Abstract
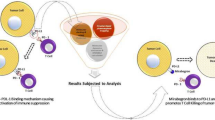
Immunotherapies are promising therapeutic options for the management of triple-negative breast cancer because of its high mutation rate and genomic instability. Of note, the blockade of the immune checkpoint protein PD-1 and its ligand PD-L1 has been proven to be an efficient and potent strategy to combat triple-negative breast cancer. To date, various anti-PD-1/anti-PD-L1 antibodies have been approved. However, the intrinsic constraints of these therapeutic antibodies significantly limit their application, making small molecules a potentially significant option for PD-1/PD-L1 inhibition. In light of this, the current study aims to use a high-throughput virtual screening technique to identify potential repurposed candidates as PD-L1 inhibitors. Thus, the present study explored binding efficiency of 2509 FDA-approved compounds retrieved from the drug bank database against PD-L1 protein. The binding affinity of the compounds was determined using the glide XP docking programme. Furthermore, prime-MM/GBSA, DFT calculations, and RF score were used to precisely re-score the binding free energy of the docked complexes. In addition, the ADME and toxicity profiles for the lead compounds were also examined to address PK/PD characteristics. Altogether, the screening process identified three molecules, namely DB01238, DB06016 and DB01167 as potential therapeutics for the PD-L1 protein. To conclude, a molecular dynamic simulation of 100 ns was run to characterise the stability and inhibitory action of the three lead compounds. The results from the simulation study confirm the robust structural and thermodynamic stability of DB01238 than other investigated molecules. Thus, our findings hypothesize that DB01238 could serve as potential PD-L1 inhibitor in the near future for triple-negative breast cancer patients.






Similar content being viewed by others
Data availability
All data to support the conclusions have been either provided or are otherwise publicly available.
Abbreviations
- PD-1:
-
Programmed cell death 1
- PD-L1:
-
Programmed cell death ligand 1
- BMS:
-
Bristol-Myers Squibb
- TME:
-
Tumour micro environment
- TIL:
-
Tumour infiltrating lymphocytes
- TMB:
-
Tumour mutation burden
- ADME:
-
Absorption, distribution, metabolism and excretion
- DFT:
-
Density functional theory
- FMO:
-
Frontier molecular orbitals
- PF/PD:
-
Pharmacokinetics/pharmacodynamics
- RMSD:
-
Root mean square deviation
- RMSF:
-
Root mean square fluctuation
- FEL:
-
Free energy landscape
References
Korlimarla A, Hari PS, Prabhu J, Ragulan C, Patil Y, Snijesh VP, Desai K, Mathews A, Appachu S, Diwakar RB, Srinath BS. Comprehensive characterization of immune landscape of Indian and Western triple negative breast cancers. Transl Oncol. 2022. https://doi.org/10.1016/j.tranon.2022.101511.
Loizides S, Constantinidou A. Triple negative breast cancer: Immunogenicity, tumor microenvironment, and immunotherapy. Front Genet. 2023. https://doi.org/10.3389/fgene.2022.1095839.
Qiu D, Zhang G, Yan X, Xiao X, Ma X, Lin S, Wu J, Li X, Wang W, Liu J, Ma Y. Prospects of immunotherapy for triple-negative breast cancer. Front Oncol. 2022. https://doi.org/10.3389/fonc.2021.797092.
Yi H, Li Y, Tan Y, Fu S, Tang F, Deng X. Immune checkpoint inhibition for triple-negative breast cancer: current landscape and future perspectives. Front Oncol. 2021. https://doi.org/10.3389/fonc.2021.648139.
Luo C, Wang P, He S, Zhu J, Shi Y, Wang J. Progress and prospect of immunotherapy for triple-negative breast cancer. Front Oncol. 2022. https://doi.org/10.3389/fonc.2022.919072.
Wang F, Ye W, Wang S, He Y, Zhong H, Wang Y, Zhu Y, Han J, Bing Z, Ji S, Liu H. Discovery of a new inhibitor targeting PD-L1 for cancer immunotherapy. Neoplasia. 2021. https://doi.org/10.1016/j.neo.2021.01.001.
Mittendorf EA, Philips AV, Meric-Bernstam F, Qiao N, Wu Y, Harrington S, Su X, Wang Y, Gonzalez-Angulo AM, Akcakanat A, Chawla A. PD-L1 expression in triple-negative breast cancer. Cancer Immunol Res. 2014. https://doi.org/10.1158/2326-6066.CIR-13-0127.
Bharadwa KR, Dasgupta K, Narayana SM, Ramachandra C, Babu SM, Rangarajan A, Kumar RV. PD-1 and PD-L1 expression in Indian women with breast cancer. Eur J Breast Health. 2022. https://doi.org/10.4274/ejbh.galenos.2021.2021-5-2.
Schmid P, Rugo HS, Adams S, Schneeweiss A, Barrios CH, Iwata H, Diéras V, Henschel V, Molinero L, Chui SY, Maiya V. Atezolizumab plus nab-paclitaxel as first-line treatment for unresectable, locally advanced or metastatic triple-negative breast cancer (IMpassion130): updated efficacy results from a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol. 2020. https://doi.org/10.1016/S1470-2045(19)30689-8.
Emens LA, Adams S, Barrios CH, Diéras V, Iwata H, Loi S, Rugo HS, Schneeweiss A, Winer EP, Patel S, Henschel V. First-line atezolizumab plus nab-paclitaxel for unresectable, locally advanced, or metastatic triple-negative breast cancer: IMpassion130 final overall survival analysis. Ann Oncol. 2021. https://doi.org/10.1016/j.annonc.2021.05.355.
Wang Y, Guo H, Feng Z, Wang S, Wang Y, He Q, Li G, Lin W, Xie XQ, Lin Z. PD-1-targeted discovery of peptide inhibitors by virtual screening, molecular dynamics simulation, and surface plasmon resonance. Molecules. 2019. https://doi.org/10.3390/molecules24203784.
Zak KM, Grudnik P, Guzik K, Zieba BJ, Musielak B, Dömling A, Dubin G, Holak TA. Structural basis for small molecule targeting of the programmed death ligand 1 (PD-L1). Oncotarget. 2016. https://doi.org/10.18632/oncotarget.8730.
Skalniak L, Zak KM, Guzik K, Magiera K, Musielak B, Pachota M, Szelazek B, Kocik J, Grudnik P, Tomala M, Krzanik S. Small-molecule inhibitors of PD-1/PD-L1 immune checkpoint alleviate the PD-L1-induced exhaustion of T-cells. Oncotarget. 2017. https://doi.org/10.18632/oncotarget.20050.
Donnelly DJ, Smith RA, Morin P, Lipovšek D, Gokemeijer J, Cohen D, Lafont V, Tran T, Cole EL, Wright M, Kim J. Synthesis and biologic evaluation of a novel 18F-labeled adnectin as a PET radioligand for imaging PD-L1 expression. J Nucl Med. 2018. https://doi.org/10.2967/jnumed.117.199596.
Li CW, Lim SO, Chung EM, Kim YS, Park AH, Yao J, Cha JH, Xia W, Chan LC, Kim T, Chang SS. Eradication of triple-negative breast cancer cells by targeting glycosylated PD-L1. Cancer Cell. 2018. https://doi.org/10.1016/j.ccell.2018.01.009.
Guzik K, Zak KM, Grudnik P, Magiera K, Musielak B, Torner R, Skalniak L, Domling A, Dubin G, Holak TA. Small-molecule inhibitors of the programmed cell death-1/programmed death-ligand 1 (PD-1/PD-L1) interaction via transiently induced protein states and dimerization of PD-L1. J Med Chem. 2017. https://doi.org/10.1021/acs.jmedchem.7b00293.
Wu X, Meng Y, Liu L, Gong G, Zhang H, Hou Y, Liu C, Wu D, Qin M. Insights into non-peptide small-molecule inhibitors of the PD-1/PD-L1 interaction: development and perspective. Bioorg Med Chem. 2021. https://doi.org/10.1016/j.bmc.2021.116038.
Hussain A, Hussain A, Sabnam N, Verma CK, Shrivastava N. Insilico exploration of the potential inhibitory activity of DrugBank compounds against CDK7 kinase using structure-based virtual screening, molecular docking, and dynamics simulation approach. Arab J Chem. 2023. https://doi.org/10.1016/j.arabjc.2022.104460.
Mittal L, Tonk RK, Awasthi A, Asthana S. Targeting cryptic-orthosteric site of PD-L1 for inhibitor identification using structure-guided approach. Arch Biochem Biophys. 2021. https://doi.org/10.1016/j.abb.2021.109059.
James N, Shanthi V, Ramanathan K. Density functional theory and molecular simulation studies for prioritizing anaplastic lymphoma kinase inhibitors. Biotechnol Appl Biochem. 2020. https://doi.org/10.1007/s12010-019-03156-1.
Sandor M, Kiss R, Keserű GM. Virtual fragment docking by Glide: a validation study on 190 protein−fragment complexes. J Chem Inf Model. 2010. https://doi.org/10.1021/ci1000407.
Jang C, Yadav DK, Subedi L, Venkatesan R, Venkanna A, Afzal S, Lee E, Yoo J, Ji E, Kim SY, Kim MH. Identification of novel acetylcholinesterase inhibitors designed by pharmacophore-based virtual screening, molecular docking and bioassay. Sci Rep. 2018. https://doi.org/10.1038/s41598-018-33354-6.
Ramesh P, Veerappapillai S. Designing novel compounds for the treatment and management of RET-positive non-small cell lung cancer—fragment based drug design strategy. Molecules. 2022. https://doi.org/10.3390/molecules27051590.
Wójcikowski M, Ballester PJ, Siedlecki P. Performance of machine-learning scoring functions in structure-based virtual screening. Sci Rep. 2017. https://doi.org/10.1038/srep46710.
Li H, Lu G, Sze KH, Su X, Chan WY, Leung KS. Machine-learning scoring functions trained on complexes dissimilar to the test set already outperform classical counterparts on a blind benchmark. Brief Bioinform. 2021. https://doi.org/10.1093/bib/bbab225.
Li H, Leung KS, Wong MH, Ballester PJ. Improving AutoDock Vina using random forest: the growing accuracy of binding affinity prediction by the effective exploitation of larger data sets. Mol Inform. 2015. https://doi.org/10.1002/minf.201400132.
Leeson PD, Young RJ. Molecular property design: does everyone get it? ACS Med Chem Lett. 2015. https://doi.org/10.1021/acsmedchemlett.5b00157.
Banerjee P, Eckert AO, Schrey AK, Preissner R. ProTox-II: a webserver for the prediction of toxicity of chemicals. Nucleic Acids Res. 2018. https://doi.org/10.1093/nar/gky318.
Aziz M, Ejaz SA, Zargar S, Akhtar N, Aborode AT, Wani TA, Batiha GE, Siddique F, Alqarni M, Akintola AA. Deep learning and structure-based virtual screening for drug discovery against NEK7: a novel target for the treatment of cancer. Molecules. 2022. https://doi.org/10.3390/molecules27134098.
Vanommeslaeghe K, Hatcher E, Acharya C, Kundu S, Zhong S, Shim J, Darian E, Guvench O, Lopes P, Vorobyov I, Mackerell AD Jr. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J Comput Chem. 2010. https://doi.org/10.1002/jcc.21367.
Wang H, Dommert F, Holm C. Optimizing working parameters of the smooth particle mesh Ewald algorithm in terms of accuracy and efficiency. J Chem Phys. 2010. https://doi.org/10.1063/1.3446812.
Allen WJ, Rizzo RC. Implementation of the Hungarian algorithm to account for ligand symmetry and similarity in structure-based design. J Chem Inform Model. 2014. https://doi.org/10.1021/ci400534h.
Abdullahi SH, Uzairu A, Shallangwa GA, Uba S, Umar AB. In-silico activity prediction, structure-based drug design, molecular docking and pharmacokinetic studies of selected quinazoline derivatives for their antiproliferative activity against triple negative breast cancer (MDA-MB231) cell line. Bull Natl Res Cent. 2022. https://doi.org/10.1186/s42269-021-00690-z.
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA, Sanschagrin PC, Mainz DT. Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein−ligand complexes. J Med Chem. 2006. https://doi.org/10.1021/jm051256o.
Azam F, Eid EE, Almutairi A. Targeting SARS-CoV-2 main protease by teicoplanin: a mechanistic insight by docking, MM/GBSA and molecular dynamics simulation. J Mol Struct. 2021. https://doi.org/10.1016/j.molstruc.2021.131124.
Genheden S, Ryde U. The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opin Drug Discov. 2015. https://doi.org/10.1517/17460441.2015.1032936.
Madhavaram M, Nampally V, Gangadhari S, Palnati MK, Tigulla P. High-throughput virtual screening, ADME analysis, and estimation of MM/GBSA binding-free energies of azoles as potential inhibitors of Mycobacterium tuberculosis H37Rv. J Recept Signal Transduct Res. 2019. https://doi.org/10.1080/10799893.2019.1660895.
Basavarajappa GM, Rehman A, Shiroorkar PN, Sreeharsha N, Anwer MK, Aloufi B. Therapeutic effects of Crataegus monogyna inhibitors against breast cancer. Front Pharmacol. 2023. https://doi.org/10.3389/fphar.2023.1187079.
Saba A, Sarwar F, Muhammad S, Ilyas M, Iqbal J, Al-Sehemi AG, Ayub K, Gilani MA, Adnan M. Insighting the inhibitory potential of novel modafinil drug derivatives against estrogen alpha (ERα) of breast cancer through a triple hybrid computational methodology. J Mol Liq. 2022. https://doi.org/10.1016/j.molliq.2022.120234.
Singh N, Chaput L, Villoutreix BO. Fast rescoring protocols to improve the performance of structure-based virtual screening performed on protein–protein interfaces. J Chem Inf Model. 2020. https://doi.org/10.1021/acs.jcim.0c00545.
Thirunavukkarasu MK, Suriya U, Rungrotmongkol T, Karuppasamy R. In silico screening of available drugs targeting non-small cell lung cancer targets: a drug repurposing approach. Pharmaceutics. 2021. https://doi.org/10.3390/pharmaceutics14010059.
Velasquez-López Y, Tejera E, Perez-Castillo Y. Can docking scoring functions guarantee success in virtual screening? Annu Rep Med Chem. 2022. https://doi.org/10.1016/bs.armc.2022.08.008.
Phengsakun G, Boonyarit B, Rungrotmongkol T, Suginta W. Structure-based virtual screening for potent inhibitors of GH-20 β-N-acetylglucosaminidase: classical and machine learning scoring functions, and molecular dynamics simulations. Comput Biol Chem. 2023. https://doi.org/10.1016/j.compbiolchem.2023.107856.
Al-Ghorbani M, Gouda MA, Baashen M, Alharbi O, Almalki FA, Ranganatha LV. Piperazine heterocycles as potential anticancer agents: a review. Pharm Chem J. 2022. https://doi.org/10.1007/s11094-022-02597-z.
Ruzic D, Ellinger B, Djokovic N, Santibanez JF, Gul S, Beljkas M, Djuric A, Ganesan A, Pavic A, Srdic-Rajic T, Petkovic M. Discovery of 1-benzhydryl-piperazine-based HDAC inhibitors with anti-breast cancer activity: synthesis, molecular modeling, in vitro and in vivo biological evaluation. Pharmaceutics. 2022. https://doi.org/10.3390/pharmaceutics14122600.
Long H, Hu X, Wang B, Wang Q, Wang R, Liu S, Xiong F, Jiang Z, Zhang XQ, Ye WC, Wang H. Discovery of Novel apigenin-piperazine hybrids as potent and selective poly (ADP-ribose) polymerase-1 (PARP-1) inhibitors for the treatment of cancer. J Med Chem. 2021. https://doi.org/10.1021/acs.jmedchem.1c00735.
Akkoç MK, Yüksel MY, Durmaz I, Atalay RÇ. Design, synthesis, and biological evaluation of indole-based 1, 4-disubstituted piperazines as cytotoxic agents. Turk J Chem. 2012. https://doi.org/10.3906/kim-1111-5.
Londhe AM, Gadhe CG, Lim SM, Pae AN. Investigation of molecular details of Keap1-Nrf2 inhibitors using molecular dynamics and umbrella sampling techniques. Molecules. 2019. https://doi.org/10.3390/molecules24224085.
Radwan A, Mahrous GM. Docking studies and molecular dynamics simulations of the binding characteristics of waldiomycin and its methyl ester analog to Staphylococcus aureus histidine kinase. PLoS ONE. 2020. https://doi.org/10.1371/journal.pone.0234215.
Ramesh P, Shin WH, Veerappapillai S. Discovery of a potent candidate for ret-specific non-small-cell lung cancer—a combined in silico and in vitro strategy. Pharmaceutics. 2021. https://doi.org/10.3390/pharmaceutics13111775.
Swapna LS, Bhaskara RM, Sharma J, Srinivasan N. Roles of residues in the interface of transient protein-protein complexes before complexation. Sci Rep. 2012. https://doi.org/10.1038/srep00334.
Mosquera-Yuqui F, Lopez-Guerra N, Moncayo-Palacio EA. Targeting the 3CLpro and RdRp of SARS-CoV-2 with phytochemicals from medicinal plants of the Andean Region: molecular docking and molecular dynamics simulations. J Biomol Struct Dyn. 2022. https://doi.org/10.1080/07391102.2020.1835716.
Chatterjee P, Karn R, Emerson IA, Banerjee S. Docking and molecular dynamics simulation revealed the potential inhibitory activity of amygdalin in triple-negative breast cancer therapeutics targeting the BRCT domain of BARD1 receptor. Mol Biotechnol. 2023. https://doi.org/10.1007/s12033-023-00680-8.
Baig MH, Sudhakar DR, Kalaiarasan P, Subbarao N, Wadhawa G, Lohani M, Khan MK, Khan AU. Insight into the effect of inhibitor resistant S130G mutant on physico-chemical properties of SHV type beta-lactamase: a molecular dynamics study. PLoS ONE. 2014. https://doi.org/10.1371/journal.pone.0112456.
Acknowledgements
The authors thank the management of Vellore Institute of Technology for providing the facilities to carry out this research work.
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Contributions
HRK performed the computational work, analysed the data and prepared tables and figures. RK conceived this study and is responsible for the overall design, interpretation, manuscript preparation and communication. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Krishnamoorthy, H.R., Karuppasamy, R. A multitier virtual screening of antagonists targeting PD-1/PD-L1 interface for the management of triple-negative breast cancer. Med Oncol 40, 312 (2023). https://doi.org/10.1007/s12032-023-02183-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-023-02183-7