Abstract
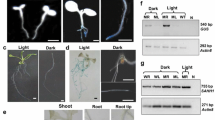
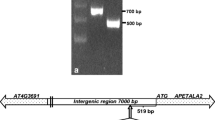
Intergenic regions of divergent gene pairs show bidirectional promoter activity but whether regulatory sequences for gene expression in opposite directions are shared is not established. In this study, promoters of divergently arranged gene pair At4g35640–At4g35650 (SERAT3;2-IDH-III) of Arabidopsis thaliana were analyzed to identify overlapping regulatory regions. Both genes showed the highest expression in flower buds and flowers. 5′ RACE experiments extended the intergenic region from 161 bp shown in TAIR annotation to 512 bp. GUS analysis of transgenic A. thaliana plants carrying the 691 bp fragment (512 bp intergenic region plus 5′ UTR of both the genes) linked to uidA gene revealed that SERAT3;2 promoter drives gene expression in the tapetum, whereas IDH-III promoter functions specifically in microspores/pollen. Serial 5′ deletion of the 691 bp fragment showed SERAT3;2 promoter extends up to −355 position, whereas IDH-III promoter encompasses the 512 bp intergenic region. In transgenics, uidA transcript levels were lower than native SERAT3;2 and IDH-III transcripts indicating presence of additional cis regulatory elements beyond the 691 bp fragment. The present study demonstrated for the first time occurrence of a nested promoter in plants and identified a novel bidirectional promoter capable of driving gene expression in tapetum and microspores/pollen.




Similar content being viewed by others
References
Trinklein, N. D., Aldred, S. F., Hartman, S. J., Schroeder, D. I., Otillar, R. P., & Myers, R. M. (2004). An abundance of bidirectional promoters in the human genome. Genome Research, 14, 62–66.
Grouse, G. F., Leys, E. J., McEwan, R. N., Frayne, E. G., & Kellems, R. E. (1985). Analysis of the mouse dhfr promoter region: Existence of a divergently transcribed gene. Molecular and Cellular Biology, 5, 1847–1858.
Herr, D. R., & Harris, G. L. (2004). Close head-to-head juxtaposition of genes favors their coordinate regulation in Drosophila melanogaster. FEBS Letters, 572, 147–153.
Shin, R., Kim, M. J., & Paek, K. H. (2003). The CaTin1 (Capsicum annuum TMV induced clone 1) and CaTin1-2 genes are linked head-to-head and share a bidirectional promoter. Plant and Cell Physiology, 44, 549–554.
Yang, M. Q., Taylor, J., & Elnitski, L. (2008). Comparative analyses of bidirectional promoters in vertebrates. BMC Bioinformatics, 9(Suppl 6), S9.
Dhadi, S. R., Krom, N., & Ramakrishna, W. (2009). Genome-wide comparative analysis of putative bidirectional promoters from rice, Arabidopsis and Populus. Gene, 429, 65–73.
Krom, N., & Ramakrishna, W. (2008). Comparative analysis of divergent and convergent gene pairs and their expression patterns in rice, Arabidopsis, and Populus. Plant Physiology, 147, 1763–1773.
Wang, Q., Wan, L., Li, D., Zhu, L., Qian, M., & Deng, M. (2009). Searching for bidirectional promoters in Arabidopsis thaliana. BMC Bioinformatics, 10(Suppl 1), S29.
Bondino, H. G., & Valle, E. M. (2009). A small intergenic region drives exclusive tissue specific expression of the adjacent genes in Arabidopsis thaliana. BMC Molecular Biology, 10, 1471–2199.
Banerjee, J., Sahoo, D. K., Dey, N., Houtz, R. L., & Maiti, I. B. (2013). An intergenic region shared by At4g35985 and At4g35987 in Arabidopsis thaliana is a tissue specific and stress inducible bidirectional promoter analyzed in transgenic Arabidopsis and tobacco plants. PLoS ONE, 8(11), e79622.
Kourmpetli, S., Lee, K., Hemsley, R., Rossignol, P., Papageorgiou, T., & Drea, S. (2013). Bidirectional promoters in seed development and related hormone/stress responses. BMC Plant Biology, 13, 187.
Mishra, R. C., & Grover, A. (2014). Intergenic sequence between Arabidopsis caseinolytic protease B cytoplasmic/heat shock protein 100 and choline kinase genes functions as a heat-inducible bidirectional promoter. Plant Physiology, 166, 1646–1658.
Liu, S. J., Yue, Q. J., & Zhang, W. (2015). Structural and functional analysis of an asymmetric bidirectional promoter in Arabidopsis thaliana. Journal of Integrative Plant Biology, 57, 162–170.
Mitra, A., Han, J., Zhang, Z. J., & Mitra, A. (2009). The intergenic region of Arabidopsis thaliana cab1 and cab2 divergent genes functions as a bidirectional promoter. Planta, 229, 1015–1022.
Liu, X., Zhou, X., Li, Y., Tian, J., Zhang, Q., Li, S., et al. (2014). Identification and functional characterization of bidirectional gene pairs and their intergenic regions in maize. BMC Genomics, 15, 338.
Wang, R., Yan, Y., Zhu, M., Yang, M., Zhou, F., Chen, H., et al. (2016). Isolation and functional characterization of bidirectional promoters in rice. Frontiers in Plant Science, 7, 766.
Murashige, T., & Skoog, F. (1962). A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiologia Plantarum, 15, 473–497.
Livak, K. J., & Schmittgen, T. D. (2001). Analysis of relative gene expression data using real–time quantitative PCR and the 2−ΔΔCT method. Methods, 25, 402–408.
Murray, M. G., & Thompson, W. F. (1980). Rapid isolation of high molecular weight plant DNA. Nucleic Acids Research, 8, 4321–4326.
Coutu, C., Brandle, J., Brown, D., Brown, K., Miki, B., Simmonds, J., et al. (2007). pORE: A modular binary vector series suited for both monocot and dicot plant transformation. Transgenic Research, 16, 771–781.
Clough, S. J., & Bent, A. F. (1998). Floral dip: A simplified method for Agrobacterium–mediated transformation of Arabidopsis thaliana. The Plant Journal, 16, 735–743.
Jefferson, R. A., Kavanagh, T. A., & Bevan, M. W. (1987). GUS fusions: β glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO Journal, 6, 3901–3907.
Higo, K., Ugawa, Y., Iwamoto, M., & Korenaga, T. (1999). Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Research, 27, 297–300.
Chow, C. N., Zheng, H. Q., Wu, N. Y., Chien, C. H., Huang, H. D., Lee, T. Y., et al. (2016). PlantPAN 2.0: An update of plant promoter analysis navigator for reconstructing transcriptional regulatory networks in plants. Nucleic Acids Research, 44(database issue), D1154–D1160.
Kawashima, C. G., Berkowitz, O., Hell, R., Noji, M., & Saito, K. (2005). Characterization and expression analysis of a serine acetyl transferase gene family involved in a key step of the sulfur assimilation pathway in Arabidopsis. Plant Physiology, 137, 220–230.
Lemaitre, T., & Hodges, M. (2006). Expression analysis of Arabidopsis thaliana NAD-dependent isocitrate dehydrogenase genes shows the presence of a functional subunit that is mainly expressed in the pollen and absent from vegetative organs. Plant and Cell Physiology, 47, 634–643.
Rogers, H. J., Bate, N., Combe, J., Sullivan, J., Sweetman, J., Swan, C., et al. (2001). Functional analysis of cis–regulatory elements within the promoter of the tobacco late pollen gene g10. Plant Molecular Biology, 45, 577–585.
Swapna, L., Khurana, R., Kumar, S. V., Tyagi, A. K., & Rao, K. V. (2011). Pollen-specific expression of Oryza sativa indica Pollen Allergen Gene (OSIPA) promoter in rice and Arabidopsis transgenic systems. Molecular Biotechnology, 48, 49–59.
Koyanagi, K. O., Hagiwara, M., Itoh, T., Gojobori, T., & Imanishi, T. (2005). Comparative genomics of bidirectional gene pairs and its implications for the evolution of a transcriptional regulation system. Gene, 353, 169–176.
Sharma, I., Srinivasan, R., Ahuja, P. S., Bhat, S. R., & Sreenivasulu, Y. (2015). Identification and characterization of a T-DNA promoter trap line of Arabidopsis thaliana uncovers an embryo sac-specific bi-directional Promoter. Plant Molecular Biology Reporter, 33, 1404–1412.
Xie, M., He, Y., & Gan, S. (2001). Bidirectionalization of polar promoters in plants. Nature Biotechnology, 19, 677–679.
Lata, S., Ranjan, A., Kushwah, N. S., Kumar, P., Dargan, S., Srinivasan, R., et al. (2016). Regulatory sequences of the Arabidopsis thaliana Rps19, a nuclear gene encoding mitochondrial ribosomal protein subunit, extend into the upstream gene. Journal of Plant Biochemistry and Biotechnology. doi:10.1007/s13562-016-0392-4.
Watanabe, M., Mochida, K., Kato, T., Tabata, S., Yoshimoto, N., Noji, M., et al. (2008). Comparative genomics and reverse genetics analysis reveal indispensable functions of the serine acetyl transferase gene family in Arabidopsis. Plant Cell, 20, 2484–2496.
Lemaitre, T., Wochniak, E. U., Flesch, V., Bismuth, E., Fernie, A. R., & Hodges, M. (2007). NAD-dependent isocitrate dehydrogenase mutants of Arabidopsis suggest the enzyme is not limiting for nitrogen assimilation. Plant Physiology, 144, 1546–1558.
Acknowledgements
The financial support for this work was obtained from the Indian Council of Agricultural Research grant under the National Agricultural Science Fund. We thank the National Phytotron Facility, Indian Agricultural Research Institute, New Delhi for providing space for growing Arabidopsis under contained conditions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
Expression profile of IDH-III gene at eFP bowser (DOC 270 kb)
Rights and permissions
About this article
Cite this article
Raipuria, R.K., Kumar, V., Guruprasad, K.N. et al. Analysis of Promoters of Arabidopsis thaliana Divergent Gene Pair SERAT3;2 and IDH-III Shows SERAT3;2 Promoter is Nested Within the IDH-III Promoter. Mol Biotechnol 59, 294–304 (2017). https://doi.org/10.1007/s12033-017-0016-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-017-0016-9