Abstract
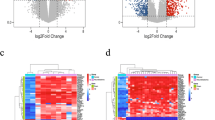
Pediatric medulloblastoma is the leading cause of cancer-related death in children. However, few studies have reported gene expression profiles of pediatric medulloblastoma and the molecular mechanism underlying this disease is unclear. To identify essential genes in pediatric medulloblastoma, we analyzed three microarray data sets from the Gene Expression Omnibus (GEO). We identified 1798 differentially expressed genes (DEGs) using the limma package. Gene set enrichment analysis demonstrated that “entrainment of circadian clock by photoperiod,” “regulation of triglyceride biosynthetic process,” and “snare complex” pathway were significantly enriched gene sets that correlated with pediatric medulloblastoma. Enriched Gene Ontology annotations of DEGs mostly included “ion-gated channel activity,” “gated channel activity,” and “channel activity.” Kyoto Encyclopedia of Genes and Genomes pathway analysis revealed that DEGs were enriched in “glutamatergic synapse,” “synaptic vesicle cycle,” and “GABAergic synapse.” Protein-protein interaction (PPI) network analysis showed that RAB5C, VAMP2, AP2M1, FNBP1, AP2A1, SYT1, SYNJ2, SYT2, HIP1R, UBB, WNT5A, SH3GL2, SYNJ1, EPN1, and DNM1 were hub genes. In conclusion, the identification of the above hub genes and pathways will help to reveal the pathogenesis of pediatric medulloblastoma and will also provide prognostic markers and therapeutic targets for pediatric medulloblastoma.






Similar content being viewed by others
References
AI-Owais MM et al (2012) Carbon monoxide mediates the anti-apoptotic effects of heme oxygenase-1 in medulloblastoma DAOY cells via K+ channel inhibition. J Biol Chem 287:24754–24764
Alejevski F et al (2019) The HisCl1 histamine receptor acts in photoreceptors to synchronize Drosophila behavioral rhythms with light-dark cycles. Nat Commun 10:252
Backes C et al (2007) GeneTrail–advanced gene set enrichment analysis. Nucleic Acids Res 35:w186–w192
Bao H, Das D, Courtney NA, Jiang Y, Briguglio JS, Lou X, Roston D, Cui Q, Chanda B, Chapman ER (2018) Dynamics and number of trans-SNARE complexes determine nascent fusion pore properties. Nature. 554:260–263
Barrett T et al (2013) NCBI GEO: archive for functional genomics data sets-update. Nucleic Acids Res 41:D991–D995
Beier CP et al (2018) Aberrant neuronal differentiation is common in glioma but is associated neither with epileptic seizures nor with better survival. Sci Rep 8:14965
Ben-Chetrit N et al (2015) Synaptojanin 2 is a druggable mediator of metastasis and the gene is overexpressed and amplified in breast cancer. Sci Signal 8:ra7
Berryer MH et al (2016) Decrease of SYNGAP1 in GABAergic cells impairs inhibitory synapse connectivity, synaptic inhibition and cognitive function. Nat Commun 7:13340
Chai C et al (2017) Metabolic circuit involving free fatty acids, microRNA122, and triglyceride synthesis in liver and muscle tissues. Gastroenterology 153:1404–1415
Chai YJ et al (2019) Comparative gene expression profiles in parathyroid adenoma and normal parathyroid tissue. J Clin Med 8:297
Chong CR et al (2013) The quest to overcome resistance to EGFR-targeted therapies in cancer. Nat Med 19:1389–1400
Dasgupta S et al (2013) SH3GL2 is frequently deleted in non-small cell lung and downregulates tumor growth by modulating EGFR signaling. J Mol Med 91:381–393
de Bont JM et al (2008) Differential expression andprognostic significance of SOX genes in pediatric medulloblastoma and ependymoma identified by microarray analysis. Neuro-Oncology 10:648–660
Dittman JS et al (2019) The control of release probability at nerve terminals. Nat Rev Neurosci 20:177–186
Farsi Z, Preobraschenski J, van den Bogaart G, Riedel D, Jahn R, Woehler A (2016) Single-vesicle imaging reveals different transport mechanisms between glutamatergic and GABAergic vesicles. Science. 351:981–984
Ferretti E, Po A (2018) Interrogating molecular data for medulloblastoma risk stratification. Lancet Oncol 19:1548–1549
Gang X et al (2017) Identification of key genes associated with rheumatoid arthritis with bioinformatics approach. Medicine 96:31
Gorello P, la Starza R, Varasano E, Chiaretti S, Elia L, Pierini V, Barba G, Brandimarte L, Crescenzi B, Vitale A, Messina M, Grammatico S, Mancini M, Matteucci C, Bardi A, Guarini A, Martelli MF, Foà R, Mecucci C (2010) Combined interphase fluorescence in situ hybridization elucidates the genetic heterogeneity of T-cell acute lymphoblastic leukemia in adults. Haematologica. 95:79–86
Kedves AT et al (2017) Recurrent ubiquitin B silencing in gynecological cancers establishes dependence on ubiquitin C. J Clin Invest 127:4554–4568
Keenan TE et al (2019) Genomic correlates of response to immune checkpoint blockade. Nat Med 25:389–402
Kool M et al (2014) Genome sequencing of SHH medulloblastoma predicts genotype–related response to smoothened inhibition. Cancer Cell 25:393–405
Korshunov A et al (2018) Molecular characterization of medulloblastomas with extensive nodularity(MBEN). Acta Neuropathol 136:303–313
Kumar V et al (2017) Challenges and recent advances in medulloblastoma therapy. Trends Pharmacol Sci 38:1061–1084
Leek JT et al (2014) svaseq: removing batch effects and other unwanted noise from sequencing data. Nucleic Acids Res 42:e161
Li P et al (2018) Key genes and integrated modules in hematopoietic differentiation of human embryonic stem cells: a comprehensive bioinformatic analysis. Stem Cell Res Ther 9:301
Liu Q et al (2018) Nanoparticle-mediated trapping of Wnt family member 5A in tumor microenvironments enhances immunotherapy for B-Raf proto-oncogene-mutant melanoma. ACS Nano 12:1250–1261
Liu S et al (2019) Identifying hub genes for heat tolerance in Water Buffalo (Bubalus bubalis) using transcriptome data. Front Genet 10:209
Mathur R et al (2018) Gene set analysis methods: a systematic comparison. BioData Mining 11:8
McNew JA et al (1999) The length of the flexible SNAREpin juxtamembrane region is a critical determinant of SNARE-dependent fusion. Mol Cell 4:415–421
Mikula M et al (2016) Genome-wide co–localization of active EGFR and downstream REK pathway kinases mirrors mitogen–inducible RNA polymerase 2 genomic occupancy. Nucleic Acids Res 44:10150–10164
Northcott PA et al (2019) Medulloblastoma. Disease Primers 5:11
Onodera Y et al (2012) Rab5c promotes AMAP1-PRKD2 complex formation to enhance β1 integrin recycling in EGF-induced cancer invasion. J Cell Biol 197:983
Pál B (2018) Involvement of extrasynaptic glutamate in Physiological and pathophysiological changes of neuronal excitability. Cell Mol Life Sci 75:2917–2949
Pasula S et al (2012) Endothelial epsin deficiency decreases tumor growth by enhancing VEGF signaling. J Clin Invest 122:4424–4438
Peng S et al (2017) paraGSEA: a scalable approach for large-scale gene expression profiling. Nucleic Acids Res 45:e155
Phi JH et al (2018) Genomic analysis reveals secondary glioblastoma after radiotherapy in a subset of recurrent medulloblastomas. Acta Neuropathol 135:939–953
Piccinin E et al (2018) Hepatic PPARγcoactivator 1βdrivers mitochondrial and anabolic signatures that contribute to hepatocellular carcinoma progression. Hepatology 67:884–898
Pilorz V et al (2014) A novel mechanism controlling resetting speed of the circadian clock to environmental stimuli. Curr Biol 24:766–773
Prada L et al (2017) Glia-to-neuron transfer of miRNAs via extracellular vesicles: a new mechanism underlying inflammation-induced synaptic alterations. Acta Neuropathol 135:529–550
Ritchie ME et al (2015) Limma powers differential expression analyses for RNA–sequencing and microarray studies. Nucleic Acids Res 43:e47
Robinson G et al (2012) Novel mutations target distinct subgroups of medulloblastoma. Nature 488:43–48
Saafan H, Foerster S, Parra-Guillen ZP, Hammer E, Michaelis M, Cinatl J Jr, Völker U, Fröhlich H, Kloft C, Ritter CA (2016) Utilising the EGFR interactome to identify mechanisms of drug resistance in non-small cell lung cancer-proof of concept towards a systems pharmacology approach. Eur J Pharm Sci 94:20–32
Salloum R et al (2019) Late morbidity and mortality among medulloblastoma survivors diagnosed across three decades: a report from the childhood cancer survivor study. J Clin Oncol 37:731–740
Servidei T et al (2009) Antitumor effect in medulloblastoma cells by gefitinib: Ectopic HER2 overexpression enhances gefitinib effects in vivo. Neuro-Oncol 11:250–259
Shukla S, Shankavaram UT, Nguyen P, Stanley BA, Smart DK (2015) Radiation–induced alteration of the brain proteome: understanding the role of the sirtuin 2 deacetylase in a murine model. J Proteome Res 14:4104–4117
Sweet-Cordero EA et al (2019) The genomic landscape of pediatric cancers: implications for diagnosis and treatment. Science 63:170–1175
Szklarczyk D et al (2018) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47:D607–D613
Teschendorf AE et al (2011) Independent surrogate variable analysis to deconvovle confounding factors in large-scale microarray profiling studies. Bioinformatics 27:1496–1505
Tsuzuki K et al (2002) Blockage of Ca2+−permeable AMPA receptors suppresses migration and induces apoptosis in human glioblastoma cells. Nat Med 8:971–978
Tuvim MJ et al (2009) Synaptotagmin 2 couples mucin granule exocytosis to Ca+ signaling from endoplasmic Reticulum. J Biol Chem 284:9781–9787
Urbina FL et al (2018) Spatiotemporal organization of exocytosis emerges during neuronal shape change. J Cell Biol 217:1113
Vasilevich AS et al (2017) How not to drown in data:a guide for biomaterial engineers. Trends Biotechnol 35:743–755
Wang J et al (2018a) Medulloblastoma:from molecular subgroups to molecular targeted therapies. Annu Rev Neurosci 41:207–232
Wang H et al (2018b) HIP1R targets PD-L1 to lysosomal degradation to alter T cell- mediated cytotoxicity. Nat Chem Biol 15:42–50
Wulff H, Christophersen P, Colussi P, Chandy KG, Yarov-Yarovoy V (2019) Antibodies and venom peptides:new modalities for ion channels. Nat Rev Drug Discov 18:339–357
Yu G et al (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS: A Journal of Integrative Biology 16:284–287
Acknowledgments
We thank Jeremy Allen, PhD, from Liwen Bianji, Edanz Group China (www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Huang, P., Guo, YD. & Zhang, HW. Identification of Hub Genes in Pediatric Medulloblastoma by Multiple-Microarray Analysis. J Mol Neurosci 70, 522–531 (2020). https://doi.org/10.1007/s12031-019-01451-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-019-01451-4