Abstract
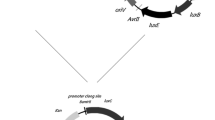
Environmental contamination by toxic organic compounds and antimicrobials is one of the causes for the recent surge of multidrug-resistant pathogenic bacteria. Monitoring contamination is therefore the first step in containment of antimicrobial resistance and requires the development of simple, sensitive, and quantitative tools that detect a broad spectrum of toxic compounds. In this study, we have engineered a new microbial biosensor based on the ttgR-regulated promoter that controls expression of the TtgABC extrusion efflux pump of Pseudomonas putida, coupled to a gfp reporter. The system was introduced in P. putida DOT-T1E, a strain characterized by its ability to survive in the presence of high concentrations of diverse toxic organic compounds. This whole-cell biosensor is capable to detect a wide range of structurally diverse antibiotics, as well as compounds such as toluene or flavonoids.




Similar content being viewed by others
References
Gross, M. (2013). Antibiotics in crisis. Current Biology: CB, 23, R1063–R1065.
CDC (2013) Antibiotic resistance threats in the United States. URL http://www.cdc.gov/drugresistance/threat-report-2013/.
World Health Organization. (2012). The evolving threat of antimicrobial resistance: Options for action. Geneva: World Health Organization.
Korsrud, G. O., Boison, J. O., Nouws, J. F., & MacNeil, J. D. (1998). Bacterial inhibition tests used to screen for antimicrobial veterinary drug residues in slaughtered animals. Journal of AOAC International, 81, 21–24.
Schenck, F. J., & Callery, P. S. (1998). Chromatographic methods of analysis of antibiotics in milk. Journal of Chromatography A, 812, 99–109.
Lee, H. J., Lee, M. H., Ryu, P. D., Lee, H., & Cho, M. H. (2001). Enzyme-linked immunosorbent assay for screening the plasma residues of tetracycline antibiotics in pigs. Journal of Veterinary Medical Science, 63, 553–556.
Loomans, E. E., Van Wiltenburg, J., Koets, M., & Van Amerongen, A. (2003). Neamin as an immunogen for the development of a generic ELISA detecting gentamicin, kanamycin, and neomycin in milk. Journal of Agriculture and Food Chemistry, 51, 587–593.
Su, L., Jia, W., Hou, C., & Lei, Y. (2011). Microbial biosensors: A review. Biosensor Bioelectronics, 26, 1788–1799.
Turner, A. P. F., Karube, I., & Wilson, G. S. (Eds.). (1987). Biosensors fundamentals and applications. Oxford: Oxford University Press.
Blum, L. J., & Coulet, P. R. (Eds.). (1991). Biosensor principles and applications. New York: Marcel Dekker.
Mulchandani, A., & Rogers, K. R. (Eds.). (1998). Enzyme and microbial biosensors: Techniques and protocols. Totowa: Humanae Press.
Nikolelis, D., Krull, U., Wang, J., & Mascini, M. (Eds.). (1998). Biosensors for direct monitoring of environmental pollutants in field. London: Kluwer Academic.
Ramsay, G. E. (1998). Commercial biosensors: Applications to clinical, bioprocess and environmental samples. Chichester: Wiley.
Rogers, K. R., & Mulchandani, A. (1998). Affinity biosensors: Techniques and protocols. Totowa: Humanae Press.
D’Souza, S. F. (2001). Microbial biosensors. Biosensors Bioelectronics, 16, 337–353.
Weber, C. C., Link, N., Fux, C., Zisch, A. H., Weber, W., & Fussenegger, M. (2005). Broad-spectrum protein biosensors for class-specific detection of antibiotics. Biotechnology and Bioengineering, 89, 9–17.
Sorensen, S. J., Burmolle, M., & Hansen, L. H. (2006). Making bio-sense of toxicity: New developments in whole-cell biosensors. Current Opinion in Biotechnology, 17, 11–16.
Ramos, J. L., Duque, E., Huertas, M. J., & Haidour, A. (1995). Isolation and expansion of the catabolic potential of a Pseudomonas putida strain able to grow in the presence of high concentrations of aromatic hydrocarbons. Journal of Bacteriology, 177, 3911–3916.
Mosqueda, G., & Ramos, J. L. (2000). A set of genes encoding a second toluene efflux system in Pseudomonas putida DOT-T1E is linked to the tod genes for toluene metabolism. Journal of Bacteriology, 182, 937–943.
Rojas, A., Duque, E., Mosqueda, G., Golden, G., Hurtado, A., Ramos, J. L., & Segura, A. (2001). Three efflux pumps are required to provide efficient tolerance to toluene in Pseudomonas putida DOT-T1E. Journal of Bacteriology, 183, 3967–3973.
Segura, A., Godoy, P., van Dillewijn, P., Hurtado, A., Arroyo, N., Santacruz, S., & Ramos, J. L. (2005). Proteomic analysis reveals the participation of energy- and stress-related proteins in the response of Pseudomonas putida DOT-T1E to toluene. Journal of Bacteriology, 187, 5937–5945.
Duque, E., Rodriguez-Herva, J. J., de la Torre, J., Dominguez-Cuevas, P., Munoz-Rojas, J., & Ramos, J. L. (2007). The RpoT regulon of Pseudomonas putida DOT-T1E and its role in stress endurance against solvents. Journal of Bacteriology, 189, 207–219.
Woodcock, D. M., Crowther, P. J., Doherty, J., Jefferson, S., DeCruz, E., Noyer-Weidner, M., et al. (1989). Quantitative evaluation of Escherichia coli host strains for tolerance to cytosine methylation in plasmid and phage recombinants. Nucleic Acids Research, 17(9), 3469–3478.
Lennox, E. S. (1955). Transduction of linked genetic characters of the host by bacteriophage P1. Virology, 2, 190–206.
Karunakaran, R., Mauchline, T. H., Hosie, A. H., & Poole, P. S. (2005). A family of promoter probe vectors incorporating autofluorescent and chromogenic reporter proteins for studying gene expression in Gram-negative bacteria. Microbiology, 151, 3249–3256.
Ramos, J. L., Duque, E., Gallegos, M. T., Godoy, P., Ramos-Gonzalez, M. I., Rojas, A., et al. (2002). Mechanisms of solvent tolerance in gram-negative bacteria. Annual Review of Microbiology, 56, 743–768.
Teran, W., Felipe, A., Segura, A., Rojas, A., Ramos, J. L., & Gallegos, M. T. (2003). Antibiotic-dependent induction of Pseudomonas putida DOT-T1E TtgABC efflux pump is mediated by the drug binding repressor TtgR. Antimicrobial Agents and Chemotherapy, 47, 3067–3072.
Duque, E., Segura, A., Mosqueda, G., & Ramos, J. L. (2001). Global and cognate regulators control the expression of the organic solvent efflux pumps TtgABC and TtgDEF of Pseudomonas putida. Molecular Microbiology, 39, 1100–1106.
Teran, W., Krell, T., Ramos, J. L., & Gallegos, M. T. (2006). Effector-repressor interactions, binding of a single effector molecule to the operator-bound TtgR homodimer mediates derepression. Journal of Biological Chemistry, 17, 7102–7109.
Guazzaroni, M. E., Teran, W., Zhang, X., Gallegos, M. T., & Ramos, J. L. (2004). TtgV bound to a complex operator site represses transcription of the promoter for the multidrug and solvent extrusion TtgGHI pump. Journal of Bacteriology, 186, 2921–2927.
Peleg, A. Y., & Hooper, D. C. (2010). Hospital-acquired infections due to gram-negative bacteria. New England Journal of Medicine, 362, 1804–1813.
Ball, P. (2000). Quinolone generations: Natural history or natural selection? Journal of Antimicrobial Chemotherapy, 46(Suppl T1), 17–24.
Falagas, M. E., Grammatikos, A. P., & Michalopoulos, A. (2008). Potential of old-generation antibiotics to address current need for new antibiotics. Expert Review of Anti-infective Therapy, 6, 593–600.
Alguel, Y., Meng, C., Teran, W., Krell, T., Ramos, J. L., Gallegos, M. T., & Zhang, X. (2007). Crystal structures of multidrug binding protein TtgR in complex with antibiotics and plant antimicrobials. Journal of Molecular Biology, 369, 829–840.
Acknowledgments
We are grateful to M.D. Jesús Villar for the information provided on the antibiotics most commonly used in Hospitals against Pseudomonas. We thank Marisa Cañadas-Garre and Virgen de las Nieves Hospital for providing the antibiotics of clinical use. This work was supported by Spanish Ministry of Economy and Competitiveness, National Programme for Recruitment and Incorporation of Human Resources, Subprogramme: Ramon y Cajal RYC-2009-04570 and grant P11-CVI-7391 from Junta de Andalucía and EFDR.
Conflict of interest
The authors declare no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
12033_2015_9849_MOESM1_ESM.tif
Supplementary material 1 (TIFF 5954 kb). Fig. S1. Toxic respond to toluene of the biosensor, via GFP expression, under UV-light illumination. A) In absence of toluene; B) In the presence of vapour phase of toluene
Rights and permissions
About this article
Cite this article
Espinosa-Urgel, M., Serrano, L., Ramos, J.L. et al. Engineering Biological Approaches for Detection of Toxic Compounds: A New Microbial Biosensor Based on the Pseudomonas putida TtgR Repressor. Mol Biotechnol 57, 558–564 (2015). https://doi.org/10.1007/s12033-015-9849-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-015-9849-2