Abstract
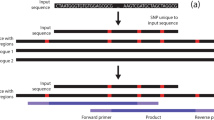
Trinucleotide repeats are common within gene coding regions and could serve as beacons to locate genes. Five of the most common trinucleotide repeats in an Actinidia (kiwifruit) expressed sequence tag (EST) database were found to be (ACC)4, (CAC)4, (CCA)4, (CTC)4, and (TGG)4. These repeats, with or without an artificial 5′-end tail, were tested by vectorette PCR against genomic DNA from Actinidia chinensis. Eighty-nine randomly selected clones showed an average insert size of 383 bp, with a maximum of 1,151 bp and a minimum of 78 bp. Two-thirds of the clones contained the artificial tail attached to the trinucleotide, showing a slight advantage of possessing such a tail during annealing and amplification. The sequences were searched against the Actinidia EST database and GenBank. Of the 89 clones, 33 had a significant hit (expect value < e−15). Twenty-four of those clones matched an Actinidia EST. Twenty-one clones contained one or more simple sequence repeats. This methodology can be applied by conventional cloning and sequencing methods or by high throughput pyrosequencing technologies to develop genetic markers and also for gene mining in species with little or no genetic/genomic resources.

Similar content being viewed by others
References
Li, Y.-C., Korol, A. B., Fahima, T., & Nevo, E. (2004). Microsatellites within genes: Structure, function, and evolution. Molecular Biology and Evolution, 21, 991–1007. doi:10.1093/molbev/msh073.
Toth, G., Gaspari, Z., & Jurka, J. (2000). Microsatellites in different eukaryotic genomes: Survey and analysis. Genome Research, 10, 967–981. doi:10.1101/gr.10.7.967.
Crowhurst, R., Gleave, A., Macrae, E., Ampomah-Dwamena, C., Atkinson, R., Beuning, L., et al. (2008). Analysis of expressed sequence tags from Actinidia: applications of a cross species EST database for gene discovery in the areas of flavor, health, color and ripening. BMC Genomics, 9, 351. doi:10.1186/1471-2164-9-351.
Morgante, M., Hanafey, M., & Powell, W. (2002). Microsatellites are preferentially associated with nonrepetitive DNA in plant genomes. Nature Genetics, 30, 194–200. doi:10.1038/ng822.
Rakoczy-Trojanowska, M., & Bolibok, H. (2004). Characteristics and comparison of three classes of microsatellite-based markers and their application in plants. Cellular & Molecular Biology Letters, 9, 221–238.
Fraser, L. G., McNeilage, M. A., Tsang, G. K., Harvey, C. F., & De Silva, H. N. (2005). Cross-species amplification of microsatellite loci within the dioecious, polyploid genus Actinidia (Actinidiaceae). Theoretical and Applied Genetics, 112, 149–157. doi:10.1007/s00122-005-0117-x.
Kuleung, C., Baenziger, P. S., & Dweikat, I. (2004). Transferability of SSR markers among wheat, rye, and triticale. Theoretical and Applied Genetics, 108, 1147–1150. doi:10.1007/s00122-003-1532-5.
Riley, J., Butler, R., Ogilvie, D., Finniear, R., Jenner, D., Powell, S., et al. (1990). A novel, rapid method for the isolation of terminal sequences from yeast artificial chromosome (YAC) clones. Nucleic Acids Research, 18, 2887–2890. doi:10.1093/nar/18.10.2887.
Lench, N. J., Norris, A., Bailey, A., Booth, A., & Markham, A. F. (1996). Vectorette PCR isolation of microsatellite repeat sequences using anchored dinucleotide repeat primers. Nucleic Acids Research, 24, 2190–2191. doi:10.1093/nar/24.11.2190.
Ujino, T., Kawahara, T., Tsumura, Y., Nagamitsu, T., Yoshimaru, H., & Ratnam, W. (1998). Development and polymorphism of simple sequence repeat DNA markers for Shorea curtisii and other Dipterocarpaceae species. Heredity, 81, 422–428. doi:10.1046/j.1365-2540.1998.00423.x.
Wang, X. W., Trigiano, R. N., Windham, M. T., Devries, R. E., Scheffler, B. E., Rinehart, T. A., et al. (2007). A simple PCR procedure for discovering microsatellites from small insert libraries. Molecular Ecology Notes, 7, 558–561. doi:10.1111/j.1471-8286.2006.01655.x.
Hayden, M. J., & Sharp, P. J. (2001). Sequence-tagged microsatellite profiling (STMP): A rapid technique for developing SSR markers. Nucleic Acids Research, 29, e43. doi:10.1093/nar/29.8.e43.
Ko, W.-Y., David, R., & Akashi, H. (2003). Molecular phylogeny of the Drosophila melanogaster species subgroup. Journal of Molecular Evolution, 57, 562–573. doi:10.1007/s00239-003-2510-x.
Brownie, J., Shawcross, S., Theaker, J., Whitcombe, D., Ferrie, R., Newton, C., et al. (1997). The elimination of primer-dimer accumulation in PCR. Nucleic Acids Research, 25, 3235–3241. doi:10.1093/nar/25.16.3235.
Sambrook, J., & Russell, D. W. (2001). Molecular cloning, a laboratory manual (3rd ed., Vol. 1). Cold Spring Harbor, NY: Cold Spring Harbor Press.
Rudd, S., Mewes, H.-W., & Mayer, K. F. X. (2003). Sputnik: A database platform for comparative plant genomics. Nucleic Acids Research, 31, 128–132. doi:10.1093/nar/gkg075.
Altschul, S. F., Madden, T. L., Schaffer, A. A., Zhang, J., Zhang, Z., Miller, W., et al. (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research, 25, 3389–3402. doi:10.1093/nar/25.17.3389.
Fraser, L. G., Harvey, C. F., Crowhurst, R. N., & De Silva, H. N. (2004). EST-derived microsatellites from Actinidia species and their potential for mapping. Theoretical and Applied Genetics, 108, 1010–1016. doi:10.1007/s00122-003-1517-4.
Celton, J., Tustin, D., Chagne, D., & Gardiner, S. (2008). Construction of a dense genetic linkage map for apple rootstocks using SSRs developed from Malus ESTs and Pyrus genomic sequences. Tree Genetics Genomics, 5, 93–107. doi:10.1007/s11295-008-017-z.
Temnykh, S., DeClerck, G., Lukashova, A., Lipovich, L., Cartinhour, S., & McCouch, S. (2001). Computational and experimental analysis of microsatellites in rice (Oryza sativa L.): Frequency, length variation, transposon associations, and genetic marker potential. Genome Research, 11, 1441–1452. doi:10.1101/gr.184001.
Acknowledgements
We thank John F. Mackay, Erik Rikkerink, Paul Datson, and Anne Gunson for critical review of the manuscript, and Ross N. Crowhurst for the bioinformatics support. This work funded by HortResearch internal funding.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hilario, E., Fraser, L.G. & McNeilage, M. Trinucleotide Repeats as Bait for Vectorette PCR: A Tool for Developing Genetic Mapping Markers. Mol Biotechnol 42, 320–326 (2009). https://doi.org/10.1007/s12033-009-9157-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-009-9157-9