Abstract
Alzheimer’s disease (AD) is a progressive cognitive disorder that occurs worldwide, and the lack of disease-modifying targets and pathways is a pressing issue. This study aimed to provide new targets and pathways by performing molecular subgroup classification. After normalizing the collected data, the subgroup number was confirmed with consensus clustering. Comparisons of clinical features among subgroups were conducted to clarify the clinical traits of each subgroup. Subgroup-specific genes were identified to perform weighted gene coexpression analysis (WGCNA). Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses were carried out. Next, gene set enrichment analysis (GSEA) was performed. Protein–protein interaction networks were built to screen core genes and in each subgroup to perform Spearman correlation analysis with clinical traits. Sequencing profiles of 1068 AD samples collected from 2 datasets were classified into 3 subgroups. Clinical comparisons revealed that patients in subgroup III tended to be younger, while their pathological grades were the most severe. WGCNA detected four gene modules, and the turquoise module, where the dopaminergic synapse pathway was enriched, was related to subgroup I. The neurotrophin signaling pathway and TGF-beta signaling pathway were robustly enriched in the blue and brown modules, respectively, in subgroup III. Moreover, 3 hub genes in subgroup I were negatively correlated with the sum of neurofibrillary tangle (Nft) density. Conversely, hub genes in subgroups II and III exhibited positive correlations with the sum of Nft density. These results provide new pathways and targets for AD treatment.
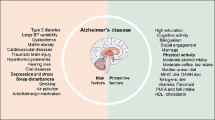
Graphic Abstract








Similar content being viewed by others
Availability of Data and Materials
All data and materials used in this work were publicly available and also available based on request.
References
2021 (2021) Alzheimer’s disease facts and figures. Alzheimers Dement 17:327–406
Akram A, Schmeidler J, Katsel P et al (2010) Increased expression of cholesterol transporter ABCA1 is highly correlated with severity of dementia in AD hippocampus. Brain Res 1318:167–177
Armstrong R (2020) Fluctuations in neurofibrillary tangle density in Alzheimer’s disease revealed by Fourier (spectral) analysis. Folia Neuropathol 58:299–306
Ashford JW (2019) The dichotomy of Alzheimer’s disease pathology: amyloid-β and Tau. Journal of Alzheimer’s Disease : JAD 68:77–83
Ballard C, Gauthier S, Corbett A et al (2011) Alzheimer’s disease. Lancet (london, England) 377:1019–1031
Barel G, Herwig R (2020) NetCore: a network propagation approach using node coreness. Nucleic Acids Res 48:e98
Barton AJ, Pearson RC, Najlerahim A et al (1993) Pre- and postmortem influences on brain RNA. J Neurochem 61
Bongiorno D, Schuetz F, Poronnik P et al (2011) Regulation of voltage-gated ion channels in excitable cells by the ubiquitin ligases Nedd4 and Nedd4–2. Channels 5:79–88
Bosse T, Nout RA, McAlpine JN et al (2018) Molecular classification of grade 3 endometrioid endometrial cancers identifies distinct prognostic subgroups. Am J Surg Pathol 42:561–568
Bowen DM, Smith CB, White P et al (1976) Neurotransmitter-related enzymes and indices of hypoxia in senile dementia and other abiotrophies. Brain : a Journal of Neurology 99:459–496
Braak H, Alafuzoff I, Arzberger T et al (2006) Staging of Alzheimer disease-associated neurofibrillary pathology using paraffin sections and immunocytochemistry. Acta Neuropathol 112:389–404
Braak H, Braak E (1991) Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol 82:239–259
Cao J, Zhong MB, Toro CA et al (2019) Endo-lysosomal pathway and ubiquitin-proteasome system dysfunction in Alzheimer’s disease pathogenesis. Neurosci Lett 703:68–78
Chandler M, Lacritz L, Hynan L et al (2005) A total score for the CERAD neuropsychological battery. Neurology 65:102–106
Chen C, Lu M, Lin S et al (2020a) The nuclear gene rpl18 regulates erythroid maturation via JAK2-STAT3 signaling in zebrafish model of Diamond-Blackfan anemia. Cell Death Dis 11:135
Chen M, Gao Y-T, Li W-X et al (2020b) FBW7 protects against spinal cord injury by mitigating inflammation-associated neuronal apoptosis in mice. Biochem Biophys Res Commun 532:576–583
Chen W, Zhuang J, Wang PP et al (2019) DNA methylation-based classification and identification of renal cell carcinoma prognosis-subgroups. Cancer Cell Int 19:185
Chen Y, Sun F, Zhang L et al (2021) miR-499a inhibits the proliferation and apoptosis of prostate cancer via targeting UBE2V2. World J Surg Oncol 19:250
Chin C-H, Chen S-H, Wu H-H et al (2014) cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8(Suppl 4):S11
Chou P-S, Wu M-N, Yang C-C et al (2019) Effect of advanced glycation end products on the progression of Alzheimer’s disease. J Alzheimers Dis 72:191–197
DeTure MA, Dickson DW (2019) The neuropathological diagnosis of Alzheimer’s disease. Mol Neurodegener 14:32
Dong X, Li S, Chen J et al (2020) Association of dietary ω-3 and ω-6 fatty acids intake with cognitive performance in older adults: National Health and nutrition examination Survey (NHANES) 2011–2014. Nutr J 19:25
Dumbacher M, Van Dooren T, Princen K et al (2018) Modifying Rap1-signalling by targeting Pde6δ is neuroprotective in models of Alzheimer’s disease. Mol Neurodegener 13:50
Ertel A, Verghese A, Byers SW et al (2006) Pathway-specific differences between tumor cell lines and normal and tumor tissue cells. Mol Cancer 5:55
Gibson GE, Luchsinger JA, Cirio R et al (2020) Benfotiamine and cognitive decline in Alzheimer’s disease: results of a randomized placebo-controlled phase IIa clinical trial. J Alzheimer's Dis 78
Gilbert JM, Brown BA, Strocchi P et al (1981) The preparation of biologically active messenger RNA from human postmortem brain tissue. J Neurochem 36:976–984
Harris LD, Jasem S, Licchesi JDF (2020) The ubiquitin system in Alzheimer’s disease. Adv Exp Med Biol 1233:195–221
Hauber AB, Johnson FR, Fillit H et al (2009) Older Americans’ risk-benefit preferences for modifying the course of Alzheimer disease. Alzheimer Dis Assoc Disord 23:23–32
Hodson R (2018) Alzheimer’s disease. Nature 559:S1
Hua Z-D, Liu X-B, Sheng J-H et al (2021) UBE2V2 positively correlates with PD-L1 expression and confers poor patient survival in lung adenocarcinoma. Applied Immunohistochemistry & Molecular Morphology : AIMM 29:585–591
Huang Z, Zhao J, Wang W et al (2020) Depletion of LncRNA NEAT1 rescues mitochondrial dysfunction through NEDD4L-dependent PINK1 degradation in animal models of Alzheimer’s disease. Front Cell Neurosci 14:28
Irwin M, Tare M, Singh A et al (2020) A positive feedback loop of hippo- and c-Jun-amino-terminal kinase signaling pathways regulates amyloid-beta-mediated neurodegeneration. Front Cell Dev Biol 8:117
Ivashko-Pachima Y, Hadar A, Grigg I et al (2021) Discovery of autism/intellectual disability somatic mutations in Alzheimer’s brains: mutated ADNP cytoskeletal impairments and repair as a case study. Mol Psychiatry 26:1619–1633
Joe E, Ringman JM (2019) Cognitive symptoms of Alzheimer’s disease: clinical management and prevention. BMJ 367:l6217
John A, Reddy PH (2021) Synaptic basis of Alzheimer’s disease: focus on synaptic amyloid beta, P-tau and mitochondria. Ageing Res Rev 65:101208
Johnson SA, McNeill T, Cordell B et al (1990) Relation of neuronal APP-751/APP-695 mRNA ratio and neuritic plaque density in Alzheimer’s disease. Science 248:854–857
Koepsell TD, Kurland BF, Harel O et al (2008) Education, cognitive function, and severity of neuropathology in Alzheimer disease. Neurology 70:1732–1739
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinform 9:559
Li GM, Zhang CL, Rui RP et al (2018) Bioinformatics analysis of common differential genes of coronary artery disease and ischemic cardiomyopathy. Eur Rev Med Pharmacol Sci 22:3553–3569
Lumpkin RJ, Baker RW, Leschziner AE et al (2020) Structure and dynamics of the ASB9 CUL-RING E3 Ligase. Nat Commun 11:2866
Luo Q, Vögeli TA (2020) A methylation-based reclassification of bladder cancer based on immune cell genes. Cancers 12
Merenlender-Wagner A, Malishkevich A, Shemer Z et al (2015) Autophagy has a key role in the pathophysiology of schizophrenia. Mol Psychiatry 20:126–132
Miyashita A, Hatsuta H, Kikuchi M et al (2014) Genes associated with the progression of neurofibrillary tangles in Alzheimer’s disease. Transl Psychiatry 4:e396
Mulder J, Zilberter M, Pasquaré SJ et al (2011) Molecular reorganization of endocannabinoid signalling in Alzheimer’s disease. Brain J Neurol 134:1041–1060
Orzan F, Pagani F, Cominelli M et al (2020) A simplified integrated molecular and immunohistochemistry-based algorithm allows high accuracy prediction of glioblastoma transcriptional subtypes. Lab Invest 100:1330–1344
Paschall JE, Oleksiak MF, VanWye JD et al (2004) FunnyBase: a systems level functional annotation of Fundulus ESTs for the analysis of gene expression. BMC Genomics 5:96
Peng XY, Wang Y, Hu H et al (2019) Identification of the molecular subgroups in coronary artery disease by gene expression profiles. J Cell Physiol
Piras IS, Krate J, Delvaux E et al (2019) Association of AEBP1 and NRN1 RNA expression with Alzheimer’s disease and neurofibrillary tangle density in middle temporal gyrus. Brain Res 1719:217–224
Qian J, Hyman BT, Betensky RA (2017) Neurofibrillary tangle stage and the rate of progression of Alzheimer symptoms: modeling using an autopsy cohort and application to clinical trial design. JAMA Neurol 74:540–548
Qin XY, Cao C, Cawley NX et al (2017) Decreased peripheral brain-derived neurotrophic factor levels in Alzheimer’s disease: a meta-analysis study (N=7277). Mol Psychiatry 22:312–320
Roalf DR, Moberg PJ, Xie SX et al (2013) Comparative accuracies of two common screening instruments for classification of Alzheimer’s disease, mild cognitive impairment, and healthy aging. Alzheimers Dement 9:529–537
Sebastian Monasor L, Müller SA, Colombo AV et al (2020) Fibrillar Aβ triggers microglial proteome alterations and dysfunction in Alzheimer mouse models. eLife 9
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Soleimani-Meigooni DN, Iaccarino L, La Joie R et al (2020) 18F-flortaucipir PET to autopsy comparisons in Alzheimer’s disease and other neurodegenerative diseases. Brain J Neurol 143:3477–3494
Vila-Castelar C, Guzmán-Vélez E, Pardilla-Delgado E et al (2020) Examining sex differences in markers of cognition and neurodegeneration in autosomal dominant Alzheimer’s disease: Preliminary Findings from the Colombian Alzheimer’s Prevention Initiative Biomarker Study. J Alzheimers Dis 77:1743–1753
Wang M, Roussos P, McKenzie A et al (2016) Integrative network analysis of nineteen brain regions identifies molecular signatures and networks underlying selective regional vulnerability to Alzheimer’s disease. Genome Med 8:104
Wang X, Yang P, Jiang Y et al (2021) UBE2D3 contributes to myocardial ischemia-reperfusion injury by regulating autophagy in dependence of p62/SQSTM1. Cell Signal 87:110118
Wu Z, Chen C, Kang SS et al (2021) Neurotrophic signaling deficiency exacerbates environmental risks for Alzheimer’s disease pathogenesis. Proc Natl Acad Sci USA 118
Yang Y, Yan R, Zhang L et al (2020) Primary glioblastoma transcriptome data analysis for screening survival-related genes. J Cell Biochem 121:1901–1910
Yang Y, Zhou X, Liu X et al (2021) Implications of FBXW7 in neurodevelopment and neurodegeneration: molecular mechanisms and therapeutic potential. Front Cell Neurosci 15:736008
Yu G, He Q-Y (2016) ReactomePA: an R/Bioconductor package for reactome pathway analysis and visualization. Mol BioSyst 12:477–479
Zhou ZD, Xie SP, Sathiyamoorthy S et al (2015) F-box protein 7 mutations promote protein aggregation in mitochondria and inhibit mitophagy. Hum Mol Genet 24:6314–6330
Acknowledgements
We would like to acknowledge colleagues for their helpful comments.
Funding
This work was supported by Jiangsu Planned Projects For Postdoctoral Research Funds (no. 1601056C).
Author information
Authors and Affiliations
Contributions
As the guarantor, Deqin Geng conceived the study. Sha Liu and Yan Lu initially drafted the manuscript and analyzed data.
Corresponding author
Ethics declarations
Ethics Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Disclaimer
The funding agencies had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the manuscript; or decision to submit the manuscript for publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Sha Liu and Yan Lu contributed equally to this work.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, S., Lu, Y. & Geng, D. Molecular Subgroup Classification in Alzheimer’s Disease by Transcriptomic Profiles. J Mol Neurosci 72, 866–879 (2022). https://doi.org/10.1007/s12031-021-01957-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-021-01957-w