Summary
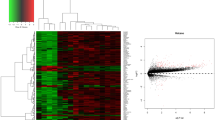
Neuroblastoma (NB) is the most common extracranial solid tumor in children. Under various treatments, some patients still have a poor prognosis. Hence, it is necessary to find new valid targets for NB therapy. In this study, a comprehensive bioinformatic analysis was used to identify differentially expressed genes (DEGs) between NB and control cells, and to select hub genes associated with NB. GSE66586 and GSE78061 datasets were downloaded from the Gene Expression Omnibus (GEO) database and DEGs were selected. Then, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses were applied to the selected DEGs. The STRING database and Cytoscape software were used to construct protein-protein interaction (PPI) networks and perform modular analysis of the DEGs. The R2 database was used for prognostic analysis. We identified a total of 238 DEGs from two microarray databases. GO enrichment analysis shows that these DEGs are mainly concentrated in the regulation of cell growth, cell migration, cell fate determination, and cell maturation. KEGG pathway analysis showed that these DEGs are mainly involved in focal adhesion, the TNF signaling pathway, cancer-related pathways, and signaling pathways regulating stem cell pluripotency. We identified the 15 most closely related DEGs from the PPI network, and performed R2 database prognostic analysis to select five hub genes – CTGF, EDN1, GATA2, LOX, and SERPINE1. This study distinguished hub genes and related signaling pathways that can potentially serve as diagnostic indicators and therapeutic biomarkers for NB, thereby improving understanding of the molecular mechanisms involved in NB.









Similar content being viewed by others
Data availability
All data is available under reasonable request.
References
Maris JM, Hogarty MD, Bagatell R, Cohn SL (2007) Neuroblastoma. Lancet 369(9579):2106–2120. https://doi.org/10.1016/S0140-6736(07)60983-0
Fonseka P, Liem M, Ozcitti C, Adda CG, Ang CS, Mathivanan S (2019) Exosomes from N-Myc amplified neuroblastoma cells induce migration and confer chemoresistance to non-N-Myc amplified cells: implications of intra-tumour heterogeneity. J Extracell Vesicles 8(1):1597614. https://doi.org/10.1080/20013078.2019.1597614
Brodeur GM (2003) Neuroblastoma: biological insights into a clinical enigma. Nat Rev Cancer 3(3):203–216. https://doi.org/10.1038/nrc1014
Scheer M, Bork K, Simon F, Nagasundaram M, Horstkorte R, Gnanapragassam VS (2020) Glycation leads to increased Polysialylation and promotes the metastatic potential of Neuroblastoma cells. Cells 9(4). https://doi.org/10.3390/cells9040868
Depuydt P, Boeva V, Hocking TD, Cannoodt R, Ambros IM, Ambros PF, Asgharzadeh S, Attiyeh EF, Combaret V, Defferrari R, Fischer M, Hero B, Hogarty MD, Irwin MS, Koster J, Kreissman S, Ladenstein R, Lapouble E, Laureys G, London WB, Mazzocco K, Nakagawara A, Noguera R, Ohira M, Park JR, Potschger U, Theissen J, Tonini GP, Valteau-Couanet D, Varesio L, Versteeg R, Speleman F, Maris JM, Schleiermacher G, De Preter K (2018) Genomic amplifications and distal 6q loss: novel markers for poor survival in high-risk Neuroblastoma patients. J Natl Cancer Inst 110(10):1084–1093. https://doi.org/10.1093/jnci/djy022
Koneru B, Lopez G, Farooqi A, Conkrite KL, Nguyen TH, Macha SJ, Modi A, Rokita JL, Urias E, Hindle A, Davidson H, McCoy K, Nance J, Yazdani V, Irwin MS, Yang S, Wheeler DA, Maris JM, Diskin SJ, Reynolds CP (2020) Telomere maintenance mechanisms define clinical outcome in high-risk neuroblastoma. Cancer Res 80:2663–2675. https://doi.org/10.1158/0008-5472.CAN-19-3068
Pinto NR, Applebaum MA, Volchenboum SL, Matthay KK, London WB, Ambros PF, Nakagawara A, Berthold F, Schleiermacher G, Park JR, Valteau-Couanet D, Pearson AD, Cohn SL (2015) Advances in risk classification and treatment strategies for Neuroblastoma. J Clin Oncol 33(27):3008–3017. https://doi.org/10.1200/JCO.2014.59.4648
Upton K, Modi A, Patel K, Kendsersky NM, Conkrite KL, Sussman RT, Way GP, Adams RN, Sacks GI, Fortina P, Diskin SJ, Maris JM, Rokita JL (2020) Epigenomic profiling of neuroblastoma cell lines. Sci Data 7(1):116. https://doi.org/10.1038/s41597-020-0458-y
Almstedt E, Elgendy R, Hekmati N, Rosen E, Warn C, Olsen TK, Dyberg C, Doroszko M, Larsson I, Sundstrom A, Arsenian Henriksson M, Pahlman S, Bexell D, Vanlandewijck M, Kogner P, Jornsten R, Krona C, Nelander S (2020) Integrative discovery of treatments for high-risk neuroblastoma. Nat Commun 11(1):71. https://doi.org/10.1038/s41467-019-13817-8
Berthold F, Faldum A, Ernst A, Boos J, Dilloo D, Eggert A, Fischer M, Fruhwald M, Henze G, Klingebiel T, Kratz C, Kremens B, Krug B, Leuschner I, Schmidt M, Schmidt R, Schumacher-Kuckelkorn R, von Schweinitz D, Schilling FH, Theissen J, Volland R, Hero B, Simon T (2020) Extended induction chemotherapy does not improve the outcome for high-risk neuroblastoma patients: results of the randomized open-label GPOH trial NB2004-HR. Ann Oncol 31(3):422–429. https://doi.org/10.1016/j.annonc.2019.11.011
Pstrag N, Ziemnicka K, Bluyssen H, Wesoly J (2018) Thyroid cancers of follicular origin in a genomic light: in-depth overview of common and unique molecular marker candidates. Mol Cancer 17(1):116. https://doi.org/10.1186/s12943-018-0866-1
Gao Y, Huo W, Zhang L, Lian J, Tao W, Song C, Tang J, Shi S, Gao Y (2019) Multiplex measurement of twelve tumor markers using a GMR multi-biomarker immunoassay biosensor. Biosens Bioelectron 123:204–210. https://doi.org/10.1016/j.bios.2018.08.060
Coyle R, Jia J, Mei Y (2016) Polymer microarray technology for stem cell engineering. Acta Biomater 34:60–72. https://doi.org/10.1016/j.actbio.2015.10.030
Toro-Dominguez D, Martorell-Marugan J, Lopez-Dominguez R, Garcia-Moreno A, Gonzalez-Rumayor V, Alarcon-Riquelme ME, Carmona-Saez P (2019) ImaGEO: integrative gene expression meta-analysis from GEO database. Bioinformatics 35(5):880–882. https://doi.org/10.1093/bioinformatics/bty721
Barrett T, Troup DB, Wilhite SE, Ledoux P, Rudnev D, Evangelista C, Kim IF, Soboleva A, Tomashevsky M, Edgar R (2007) NCBI GEO: mining tens of millions of expression profiles--database and tools update. Nucleic Acids Res 35(Database issue):D760–D765. https://doi.org/10.1093/nar/gkl887
Hart LS, Rader J, Raman P, Batra V, Russell MR, Tsang M, Gagliardi M, Chen L, Martinez D, Li Y, Wood A, Kim S, Parasuraman S, Delach S, Cole KA, Krupa S, Boehm M, Peters M, Caponigro G, Maris JM (2017) Preclinical therapeutic synergy of MEK1/2 and CDK4/6 inhibition in Neuroblastoma. Clin Cancer Res 23(7):1785–1796. https://doi.org/10.1158/1078-0432.CCR-16-1131
Gu L, Chu P, Lingeman R, McDaniel H, Kechichian S, Hickey RJ, Liu Z, Yuan YC, Sandoval JA, Fields GB, Malkas LH (2015) The mechanism by which MYCN amplification confers an enhanced sensitivity to a PCNA-derived cell permeable peptide in Neuroblastoma cells. EBioMedicine 2(12):1923–1931. https://doi.org/10.1016/j.ebiom.2015.11.016
Clough E, Barrett T (2016) The gene expression omnibus database. Methods Mol Biol 1418:93–110. https://doi.org/10.1007/978-1-4939-3578-9_5
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4(2):249–264. https://doi.org/10.1093/biostatistics/4.2.249
Gautier L, Cope L, Bolstad BM, Irizarry RA (2004) Affy--analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20(3):307–315. https://doi.org/10.1093/bioinformatics/btg405
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47. https://doi.org/10.1093/nar/gkv007
Diboun I, Wernisch L, Orengo CA, Koltzenburg M (2006) Microarray analysis after RNA amplification can detect pronounced differences in gene expression using limma. BMC Genomics 7:252. https://doi.org/10.1186/1471-2164-7-252
Nie K, Shi L, Wen Y, Pan J, Li P, Zheng Z, Liu F (2019) Identification of hub genes correlated with the pathogenesis and prognosis of gastric cancer via bioinformatics methods. Minerva Med. https://doi.org/10.23736/S0026-4806.19.06166-4
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. The gene ontology consortium. Nat Genet 25(1):25–29. https://doi.org/10.1038/75556
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30. https://doi.org/10.1093/nar/28.1.27
Ogata H, Goto S, Sato K, Fujibuchi W, Bono H, Kanehisa M (1999) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 27(1):29–34. https://doi.org/10.1093/nar/27.1.29
Sherman BT, da Huang W, Tan Q, Guo Y, Bour S, Liu D, Stephens R, Baseler MW, Lane HC, Lempicki RA (2007) DAVID knowledgebase: a gene-centered database integrating heterogeneous gene annotation resources to facilitate high-throughput gene functional analysis. BMC Bioinformatics 8:426. https://doi.org/10.1186/1471-2105-8-426
Doncheva NT, Morris JH, Gorodkin J, Jensen LJ (2019) Cytoscape StringApp: network analysis and visualization of proteomics data. J Proteome Res 18(2):623–632. https://doi.org/10.1021/acs.jproteome.8b00702
Park JA, Cheung NV (2020) Targets and antibody formats for immunotherapy of Neuroblastoma. J Clin Oncol:JCO1901410. https://doi.org/10.1200/JCO.19.01410
Xia XQ, Jia Z, Porwollik S, Long F, Hoemme C, Ye K, Muller-Tidow C, McClelland M, Wang Y (2010) Evaluating oligonucleotide properties for DNA microarray probe design. Nucleic Acids Res 38(11):e121. https://doi.org/10.1093/nar/gkq039
Pounds S, Morris SW (2003) Estimating the occurrence of false positives and false negatives in microarray studies by approximating and partitioning the empirical distribution of p-values. Bioinformatics 19(10):1236–1242. https://doi.org/10.1093/bioinformatics/btg148
Kupfer P, Guthke R, Pohlers D, Huber R, Koczan D, Kinne RW (2012) Batch correction of microarray data substantially improves the identification of genes differentially expressed in rheumatoid arthritis and osteoarthritis. BMC Med Genet 5:23. https://doi.org/10.1186/1755-8794-5-23
Karstens KF, Bellon E, Polonski A, Wolters-Eisfeld G, Melling N, Reeh M, Izbicki JR, Tachezy M (2020) Expression and serum levels of the neural cell adhesion molecule L1-like protein (CHL1) in gastrointestinal stroma tumors (GIST) and its prognostic power. Oncotarget 11(13):1131–1140. https://doi.org/10.18632/oncotarget.27525
Beltran-Anaya FO, Romero-Cordoba S, Rebollar-Vega R, Arrieta O, Bautista-Pina V, Dominguez-Reyes C, Villegas-Carlos F, Tenorio-Torres A, Alfaro-Riuz L, Jimenez-Morales S, Cedro-Tanda A, Rios-Romero M, Reyes-Grajeda JP, Tagliabue E, Iorio MV, Hidalgo-Miranda A (2019) Expression of long non-coding RNA ENSG00000226738 (LncKLHDC7B) is enriched in the immunomodulatory triple-negative breast cancer subtype and its alteration promotes cell migration, invasion, and resistance to cell death. Mol Oncol 13(4):909–927. https://doi.org/10.1002/1878-0261.12446
Villalobo A, Berchtold MW (2020) The role of Calmodulin in tumor cell migration, invasiveness, and metastasis. Int J Mol Sci 21(3). https://doi.org/10.3390/ijms21030765
Haug BH, Hald OH, Utnes P, Roth SA, Lokke C, Flaegstad T, Einvik C (2015) Exosome-like extracellular vesicles from MYCN-amplified Neuroblastoma cells contain oncogenic miRNAs. Anticancer Res 35(5):2521–2530
Ma W, Chen X, Wu X, Li J, Mei C, Jing W, Teng L, Tu H, Jiang X, Wang G, Chen Y, Wang K, Wang H, Wei Y, Liu Z, Yuan Y (2020) Long noncoding RNA SPRY4-IT1 promotes proliferation and metastasis of hepatocellular carcinoma via mediating TNF signaling pathway. J Cell Physiol. https://doi.org/10.1002/jcp.29438
Li M, Ren CX, Zhang JM, Xin XY, Hua T, Wang HB, Wang HB (2018) The effects of miR-195-5p/MMP14 on proliferation and invasion of cervical carcinoma cells through TNF signaling pathway based on bioinformatics analysis of microarray profiling. Cell Physiol Biochem 50(4):1398–1413. https://doi.org/10.1159/000494602
Song Y, Kim JS, Choi EK, Kim J, Kim KM, Seo HR (2017) TGF-beta-independent CTGF induction regulates cell adhesion mediated drug resistance by increasing collagen I in HCC. Oncotarget 8(13):21650–21662. https://doi.org/10.18632/oncotarget.15521
Waddell JM, Evans J, Jabbour HN, Denison FC (2011) CTGF expression is up-regulated by PROK1 in early pregnancy and influences HTR-8/Svneo cell adhesion and network formation. Hum Reprod 26(1):67–75. https://doi.org/10.1093/humrep/deq294
Ball DK, Rachfal AW, Kemper SA, Brigstock DR (2003) The heparin-binding 10 kDa fragment of connective tissue growth factor (CTGF) containing module 4 alone stimulates cell adhesion. J Endocrinol 176(2):R1–R7. https://doi.org/10.1677/joe.0.176r001
Song ZM, Liu F, Chen YM, Liu YJ, Wang XD, Du SY (2019) CTGF-mediated ERK signaling pathway influences the inflammatory factors and intestinal flora in ulcerative colitis. Biomed Pharmacother 111:1429–1437. https://doi.org/10.1016/j.biopha.2018.12.063
Wang M, Liu Y, Zou J, Yang R, Xuan F, Wang Y, Gao N, Cui H (2015) Transcriptional co-activator TAZ sustains proliferation and tumorigenicity of neuroblastoma by targeting CTGF and PDGF-beta. Oncotarget 6(11):9517–9530. https://doi.org/10.18632/oncotarget.3367
Yuan W, Qian M, Li ZX, Zhao CL, Zhao J, Xiao JR (2019) Endothelin-1 activates the notch signaling pathway and promotes tumorigenesis in Giant cell tumor of the spine. Spine (Phila Pa 1976) 44(17):E1000–E1009. https://doi.org/10.1097/BRS.0000000000003044
Basurto L, Sanchez L, Diaz A, Valle M, Robledo A, Martinez-Murillo C (2019) Differences between metabolically healthy and unhealthy obesity in PAI-1 level: fibrinolysis, body size phenotypes and metabolism. Thromb Res 180:110–114. https://doi.org/10.1016/j.thromres.2019.06.013
Tang W, Dong K, Li K, Dong R, Zheng S (2016) MEG3, HCN3 and linc01105 influence the proliferation and apoptosis of neuroblastoma cells via the HIF-1alpha and p53 pathways. Sci Rep 6:36268. https://doi.org/10.1038/srep36268
Chen SJ, Hoffman NE, Shanmughapriya S, Bao L, Keefer K, Conrad K, Merali S, Takahashi Y, Abraham T, Hirschler-Laszkiewicz I, Wang J, Zhang XQ, Song J, Barrero C, Shi Y, Kawasawa YI, Bayerl M, Sun T, Barbour M, Wang HG, Madesh M, Cheung JY, Miller BA (2014) A splice variant of the human ion channel TRPM2 modulates neuroblastoma tumor growth through hypoxia-inducible factor (HIF)-1/2alpha. J Biol Chem 289(52):36284–36302. https://doi.org/10.1074/jbc.M114.620922
Wang Q, Xu Z, An Q, Jiang D, Wang L, Liang B, Li Z (2015) TAZ promotes epithelial to mesenchymal transition via the upregulation of connective tissue growth factor expression in neuroblastoma cells. Mol Med Rep 11(2):982–988. https://doi.org/10.3892/mmr.2014.2818
Mendes-de-Almeida DP, Andrade FG, Borges G, Dos Santos-Bueno FV, Vieira IF, da Rocha L, Mendes-da-Cruz DA, Zancope-Oliveira RM, Calado RT, Pombo-de-Oliveira MS (2019) GATA2 mutation in long stand Mycobacterium kansasii infection, myelodysplasia and MonoMAC syndrome: a case-report. BMC Med Genet 20(1):64. https://doi.org/10.1186/s12881-019-0799-6
Hoene V, Fischer M, Ivanova A, Wallach T, Berthold F, Dame C (2009) GATA factors in human neuroblastoma: distinctive expression patterns in clinical subtypes. Br J Cancer 101(8):1481–1489. https://doi.org/10.1038/sj.bjc.6605276
Wei JS, Johansson P, Chen L, Song YK, Tolman C, Li S, Hurd L, Patidar R, Wen X, Badgett TC, Cheuk AT, Marshall JC, Steeg PS, Vaque Diez JP, Yu Y, Gutkind JS, Khan J (2013) Massively parallel sequencing reveals an accumulation of de novo mutations and an activating mutation of LPAR1 in a patient with metastatic neuroblastoma. PLoS One 8(10):e77731. https://doi.org/10.1371/journal.pone.0077731
Li Q, Zhu CC, Ni B, Zhang ZZ, Jiang SH, Hu LP, Wang X, Zhang XX, Huang PQ, Yang Q, Li J, Gu JR, Xu J, Luo KQ, Zhao G, Zhang ZG (2019) Lysyl oxidase promotes liver metastasis of gastric cancer via facilitating the reciprocal interactions between tumor cells and cancer associated fibroblasts. EBioMedicine 49:157–171. https://doi.org/10.1016/j.ebiom.2019.10.037
Redova M, Chlapek P, Loja T, Zitterbart K, Hermanova M, Sterba J, Veselska R (2010) Influence of LOX/COX inhibitors on cell differentiation induced by all-trans retinoic acid in neuroblastoma cell lines. Int J Mol Med 25(2):271–280
Chlapek P, Redova M, Zitterbart K, Hermanova M, Sterba J, Veselska R (2010) Enhancement of ATRA-induced differentiation of neuroblastoma cells with LOX/COX inhibitors: an expression profiling study. J Exp Clin Cancer Res 29:45. https://doi.org/10.1186/1756-9966-29-45
Acknowledgements
We thank the staff from Medical Research Center of Shengjing Hospital who gave us support throughout the experiments.
Funding
The work was supported by the National Natural Science Foundation of China (No. 81972515, 81,472,359), Key Research and Development Foundation of Liaoning Province (2019JH8/10300024), 2013 Liaoning Climbing Scholar Foundation, and 345 Talent Project of Shengjing Hospital of China Medical University.
Author information
Authors and Affiliations
Contributions
Bo Chen and Peng Ding had full access to all of the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis. Both contributed equally to the study and are co-first authors; Zhongyan Hua and Xiuni Qin contributed to collection of data; Zhijie Li contributed to the study design, interpretation of the data, the writing of the manuscript, and the submission of the manuscript for publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
For this type of study, formal consent is not required.
Consent to participate
Not applicable.
Consent for publication
All authors consent to the publication of this study.
Code availability
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, B., Ding, P., Hua, Z. et al. Analysis and identification of novel biomarkers involved in neuroblastoma via integrated bioinformatics. Invest New Drugs 39, 52–65 (2021). https://doi.org/10.1007/s10637-020-00980-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10637-020-00980-9