Abstract
Purpose
Type 1 diabetes mellitus (T1DM) is a chronic autoimmune disease characterized by the destruction of pancreatic β cells. The goal of this study was to explore potential biological biomarkers for T1DM.
Methods
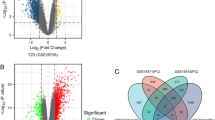
Two microarray datasets (GSE55098 and GSE156035) about human peripheral blood mononuclear cells (PBMCs) were systematically extracted from the Gene Expression Omnibus (GEO) database. Common genes were identified from the perspective of differentially expressed genes (DEGs) and weighted gene co-expression network analysis (WGCNA) respectively, and hub genes were identified by least absolute shrinkage and selection operator (LASSO) analysis. We also observed the expression of these hub genes in some common autoimmune diseases and predicted transcription factors (TFs) that might be associated with these genes.
Results
Seven hub genes (DDIT4, ESCO2, SH3BP4, PRICKLE1, EPM2AIP1, KCNJ15 and GRM8) were finally identified. Receiver operating characteristic (ROC) analysis showed that the high expression of these genes could well predict the occurrence of T1DM. Gene set enrichment analysis (GSEA) suggested that most of these hub genes may be mainly involved in the changes of biological functions such as inflammation, infection, immunity, cancer, and apoptosis. Further, compared with the control group, the expression levels of these hub genes also changed in some other autoimmune diseases, such as rheumatoid arthritis (RA), systemic lupus erythematosus (SLE), primary biliary cholangitis (PBC), etc., indicating that they might be the common targets of these autoimmune diseases.
Conclusions
The present study identified novel genes associated with T1DM from the PBMCs perspective that might provide new ideas for the early diagnosis, monitoring, evaluation, and prediction of T1DM.







Similar content being viewed by others
Data availability
Data was collected from the GEO database.
References
H. Sun, P. Saeedi, S. Karuranga, M. Pinkepank, K. Ogurtsova, B.B. Duncan, C. Stein, A. Basit, J.C.N. Chan, J.C. Mbanya, M.E. Pavkov, A. Ramachandaran, S.H. Wild, S. James, W.H. Herman, P. Zhang, C. Bommer, S. Kuo, E.J. Boyko, D.J. Magliano, (2022) IDF Diabetes Atlas: Global, regional and country-level diabetes prevalence estimates for 2021 and projections for 2045. Diabetes Res Clin Pract. 183(109119). https://doi.org/10.1016/j.diabres.2021.109119
A. Katsarou, S. Gudbjörnsdottir, A. Rawshani, D. Dabelea, E. Bonifacio, B.J. Anderson, L.M. Jacobsen, D.A. Schatz, Å Lernmark, Type 1 diabetes mellitus. Nat. Rev. Dis. Primers. 31, 7016 (2017). https://doi.org/10.1038/nrdp.2017.16
R. Barnett, Type 1 diabetes. Lancet 391(10117), 195 (2018). https://doi.org/10.1016/s0140-6736(18)30024-2
J.S. Skyler, Hope vs hype: where are we in type 1 diabetes. Diabetologia 61(3), 509–516 (2018). https://doi.org/10.1007/s00125-017-4530-x
L.A. DiMeglio, C. Evans-Molina, R.A. Oram, Type 1 diabetes. Lancet 391(10138), 2449–2462 (2018). https://doi.org/10.1016/s0140-6736(18)31320-5
X. Li, M. Liao, J. Guan, L. Zhou, R. Shen, M. Long, J. Shao, Identification of key genes and pathways in peripheral blood mononuclear cells of type 1 diabetes mellitus by integrated bioinformatics analysis. Diabetes Metab. J. (2022). https://doi.org/10.4093/dmj.2021.0018
P. Sen, E. Kemppainen, M. Orešič, Perspectives on systems modeling of human peripheral blood mononuclear cells. Front. Mol. Biosci. 4, 96 (2017). https://doi.org/10.3389/fmolb.2017.00096
R. Bomprezzi, M. Ringnér, S. Kim, M.L. Bittner, J. Khan, Y. Chen, A. Elkahloun, A. Yu, B. Bielekova, P.S. Meltzer, R. Martin, H.F. McFarland, J.M. Trent, Gene expression profile in multiple sclerosis patients and healthy controls: identifying pathways relevant to disease. Hum. Mol. Genet 12(17), 2191–2199 (2003). https://doi.org/10.1093/hmg/ddg221
S. Axelsson, M. Faresjö, M. Hedman, J. Ludvigsson, R. Casas, Cryopreserved peripheral blood mononuclear cells are suitable for the assessment of immunological markers in type 1 diabetic children. Cryobiology 57(3), 201–208 (2008). https://doi.org/10.1016/j.cryobiol.2008.08.001
E.C. Kaizer, C.L. Glaser, D. Chaussabel, J. Banchereau, V. Pascual, P.C. White, Gene expression in peripheral blood mononuclear cells from children with diabetes. J. Clin. Endocrinol. Metab. 92(9), 3705–3711 (2007). https://doi.org/10.1210/jc.2007-0979
S.C. Kent, Y. Chen, S.M. Clemmings, V. Viglietta, N.S. Kenyon, C. Ricordi, B. Hering, D.A. Hafler, Loss of IL-4 secretion from human type 1a diabetic pancreatic draining lymph node NKT cells. J. Immunol. 175(7), 4458–4464 (2005). https://doi.org/10.4049/jimmunol.175.7.4458
P.A. Ott, B.R. Berner, B.A. Herzog, R. Guerkov, N.L. Yonkers, I. Durinovic-Bello, M. Tary-Lehmann, P.V. Lehmann, D.D. Anthony, CD28 costimulation enhances the sensitivity of the ELISPOT assay for detection of antigen-specific memory effector CD4 and CD8 cell populations in human diseases. J. Immunol. Methods 285(2), 223–235 (2004). https://doi.org/10.1016/j.jim.2003.12.007
P. Sen, A.M. Dickens, M.A. López-Bascón, T. Lindeman, E. Kemppainen, S. Lamichhane, T. Rönkkö, J. Ilonen, J. Toppari, R. Veijola, H. Hyöty, T. Hyötyläinen, M. Knip, M. Orešič, Metabolic alterations in immune cells associate with progression to type 1 diabetes. Diabetologia 63(5), 1017–1031 (2020). https://doi.org/10.1007/s00125-020-05107-6
J.T. Leek, W.E. Johnson, H.S. Parker, A.E. Jaffe, J.D. Storey, The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28(6), 882–883 (2012). https://doi.org/10.1093/bioinformatics/bts034
M.E. Ritchie, B. Phipson, D. Wu, Y. Hu, C.W. Law, W. Shi, G.K. Smyth, limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7), e47 (2015). https://doi.org/10.1093/nar/gkv007
G. Yu, L.G. Wang, Y. Han, Q.Y. He, clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16(5), 284–287 (2012). https://doi.org/10.1089/omi.2011.0118
P. Langfelder, S. Horvath, WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008). https://doi.org/10.1186/1471-2105-9-559
D. Szklarczyk, A.L. Gable, K.C. Nastou, D. Lyon, R. Kirsch, S. Pyysalo, N.T. Doncheva, M. Legeay, T. Fang, P. Bork, L.J. Jensen, C. von Mering, The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 49(D1), D605–d612 (2021). https://doi.org/10.1093/nar/gkaa1074
P. Shannon, A. Markiel, O. Ozier, N.S. Baliga, J.T. Wang, D. Ramage, N. Amin, B. Schwikowski, T. Ideker, Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11), 2498–2504 (2003). https://doi.org/10.1101/gr.1239303
S. Engebretsen, J. Bohlin, Statistical predictions with glmnet. Clin. Epigenetics 11(1), 123 (2019). https://doi.org/10.1186/s13148-019-0730-1
A.J. McEligot, V. Poynor, R. Sharma, A. Panangadan, Logistic LASSO Regression for Dietary Intakes and Breast Cancer. Nutrients 12, 9 (2020). https://doi.org/10.3390/nu12092652
O. Fornes, J.A. Castro-Mondragon, A. Khan, R. van der Lee, X. Zhang, P.A. Richmond, B.P. Modi, S. Correard, M. Gheorghe, D. Baranašić, W. Santana-Garcia, G. Tan, J. Chèneby, B. Ballester, F. Parcy, A. Sandelin, B. Lenhard, W.W. Wasserman, A. Mathelier, JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acids Res 48(D1), D87–d92 (2020). https://doi.org/10.1093/nar/gkz1001
A. Subramanian, P. Tamayo, V.K. Mootha, S. Mukherjee, B.L. Ebert, M.A. Gillette, A. Paulovich, S.L. Pomeroy, T.R. Golub, E.S. Lander, J.P. Mesirov, Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102(43), 15545–15550 (2005). https://doi.org/10.1073/pnas.0506580102
A.S. Santos, E. Cunha-Neto, N.V. Gonfinetti, F.B. Bertonha, P. Brochet, A. Bergon, C.A. Moreira-Filho, C. Chevillard, M.E.R. da Silva, Prevalence of Inflammatory Pathways Over Immuno-Tolerance in Peripheral Blood Mononuclear Cells of Recent-Onset Type 1 Diabetes. Front Immunol 12, 765264 (2021). https://doi.org/10.3389/fimmu.2021.765264
G.S. Eisenbarth, Type 1 diabetes: molecular, cellular and clinical immunology. Adv. Exp. Med. Biol. 552, 306–10 (2004).
Y.J. Lan, N.B. Sam, M.H. Cheng, H.F. Pan, J. Gao, Progranulin as a Potential Therapeutic Target in Immune-Mediated Diseases. J. Inflamm. Res. 14, 6543–146556 (2021). https://doi.org/10.2147/jir.S339254
G. Geng, Z. Zhang, L. Cheng, Identification of a Multi-Long Noncoding RNA Signature for the Diagnosis of Type 1 Diabetes Mellitus. Front. Bioeng. Biotechnol. 8, 553 (2020). https://doi.org/10.3389/fbioe.2020.00553
Y. Gao, R. Zhang, S. Dai, X. Zhang, X. Li, C. Bai, Role of TGF-β/Smad pathway in the transcription of pancreas-specific genes during Beta cell differentiation. Front. Cell Dev. Biol. 7, 351 (2019). https://doi.org/10.3389/fcell.2019.00351
W.P. Miller, C. Yang, M.L. Mihailescu, J.A. Moore, W. Dai, A.J. Barber, M.D. Dennis, Deletion of the Akt/mTORC1 repressor REDD1 prevents visual dysfunction in a rodent model of type 1 diabetes. Diabetes 67(1), 110–119 (2018). https://doi.org/10.2337/db17-0728
G. Wang, Y. Yan, Z. Zheng, T., Zhang The mechanism of hsa-miR-424-5 combining PD-1 through mTORC signaling pathway to stimulate immune effect and participate in Type 1 diabetes. Biosci. Rep. 40, 3 (2020). https://doi.org/10.1042/bsr20193800
Y.M. Kim, D.H. Kim, dRAGging amino acid-mTORC1 signaling by SH3BP4. Mol. Cells 35(1), 1–6 (2013). https://doi.org/10.1007/s10059-013-2249-1
B. Xu, M. Gogol, K. Gaudenz, J.L. Gerton, Improved transcription and translation with L-leucine stimulation of mTORC1 in Roberts syndrome. BMC Genomics 17, 25 (2016). https://doi.org/10.1186/s12864-015-2354-y
A.M. Daulat, F. Bertucci, S. Audebert, A. Sergé, P. Finetti, E. Josselin, R. Castellano, D. Birnbaum, S. Angers, J.P. Borg, PRICKLE1 contributes to cancer cell dissemination through its interaction with mTORC2. Dev. Cell 37(4), 311–325 (2016). https://doi.org/10.1016/j.devcel.2016.04.011
M. Blandino-Rosano, R. Barbaresso, M. Jimenez-Palomares, M. Bozadjieva N., J.P. Werneck-de-Castro, M. Hatanaka, R.G. Mirmira, N. Sonenberg, M. Liu, M.A. Rüegg, M.N. Hall, E. Bernal-Mizrachi, Loss of mTORC1 signalling impairs β-cell homeostasis and insulin processing. Nat. Commun. 8, 16014 (2017). https://doi.org/10.1038/ncomms16014
T. Suto, T. Karonitsch, The immunobiology of mTOR in autoimmunity. J Autoimmun 110, 102373 (2020). https://doi.org/10.1016/j.jaut.2019.102373
C.P. Martins, L.A. New, E.C. O’Connor, D.M. Previte, K.R. Cargill, I.L. Tse, S. Sims-Lucas, J.D. Piganelli, Glycolys is inhibition induces functional and metabolic exhaustion of CD4(+) T cells in Type 1 Diabetes. Front. Immunol. 12, 669456 (2021). https://doi.org/10.3389/fimmu.2021.669456
X. Bi, F. Li, S. Liu, Y. Jin, X. Zhang, T. Yang, Y. Dai, X. Li, A.Z. Zhao, ω-3 polyunsaturated fatty acids ameliorate type 1 diabetes and autoimmunity. J. Clin. Invest 127(5), 1757–1771 (2017). https://doi.org/10.1172/jci87388
K.M. Irvine, P. Gallego, X. An, S.E. Best, G. Thomas, C. Wells, M. Harris, A. Cotterill, R. Thomas, Peripheral blood monocyte gene expression profile clinically stratifies patients with recent-onset type 1 diabetes. Diabetes 61(5), 1281–1290 (2012). https://doi.org/10.2337/db11-1549
J. Turnbull, E. Tiberia, S. Pereira, X. Zhao, N. Pencea, A.L. Wheeler, W.Q. Yu, A. Ivovic, T. Naranian, N. Israelian, A. Draginov, M. Piliguian, P.W. Frankland, P. Wang, C.A. Ackerley, A. Giacca, B.A. Minassian, Deficiency of a glycogen synthase-associated protein, Epm2aip1, causes decreased glycogen synthesis and hepatic insulin resistance. J. Biol. Chem. 288(48), 34627–34637 (2013). https://doi.org/10.1074/jbc.M113.483198
K. Okamoto, N. Iwasaki, C. Nishimura, K. Doi, E. Noiri, S. Nakamura, M. Takizawa, M. Ogata, R. Fujimaki, N. Grarup, C. Pisinger, K. Borch-Johnsen, T. Lauritzen, A. Sandbaek, T. Hansen, K. Yasuda, H. Osawa, K. Nanjo, T. Kadowaki, M. Kasuga, O. Pedersen, T. Fujita, N. Kamatani, Y. Iwamoto, K. Tokunaga, Identification of KCNJ15 as a susceptibility gene in Asian patients with type 2 diabetes mellitus. Am. J. Hum. Genet 86(1), 54–64 (2010). https://doi.org/10.1016/j.ajhg.2009.12.009
K. Okamoto, N. Iwasaki, K. Doi, E. Noiri, Y. Iwamoto, Y. Uchigata, T. Fujita, K. Tokunaga, Inhibition of glucose-stimulated insulin secretion by KCNJ15, a newly identified susceptibility gene for type 2 diabetes. Diabetes 61(7), 1734–1741 (2012). https://doi.org/10.2337/db11-1201
Z. Hawi, T.D. Cummins, J. Tong, B. Johnson, R. Lau, W. Samarrai, M.A. Bellgrove, The molecular genetic architecture of attention deficit hyperactivity disorder. Mol. Psychiatry 20(3), 289–297 (2015). https://doi.org/10.1038/mp.2014.183
Acknowledgements
We want to acknowledge all participants of this study and the technical support provided by the Affiliated Hospital of Jiangsu University.
Funding
This study was funded by the National Natural Science Foundation of China (81870548 and 81570721), the Social Development Project of Jiangsu Province (BE2018692), the Natural Science Foundation of Jiangsu Province, China (BK20191222), the Key project for Medical Education Collaborative Innovation Fund of Jiangsu University (JDY2022005).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Z., Zhang, L., Tang, F. et al. Transcriptome analysis of peripheral blood mononuclear cells in patients with type 1 diabetes mellitus. Endocrine 78, 270–279 (2022). https://doi.org/10.1007/s12020-022-03163-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12020-022-03163-z