fusion with large enzymes [22, 23]. Post-translational modifications are important for antibodies to have biological functions, especially glycosylation, and it has been reported that the lack of glycosylation actually lowers the affinity of antibodies to Fcγ receptors [24]. The E. coli expression system is simple and capable of expressing a large amount of protein, but not suitable for IgG production due to lack of the post-translational modifications such as disulfide bonds and glycosylation. Hence, the IgG is typically produced by transfecting expression plasmids into mammalian cells, such as CHO cells or HEK293 cells [25, 26]. However, the selection of stable cell lines expressing high amount of IgG is time-consuming, and transfection of large amount of plasmid is expensive when the commercially available kits are used. The baculovirus expression vector system (BEVS) using cultured insect cells is suitable for expressing eukaryotic proteins because of its post-translational modification is similar to that for mammalian cells. The BEVS using silkworm (silkworm-BEVS) is a more efficient system than that using the insect cells for expressing heterologous proteins. It is easy to scale up without evaluating growth conditions; simply increasing the number of silkworms rearing. In fact, our laboratory has successfully used the silkworm-BEVS to produce a variety of useful proteins [27,28,29].
In this study, we proposed that silkworm-BEVS is effective for rapid delivery of therapeutic and diagnostic agents against emerging infectious diseases, using the production of recombinant CR3022 antibody as an example. We produced more than 1 mg of CR3022 in different formats, scFv, Fab, and IgG, from silkworm serum and evaluated its binding affinity to the S protein of SARS-CoV-2. The results showed that silkworm-BEVS was effective in producing high quality and functional CR3022 antibodies.
Materials and Methods
Cell Culture and Silkworm Strain
Insect culture cells, BmN (Bombyx mori-derived cells) cells (Funakoshi Inc.), were cultured at 27 °C using IPL-41 insect medium (Sigma, St. Louis, MO) containing 10% Fatal Bovine Serum (Gibco, Grand Island, NY). The f38 strain of silkworm used in this study was provided by the Genetic Resources Development Center, Faculty of Agriculture, Kyushu University supported by the National BioResource Project (NBRP). Silkworm larvae were reared by fresh mulberry leaves at 25 to 27 °C.
Construction of Recombinant Baculovirus
The single chain variable fragments (scFv) of CR3022 heavy chain (Hc) (GenBank: ABA54613.1) and light chain (Lc) (GenBank: ABA54614.1) linked by a flexible peptide linker (GGGGS × 3) was chemically synthesized (Integrated DNA Technologies Inc.). To construct gateway-based entry clones, the coding region of scFvs (VHVL, VLVH) were amplified by polymerase chain reaction (PCR) with KOD-Plus-Neo DNA polymerase (TOYOBO, Tokyo, Japan) using the primer sets (VHVL-F, R and VLVH-F, R, Table 1). The SalI and XhoI-digested amplicons were inserted into the modified pENTR11 (XhoI-digested pENTRL21c-TEV-H8S amplicon; our laboratory stocks) vector by Ligation High (TOYOBO, Tokyo, Japan). As described in our previous report [27], the lobster L21 sequence (L21) was used to enhance translation, the silkworm 30 K-signal peptide (30 K) to secrete CR3022 antibodies into silkworm sera, and the C-terminal His8-STREP double tag is used to facilitate purification. The resulting constructs were designated pENTR11-L21-30K-VH(G4S)VL-TEV-H8S and pENTR11-L21-30K-VL(G4S)VH-TEV-H8S, respectively. To generate the Fab (HFab and Lc) expression vectors, CH1 and CL region were obtained by PCR using the CH1-G-R, F and CL-G-R, F primer sets (Table 1) from the vector of our previous study [30], and then these regions were connected to VH and VL gene by the Gibson assembly method [31], respectively. Regarding the Hc of IgG (HFab-Fc) expression vector, the Fc region was connected to the HFab by PCR using Fc-G-F and Fc-G-R primers in the same manner as above. The resulting constructs were designated pENTR11-L21-30K-HFab-TEV-H8S, pENTR11-L21-30K-Lc-TEV-H8S, and pENTR11-L21-30K-Hc-TEV-H8S respectively. As illustrated in Fig. 1b, the resulting constructs were then transferred to the modified pDEST8/v-cath-EGFP vector (Invitrogen, Carlsbad, CA, our laboratory stock) by the Gateway LR reaction to generate baculovirus transfer plasmid following the manufacturer’s protocol. The five recombinant baculoviruses were created using BmNPV/T3 bacmid DNA according to the previous study [32]. The bacmid DNA was then transfected into the BmN cell using Avalanche®-Everyday transfection reagent (EZ Biosystems, College Park, MD, USA) to produce the recombinant virus particles. The culture supernatant was harvested as the P1 virus on the 5th day after cell transfection. The high-titer virus (P3) stock (~ 1 × 107 plaque-forming unit/ml) was prepared through a serial infection of baculovirus in cultured cells. All viruses for silkworm infections were kept at 4 °C in the dark until use.
a Schematic diagram of the three formats of CR3022 antibodies (Hc: heavy chain, Lc: light chain, HFab: N-terminal half of one heavy chain). In the scFvs (VHVL and VLVH), the antibody regions of the VH and VL are linked by a linker (3 × G4S). All three formats have a purification tag for facilitating purification. b Construction of the recombinant baculoviruses expressing the three formats of CR3022 antibodies (scFv, Fab and IgG). The expression of them were under the control of the polyhedrin promoter (PH) and follow by an SV40 polyadenylation signal (SV40). B1 and B2: recombination sites for Gateway cloning; L21: leader sequence for enhancing translation efficiency; 30 K: signal peptide of silkworm 30 kDa protein; TEV: tobacco etch virus protease cleavage site; 8xHis: 8 × Histidine tag; Strep: Strep tag
Expression of Recombinant scFv, Fab and IgG in Silkworm Hemolymph
The recombinant baculovirus expressing the scFv (~ 1 × 105 plaque-forming unit per larvae) were injected into the 5th instar silkworm larvae (day 3). In the case of Fab and IgG, the two recombinant baculoviruses, expressing HFab and Lc or Hc and Lc, respectively, in different ratio were infected to BmN cells and all samples were prepared based on the method according to the previous study [27]. After optimizing the co-infection ratio of two recombinant baculoviruses using BmN cells, the recombinant baculoviruses expressing HFab and Lc or Hc and Lc were injected into the 5th instar silkworm larvae (day 3) at a ratio of 3:1. Silkworm hemolymph was collected into a tube containing 20 mM 1-phenyl-2-thiourea at 4 dpi and centrifuged at 9000 rpm for 10 min at 4 °C to remove insoluble matter. The supernatant sample was immediately frozen at − 80 °C.
SDS-PAGE and Western Blotting Analysis
The expression of CR3022 antibodies in all samples was analyzed using sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) followed by Western blotting analysis. The samples were separated by 5–20% or 12% SDS-PAGE under the reducing or non-reducing conditions and transferred onto polyvinylidene difluoride (PVDF) membrane (Millipore, Billerica, MA), and further blocked in TBST buffer (20 mM Tris–HCl pH 7.6; 500 mM NaCl; 0.1% w/v Tween-20) with 5% w/v skim milk. The membrane was incubated with HisProbe-HRP (1:5000) (Thermo Fisher Scientific, Waltham, MA) at 4 °C. The specific protein bands were visualized using the Super Signal West Pico Chemiluminescent Substrate (Thermo Fisher Scientific, Waltham, MA).
Purification of Recombinant CR3022 Antibodies
The purification procedure was performed using HisTrap excel column (GE Healthcare Bioscience, Piscataway, NJ) and Strep-Tactin HP column (IBA GmbH, Germany). This protocol for two-step affinity chromatography was previously optimized in our laboratory [27]. The clarified silkworm serum was diluted up to 50 mL by binding buffer (20 mM Tris–HCl pH 7.5; 0.5 M NaCl), and filtered through 0.45 μm filters (Millipore, Billerica, MA) after centrifuged at 9000 rpm for 10 min at 4 °C. The sample was loaded onto a 5 mL HisTrap excel column, and after washing with 25 mL binding buffer containing 30 mM imidazole, the proteins were eluted with 12 mL and 15 mL binding buffer containing 100 mM and 500 mM imidazole, respectively. The eluted fractions containing antibodies were collected and concentrated to 5 mL by ultrafiltration using Amicon Ultra-15 10 K filters (Millipore, Billerica, MA).
Then, the sample removing excess imidazole was further diluted up to 50 mL by PBS and load onto a 5 mL Strep-Tactin column. After the wash with 25 mL of PBS, the proteins were eluted with 24 mL of PBS supplemented with 2.5 mM desthiobiotin, then the fractions containing antibodies were collected and concentrated to 2 mL by ultrafiltration using Amicon Ultra-15 10 K filters. The concentrated sample was dialyzed by PBS, and the final concentration of the protein was quantified by ImageJ using bovine serum albumin (BSA) as standard.
Spike-Antibody Interaction Analysis by Surface Plasmon Resonance (SPR) Experiments
SPR experiments were performed using a Biacore X100 (GE Healthcare Bioscience, Piscataway, NJ) at 25 °C. SARS-CoV-2 spike proteins were immobilized on a CM5 sensor chip (Cytiva, Tokyo, Japan) using standard amine-coupling chemistry. This SARS-CoV-2 Spike protein was expressed using a silkworm-baculovirus expression vector system and purified from silkworm serum using the two-step chromatography as well as the recombinant antibodies. Immobilization densities on flow cell 2 are ~ 2000 for CR3022 VHVL, VLVH, and Fab and ~ 1000 RU for IgG, respectively. Flow cell 1 was left blank to serve as a reference surface. All the surfaces were blocked with 1 M ethanolamine, pH 8.5. A two-fold dilution series of the antibodies (15.6–250 nM of VHVL, VLVH and Fab, 0.63–10 nM of IgG) diluted in a running buffer (PBS with 0.005% Tween-20) was injected with a flow rate of 30 μL/min for 3 min followed by 10 min of dissociation step. The surfaces were regenerated with 30 s injection of 10 mM Gly-HCl, pH 2.5 followed by 30 s injection of 3 M MgCl2. The data were fit to a simple 1:1 interaction model using the global data analysis available within Biacore X100 evaluation software.
ELISA
ELISA was performed to examine the binding affinity of CR3022 IgG to S protein. The 96-well plate were coated with four concentrations of S protein (0.1, 1, 10, 100 ng) and incubated overnight at 4 °C. The plates were washed four times with 300 μL PBST (PBS with 0.1% Tween-20), blocked with 5% skim milk in PBST for 2 h at room temperature, and then the four concentrations of purified CR3022 IgG (0.01, 0.1, 1, 10 μg/mL) were added to the wells and incubated for 1 h at 37 °C. The plates were washed four times and incubated with Anti-Human IgG Fc which conjected with HRP (1:10,000) (#ab98624, Abcam, Cambridge, UK). After 1 h incubation at 37 °C, the plate was washed, developed with TMB substrate and the reaction was stopped by the addition of 2 N H2SO4. Then the absorbance read at 450 nm and 620 nm.
Results
Construction of the Recombinant Baculoviruses and Expression of scFv, Fab and IgG
To establish the low cost and efficient expression system of CR3022 scFv, Fab and IgG, we have generated 5 recombinant baculoviruses shown in Fig. 1b as described under the Materials and methods. The Heavy chain (Hc) and Light chain (Lc) of CR3022 sequence were codon-optimized for B. mori s and were chemically synthesized. A signal peptide of a silkworm 30 kDa protein (30 K-signal peptide) was fused to the N-termini of the target proteins, and a lobster L21 sequence was inserted upstream of the 30 K-signal peptide to increase the amount of protein translation. In addition, TEV cleavage site, His and STREP tag were fused to the C-termini of all constructs for purification, except for the Lc expression vector.
For scFvs (VHVL, VLVH) expression, the 5th instar silkworm larvae were infected with the corresponding recombinant baculoviruses. The expression of these antibodies in silkworm hemolymph was then confirmed by Western blotting analysis. As shown in Fig. 2a, b, VHVL and VLVH were expressed in silkworm hemolymph and their predicted molecular weight were approximately 30 kDa regardless of reducing or non-reducing conditions, respectively.
To optimize the expression of Fab and IgG as heterodimer or heterotetramer, respectively, we varied the ratio of two viruses co-infected into BmN cells and confirmed the expression of them by Western blotting at 3 dpi (the virus combinations to express Fab and IgG are shown in Fig. 1b). Under the non-reducing conditions, it is demonstrated that the optimal infection ratio of the two recombinant baculoviruses, HFab and Lc for Fab or Hc and Lc for IgG, was 3:1 in the medium of cultured BmN cells (Supplemental Fig. S1a, b). Therefore, this 3:1 of recombinant baculoviruses co-infection ratio was used for the efficient expression of Fab and IgG in silkworm larvae. As shown in Fig. 2a, Lanes 3 and 4, the expression of the HFab of Fab or Hc of IgG with molecular weight of approximately 27 or 53 kDa, respectively, were confirmed by Western blotting (reducing conditions) in hemolymph of the 5th instar larvae at 4dpi. Furthermore, the assembled Fab and IgG were observed in the non-reducing conditions (Fig. 2b, Lanes 3 and 4). When the hemolymph of silkworm f38 strain was subjected to Western blotting analysis using His-probe, the non-specific background signal was detected around 100 kDa in all lanes (Fig. 2, asterisk).
Purification of CR3022 Recombinant Antibodies from Silkworm Hemolymph
All four CR3022-derived antibodies successfully expressed in silkworm hemolymph were purified by two-step affinity chromatography. As shown in Fig. 3, the affinity purification through His tag and Strep tag chromatography is simple and highly efficient; CBB staining of SDS-PAGE gel clearly shows that the recombinant CR3022 antibodies could be purified with high purity. In the cases of Fab and IgG, the Lc without purification tags were detected in the same elution fractions with HFab and Hc at a molecular ratio of approximately 1:1 (Fig. 3c, d), suggesting that disulfide bonds between Lc and HFab or Hc are correctly formed. Moreover, in the Histidine affinity purification, the scFvs and Fab begin to elute in the binding buffer containing 100 mM imidazole, whereas most of the IgG elutes in the binding buffer containing 500 mM imidazole (Fig. 3d, Lane 5 and 6). Since the IgG molecule forming homodimer by disulfide bonds has two purification tags, it is expected to have strong binding affinity to nickel conjected agarose beads. These purified proteins were dialyzed against 1 × PBS and quantified. As a result, 2.34 mg of VHVL, 2.64 mg of VLVH, 1.44 mg of Fab, and 1.30 mg of IgG were purified from 10 ml of silkworm hemolymph, and the SDS-PAGE verifications for each purified antibody demonstrated that they were achieved in good purity, 95% for VHVL, 98% for VLVH, 93% for Fab, and 84% for IgG, respectively, as judged by image analysis of gels.
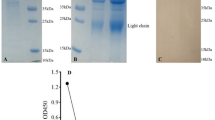
Purification of the CR3022 antibodies from hemolymph of silkworm larvae infected or co-infected with recombinant baculoviruses using the two-step affinity chromatography (a VHVL, b VLVH, c Fab, d IgG) as described in Materials and Methods. The arrows indicate the each of recombinant antibodies. IP: input; FT: flow-through; WS: wash fraction; lanes 1–4: elution fractions by 100 mM imidazole; lanes 5–10: elution fractions by 500 mM imidazole; lanes 11–19: elution fractions by 2.5 mM desthiobiotin. Protein samples from each step were resolved in 12% SDS-PAGE and visualized by Coomassie Brilliant Blue (CBB) R-250 staining
Binding Analysis of CR3022 Recombinant Antibodies
The binding affinity of CR3022 scFvs, Fab and IgG with SARS-CoV-2 Spike (S) protein was measured using surface plasmon resonance (SPR) analysis (Fig. 4). The dissociation constants (KD) of the VHVL, VLVH and Fab with the ectodomain of SARS-CoV-2 S protein (a.a. 1–1208; without furin cleavage site (residues 682–685) and proline substitutions at residues 986 and 987) [33] were 20.1 nM, 27.8 nM and 45.4 nM, respectively (Fig. 4a–c). On the other hand, CR3022 exhibited a much higher affinity for S protein with the KD of 0.21 nM, which was approximately 50–100 greater than Fab and scFvs (Fig. 4d). Furthermore, to evaluate whether the CR3022 IgG produced in silkworm are usable for ELISA, different amounts of S protein (0.1, 1, 10, and 100 ng) were immobilized on a 96-well plate and its binding with CR3022 IgG was measured. As shown in Fig. 5, the CR3022 IgG was able to detect SARS-CoV-2 S protein in a dose-dependent manner. With 100 ng of S protein on the plate, the binding signal reached the plateau at a concentration of 1 μg/mL CR3022 IgG. In addition, when using 10 μg/mL IgG, the detection limit was ~ 100 pg of S protein (Fig. 5, inlet).
Binding affinity of CR3022 antibodies with S protein. The binding affinity of CR3022 VHVL (a), VLVH (b), Fab (c) and IgG (d) with S protein were measured using surface plasmon resonance as described in Materials and Methods. The data were fit to a simple 1:1 interaction model using the global data analysis available within Biacore X100 evaluation software
Discussion
Upon the emerging of infectious diseases spreading all over the world, it is important to urgently proceed with basic scientific research on such pathogenic the viruses and microorganisms. These basic studies are indispensable for the development of diagnostic and therapeutic agents to prevent the diseases and protect our health. In particular, antibodies specific for pathogenic microorganisms are one of the most important tools in these studies.
Using a Silkworm-BEVS, we produced three formats of CR3022 antibodies (scFvs, Fab and IgG) which have been first reported as a neutralizing antibody against the RBD of SARS-CoV and recently reported to bind strongly to that of SARS-CoV-2. The S protein, the major antigen of SARS-CoV-2, is very similar to that of the previously prevalent SARS-CoV, with 73% homology in amino acid sequences of RBD, a target for diagnostic and therapeutic agents [34].
In this study, we produced more than 1 mg of four different recombinant CR3022 antibodies from 10 mL of silkworm hemolymph within a month (Fig. 3). Several expression systems were used to produce human scFv [35,36,37] and the E. coli expression system is one of the most suitable expression systems for mass production of scFv [38]. However, it is time-consuming to purify and refold the scFv expressed in E. coli from the inclusion body [39], whereas the silkworm-BEVS is capable of producing scFvs comparable to the E. coli expression system without refolding. In addition, it is very difficult to produce Fab or IgG in E. coli which lacks post-translational modifications. Cultured mammalian cells also provide a platform for producing recombinant antibodies in large scale, but the screening of cell lines with high-expression and the modulation of culture conditions are time-consuming (minimum 12 months) [25]. Using silkworm-BEVS, it is possible to express and purify the biologically active Fab and IgG with correct disulfide bond formation in a much shorter time than mammalian cell expression systems. For CR3022 IgG, it has been reported that an expression level of 130 μg per gram of fresh leaf weight can be obtained using a transient expression system in plants [40]. In this study, 130 μg of purified CR3022 IgG was eventually purified and recovered from 1 mL of silkworm serum using silkworm-BEVS. Although it is difficult to make a simple comparison, these results suggest that the expression level is comparable or more equivalent to that produced in plants.
The SPR measurement showed that CR3022 IgG has higher affinity (KD = 0.21 nM) than VHVL, VLVH and Fab (KD = 20.1 nM, 27.8 nM and 45.4 nM, respectively). The previous study measuring the affinity of CR3022 antibodies produced using the different expression system [14] revealed that the CR3022 scFv produced by E. coli had a KD of 6.3 nM with the RBD, which is higher than that produced by the silkworm-BEVS. This result may be caused by the differences in the antigens or in coupling methods; the S proteins were immobilized by amine-coupling in this study, and RBD was enzymatic biotinylated and immobilized onto a strept-avidin chip in the other studies. The VHVL and VLVH of CR3022 produced using silkworm-BEVS showed almost equivalent productivity of 2.34 mg and 2.64 mg per 10 mL of silkworm serum, respectively, while the affinity of VHVL was slightly higher than that of VLVH. Previous studies have shown that the productivity and affinity of VHVL and VLVH differ depending on the type of antibody [41, 42], and for CR3022, VHVL was considered to be a more suitable form of scFv for diagnostic and pharmaceutical applications than VLVH. It was also reported that the affinity of CR3022 Fab or IgG produced in mammalian cells with the RBD is 115 nM and < 0.1 nM, respectively [17]. The affinity of CR3022 IgG produced using silkworm-BEVS was almost the same as that in the mammalian cells at 0.21 nM. In contrast, the KD of CR3022 Fab produced using silkworm-BEVS (45.4 nM) is higher than that of produced using mammalian system (115 nM). The CR3022 Fab and IgG used in this study were designed to have two disulfide bonds (usually one) connecting CH1 and CL to enhance the productivity of the recombinant antibodies. The KD of CR3022 Fab with two disulfide bonds produced in this study was 2.5-fold lower than that of mammalian cell-produced CR3022 Fab with single disulfide bond, suggesting that an additional disulfide bond probably contributes to increase the binding activity of CR3022 Fab in the absence of Fc regions.
Dose-dependent binding of the CR3022 IgG produced using silkworm-BEVS were shown by ELISA, but the reaction plateaued at a concentration of 100 ng of S protein per well (96-well plate). The detection limit in ELISA was ~ 100 pg (0.7 pM) of S protein, which is less sensitive than other detection systems, as described in previous report [43], and would not be enough to detect the virus protein in sera with low viral titers in early infected patients. To use the IgG as a diagnostic reagent to detect the SARS-CoV-2 from patient sera, the sensitivity of detection system such as an immunochromatography system needs to be improved further. Since several SARS-CoV-2 antibodies recognizing the different regions of RBD were reported [20, 21], it is possible to develop a highly sensitive detection system by combining the multiple antibodies without competition for epitope binding.
Recently, the antibody-dependent cellular cytotoxicity (ADCC) activity of monoclonal antibodies against several viruses-infected cells has been highlighted because of its potential to decrease viral replication and reduce clinical symptoms [44, 45]. ADCC is one of the antiviral activity that induces the cellular cytotoxicity through a natural killer (NK) cell expressing Fc receptors, which binds to Fc region of IgG. [46]. Therefore, this ability of antibodies to induce strong ADCC would be advantageous when used as therapeutic agents. The induction of ADCC is enhanced by the lack of α-1,6-fucose in the N-linked glycan (Asn-297 in the CH2 domain) of the Fc region [47, 48]. Previous studies showed that the antibodies secreted form the transgenic silkworm lack this fucose and is demonstrated to have the ability to induce the strong ADCC [49]. Considering these reports, it is highly likely that CR3022 IgG produced using silkworm-BEVS can induce stronger ADCC than those produced by other expression systems, and is expected to be highly effective in clinical applications.
Conclusion
In conclusion, we demonstrated that the silkworm-BEVS can produce antibodies in three different formats of CR3022 easily in a short-term. Furthermore, the purified CR3022 antibodies showed almost the same affinity for the S protein as those produced in mammalian cells. Thus, our study suggests that the silkworm-BEVS is suitable as an expression system for producing diagnostic and therapeutic agents that can rapidly respond to emerging of infectious diseases including COVID-19.
References
Zu, Z. Y., Jiang, M. D., Xu, P. P., Chen, W., Ni, Q. Q., Lu, G. M., & Zhang, L. J. (2020). Coronavirus disease 2019 (COVID-19): A perspective from China. Radiology. https://doi.org/10.1148/radiol.2020200490
Coronavirus disease (COVID-19). (n.d.). Retrieved February 26, 2021, from https://www.who.int/emergencies/diseases/novel-coronavirus-2019.
Drosten, C., Günther, S., Preiser, W., van der Werf, S., Brodt, H.-R., Becker, S., et al. (2003). Identification of a novel Coronavirus in patients with severe acute respiratory syndrome. New England Journal of Medicine, 348(20), 1967–1976. https://doi.org/10.1056/nejmoa030747
Zaki, A. M., van Boheemen, S., Bestebroer, T. M., Osterhaus, A. D. M. E., & Fouchier, R. A. M. (2012). Isolation of a novel Coronavirus from a man with pneumonia in Saudi Arabia. New England Journal of Medicine, 367(19), 1814–1820. https://doi.org/10.1056/nejmoa1211721
Fan, Y., Zhao, K., Shi, Z. L., & Zhou, P. (2019). Bat coronaviruses in China. Viruses. 11(3), 210
Cui, J., Li, F., & Shi, Z. L. (2019). Origin and evolution of pathogenic coronaviruses. Nature Reviews Microbiology, 17(3), 181–192. https://doi.org/10.1038/s41579-018-0118-9
Zhou, P., Yang, X. L., Wang, X. G., Hu, B., Zhang, L., Zhang, W., et al. (2020). A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature, 579(7798), 270–273. https://doi.org/10.1038/s41586-020-2012-7
Tortorici, M. A., & Veesler, D. (2019). Structural insights into coronavirus entry. Advances in Virus Research, 105, 93–116. https://doi.org/10.1016/bs.aivir.2019.08.002
Zhang, H., Penninger, J. M., Li, Y., Zhong, N., & Slutsky, A. S. (2020). Angiotensin-converting enzyme 2 (ACE2) as a SARS-CoV-2 receptor: Molecular mechanisms and potential therapeutic target. Intensive Care Medicine, 46(4), 586–590. https://doi.org/10.1007/s00134-020-05985-9
Walls, A. C., Park, Y. J., Tortorici, M. A., Wall, A., McGuire, A. T., & Veesler, D. (2020). Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell, 181(2), 281-292.e6. https://doi.org/10.1016/j.cell.2020.02.058
Walls, A. C., Xiong, X., Park, Y. J., Tortorici, M. A., Snijder, J., Quispe, J., et al. (2019). Unexpected receptor functional mimicry elucidates activation of coronavirus fusion. Cell, 176(5), 1026-1039.e15. https://doi.org/10.1016/j.cell.2018.12.028
Li, T., Zheng, Q., Yu, H., Wu, D., Xue, W., Zhang, Y., et al. (2020). Characterization of the SARS-CoV-2 spike in an early prefusion conformation. bioRxiv. https://doi.org/10.1101/2020.03.16.994152
Henderson, R., Edwards, R., Mansouri, K., Janowska, K., Stalls, V., Kopp, M., et al. (2020). Glycans on the SARS-CoV-2 spike control the receptor binding domain conformation. bioRxiv. https://doi.org/10.1101/2020.06.26.173765
Tian, X., Li, C., Huang, A., Xia, S., Lu, S., Shi, Z., et al. (2020). Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerging Microbes and Infections, 9(1), 382–385. https://doi.org/10.1080/22221751.2020.1729069
van den Brink, E. N., ter Meulen, J., Cox, F., Jongeneelen, M. A. C., Thijsse, A., Throsby, M., et al. (2005). Molecular and biological characterization of human monoclonal antibodies binding to the spike and nucleocapsid proteins of severe acute respiratory syndrome coronavirus. Journal of Virology, 79(3), 1635–1644. https://doi.org/10.1128/jvi.79.3.1635-1644.2005
ter Meulen, J., van den Brink, E. N., Poon, L. L. M., Marissen, W. E., Leung, C. S. W., Cox, F., et al. (2006). Human monoclonal antibody combination against SARS coronavirus: Synergy and coverage of escape mutants. PLoS Medicine, 3(7), e237. https://doi.org/10.1371/journal.pmed.0030237
Yuan, M., Wu, N. C., Zhu, X., Lee, C. C. D., So, R. T. Y., Lv, H., et al. (2020). A highly conserved cryptic epitope in the receptor binding domains of SARS-CoV-2 and SARS-CoV. Science, 368(6491), 630–633. https://doi.org/10.1126/science.abb7269
Huo, J., Zhao, Y., Ren, J., Zhou, D., Duyvesteyn, H. M. E., Ginn, H. M., et al. (2020). Neutralization of SARS-CoV-2 by destruction of the prefusion spike. Cell Host and Microbe, 28(3), 445-454.e6. https://doi.org/10.1016/j.chom.2020.06.010
Wu, N. C., Yuan, M., Bangaru, S., Huang, D., Zhu, X., Lee, C.-C.D., et al. (2020). A natural mutation between SARS-CoV-2 and SARS-CoV determines neutralization by a cross-reactive antibody. PLOS Pathogens, 16(12), e1009089. https://doi.org/10.1371/journal.ppat.1009089
Shi, R., Shan, C., Duan, X., Chen, Z., Liu, P., Song, J., et al. (2020). A human neutralizing antibody targets the receptor-binding site of SARS-CoV-2. Nature, 584(7819), 120–124. https://doi.org/10.1038/s41586-020-2381-y
Wu, Y., Wang, F., Shen, C., Peng, W., Li, D., Zhao, C., et al. (2020). A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science, 368(6496), 1274–1278. https://doi.org/10.1126/science.abc2241
Ahmad, Z. A., Yeap, S. K., Ali, A. M., Ho, W. Y., Alitheen, N. B. M., & Hamid, M. (2012). ScFv antibody: Principles and clinical application. Clinical and Developmental Immunology, 2012, 980250. https://doi.org/10.1155/2012/980250
Bera, T. K., Onda, M., Kreitman, R. J., & Pastan, I. (2014). An improved recombinant Fab-immunotoxin targeting CD22 expressing malignancies. Leukemia Research, 38(10), 1224–1229. https://doi.org/10.1016/j.leukres.2014.06.014
Jefferis, R. (2008). Glycosylation of recombinant antibody therapeutics. Biotechnology Progress, 21(1), 11–16. https://doi.org/10.1021/bp040016j
Wurm, F. M. (2004). Production of recombinant protein therapeutics in cultivated mammalian cells. Nature Biotechnology, 22(11), 1393–1398. https://doi.org/10.1038/nbt1026
Jäger, V., Büssow, K., Wagner, A., Weber, S., Hust, M., Frenzel, A., & Schirrmann, T. (2013). High level transient production of recombinant antibodies and antibody fusion proteins in HEK293 cells. BMC Biotechnology, 13(1), 1–20. https://doi.org/10.1186/1472-6750-13-52
Morifuji, Y., Xu, J., Karasaki, N., Iiyama, K., Morokuma, D., Hino, M., et al. (2018). Expression, purification, and characterization of recombinant human α1-antitrypsin produced using silkworm-baculovirus expression system. Molecular Biotechnology, 60(12), 924–934. https://doi.org/10.1007/s12033-018-0127-y
Kakino, K., Masuda, A., Hino, M., Ebihara, T., Xu, J., Mon, H., et al. (2020). Efficient production of recombinant T7 endonuclease I using silkworm-baculovirus expression vector system. Journal of Asia-Pacific Entomology, 23(3), 694–700. https://doi.org/10.1016/j.aspen.2020.05.001
Fujita, R., Hino, M., Ebihara, T., Nagasato, T., Masuda, A., Lee, J. M., et al. (2020). Efficient production of recombinant SARS-CoV-2 spike protein using the baculovirus-silkworm system. Biochemical and Biophysical Research Communications, 529(2), 257–262. https://doi.org/10.1016/j.bbrc.2020.06.020
Shiroishi, M., Ito, Y., Shimokawa, K., Lee, J. M., Kusakabe, T., & Ueda, T. (2018). Structure–function analyses of a stereotypic rheumatoid factor unravel the structural basis for germline-encoded antibody autoreactivity. Journal of Biological Chemistry, 293(18), 7008–7016. https://doi.org/10.1074/jbc.M117.814475
Gibson, D. G., Young, L., Chuang, R. Y., Venter, J. C., Hutchison, C. A., & Smith, H. O. (2009). Enzymatic assembly of DNA molecules up to several hundred kilobases. Nature Methods, 6(5), 343–345. https://doi.org/10.1038/nmeth.1318
Ono, C., Nakatsukasa, T., Nishijima, Y., Asano, S. I., Sahara, K., & Bando, H. (2007). Construction of the BmNPVT3 bacmid system and its application to the functional analysis of BmNPV he65. Journal of Insect Biotechnology and Sericology, 76(3), 161–167. https://doi.org/10.11416/jibs.76.3_161
Wrapp, D., Wang, N., Corbett, K. S., Goldsmith, J. A., Hsieh, C. L., Abiona, O., et al. (2020). Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science, 367(6483), 1260–1263. https://doi.org/10.1126/science.abb2507
Grifoni, A., Sidney, J., Zhang, Y., Scheuermann, R. H., Peters, B., & Sette, A. (2020). A sequence homology and bioinformatic approach can predict candidate targets for immune responses to SARS-CoV-2. Cell Host and Microbe, 27(4), 671-680.e2. https://doi.org/10.1016/j.chom.2020.03.002
Stöger, E., Vaquero, C., Torres, E., Sack, M., Nicholson, L., Drossard, J., et al. (2000). Cereal crops as viable production and storage systems for pharmaceutical scFv antibodies. Plant Molecular Biology, 42(4), 583–590. https://doi.org/10.1023/A:1006301519427
Damasceno, L. M., Pla, I., Chang, H. J., Cohen, L., Ritter, G., Old, L. J., & Batt, C. A. (2004). An optimized fermentation process for high-level production of a single-chain Fv antibody fragment in Pichia pastoris. Protein Expression and Purification, 37(1), 18–26. https://doi.org/10.1016/j.pep.2004.03.019
Kurasawa, J. H., Shestopal, S. A., Jha, N. K., Ovanesov, M. V., Lee, T. K., & Sarafanov, A. G. (2013). Insect cell-based expression and characterization of a single-chain variable antibody fragment directed against blood coagulation factor VIII. Protein Expression and Purification, 88(2), 201–206. https://doi.org/10.1016/j.pep.2012.12.008
Miller, K. D., Weaver-Feldhaus, J., Gray, S. A., Siegel, R. W., & Feldhaus, M. J. (2005). Production, purification, and characterization of human scFv antibodies expressed in Saccharomyces cerevisiae, Pichia pastoris, and Escherichia coli. Protein Expression and Purification, 42(2), 255–267. https://doi.org/10.1016/j.pep.2005.04.015
Sarker, A., Rathore, A. S., & Gupta, R. D. (2019). Evaluation of scFv protein recovery from E. coli by in vitro refolding and mild solubilization process. Microbial Cell Factories, 18(1), 5. https://doi.org/10.1186/s12934-019-1053-9
Rattanapisit, K., Shanmugaraj, B., Manopwisedjaroen, S., Purwono, P. B., Siriwattananon, K., Khorattanakulchai, N., et al. (2020). Rapid production of SARS-CoV-2 receptor binding domain (RBD) and spike specific monoclonal antibody CR3022 in Nicotiana benthamiana. Scientific Reports. https://doi.org/10.1038/s41598-020-74904-1
Lu, D., Jimenez, X., Witte, L., & Zhu, Z. (2004). The effect of variable domain orientation and arrangement on the antigen-binding activity of a recombinant human bispecific diabody. Biochemical and Biophysical Research Communications, 318(2), 507–513. https://doi.org/10.1016/j.bbrc.2004.04.060
Riaño-Umbarila, L., Rojas-Trejo, V. M., Romero-Moreno, J. A., Costas, M., Utrera-Espíndola, I., Olamendi-Portugal, T., et al. (2020). Comparative assessment of the VH-VL and VL-VH orientations of single-chain variable fragments of scorpion toxin-neutralizing antibodies. Molecular Immunology, 122, 141–147. https://doi.org/10.1016/j.molimm.2020.04.015
Yan, Y., Chang, L., & Wang, L. (2020). Laboratory testing of SARS-CoV, MERS-CoV, and SARS-CoV-2 (2019-nCoV): Current status, challenges, and countermeasures. Reviews in Medical Virology, 30(3), e2106. https://doi.org/10.1002/rmv.2106
Milligan, C., Richardson, B. A., John-Stewart, G., Nduati, R., & Overbaugh, J. (2015). Passively acquired antibody-dependent cellular cytotoxicity (ADCC) activity in HIV-infected infants is associated with reduced mortality. Cell Host and Microbe, 17(4), 500–506. https://doi.org/10.1016/j.chom.2015.03.002
Jegaskanda, S., Luke, C., Hickman, H. D., Sangster, M. Y., Wieland-Alter, W. F., McBride, J. M., et al. (2016). Generation and protective ability of influenza virus-specific antibody-dependent cellular cytotoxicity in humans elicited by vaccination, natural infection, and experimental challenge. Journal of Infectious Diseases, 214(6), 945–952. https://doi.org/10.1093/infdis/jiw262
Carter, P. (2001). Improving the efficacy of antibody-based cancer therapies. Nature Reviews Cancer, 1(2), 118–129. https://doi.org/10.1038/35101072
Iida, S., Misaka, H., Inoue, M., Shibata, M., Nakano, R., Yamane-Ohnuki, N., et al. (2006). Nonfucosylated therapeutic IgG1 antibody can evade the inhibitory effect of serum immunoglobulin G on antibody-dependent cellular cytotoxicity through its high binding to FcγRIIIa. Clinical Cancer Research, 12(9), 2879–2887. https://doi.org/10.1158/1078-0432.CCR-05-2619
Iida, S., Kuni-Kamochi, R., Mori, K., Misaka, H., Inoue, M., Okazaki, A., et al. (2009). Two mechanisms of the enhanced antibody-dependent cellular cytotoxicity (ADCC) efficacy of non-fucosylated therapeutic antibodies in human blood. BMC Cancer, 9(1), 58. https://doi.org/10.1186/1471-2407-9-58
Tada, M., Tatematsu, K., Ishii-Watabe, A., Harazono, A., Takakura, D., Hashii, N., et al. (2015). Characterization of anti-CD20 monoclonal antibody produced by transgenic silkworms (Bombyx mori). MAbs, 7(6), 1138–1150. https://doi.org/10.1080/19420862.2015.1078054
Acknowledgements
This work was supported in part by Cabinet Office, Government of Japan, Cross-ministerial Strategic Innovation Promotion Program (SIP), “Technologies for Smart Bio-industry and Agriculture” (funding agency: Bio-oriented Technology Research Advancement Institution, NARO). This work was partially supported by Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)) from Japan Agency for Medical Research and Development (AMED) under Grant Number JP20am0101091.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ebihara, T., Masuda, A., Takahashi, D. et al. Production of scFv, Fab, and IgG of CR3022 Antibodies Against SARS-CoV-2 Using Silkworm-Baculovirus Expression System. Mol Biotechnol 63, 1223–1234 (2021). https://doi.org/10.1007/s12033-021-00373-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-021-00373-0