Abstract
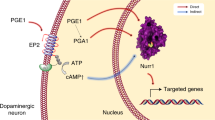
The orphan nuclear receptor Nurr1 is critical for the development, maintenance, and protection of midbrain dopaminergic neurons. Recently, we demonstrated that prostaglandins E1 (PGE1) and PGA1 directly bind to the ligand-binding domain (LBD) of Nurr1 and stimulate its transcriptional activation function. In this direction, here we report the transcriptional activation of Nurr1 by PGA2, a dehydrated metabolite of PGE2, through physical binding ably supported by NMR titration and crystal structure. The co-crystal structure of Nurr1-LBD bound to PGA2 revealed the covalent coupling of PGA2 with Nurr1-LBD through Cys566. PGA2 binding also induces a 21° shift of the activation function 2 (AF-2) helix H12 away from the protein core, similar to that observed in the Nurr1-LBD-PGA1 complex. We also show that PGA2 can rescue the locomotor deficits and neuronal degeneration in LRRK2 G2019S transgenic fly models.




Similar content being viewed by others
Data Availability
Coordinates and structure factors for Nurr1-LBD – PGA2 complex have been deposited in the Protein Data Bank with accession number 5YD6.
References
Aarnisalo, P., Kim, C. H., Lee, J. W., & Perlmann, T. (2002). Defining requirements for heterodimerization between the retinoid X receptor and the orphan nuclear receptor Nurr1. Journal of Biological Chemistry, 277(38), 35118–35123. https://doi.org/10.1074/jbc.M201707200
Baker, K. D., Shewchuk, L. M., Kozlova, T., Makishima, M., Hassell, A., Wisely, B., et al. (2003). The Drosophila orphan nuclear receptor DHR38 mediates an atypical ecdysteroid signaling pathway. Cell, 113(6), 731–742. https://doi.org/10.1016/s0092-8674(03)00420-3
Battye, T. G., Kontogiannis, L., Johnson, O., Powell, H. R., & Leslie, A. G. (2011). iMOSFLM: A new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallographica Section D Biological Crystallography, 67(Pt 4), 271–281. https://doi.org/10.1107/s0907444910048675
Bruning, J. M., Wang, Y., Oltrabella, F., Tian, B., Kholodar, S. A., Liu, H., et al. (2019). Covalent Modification and Regulation of the Nuclear Receptor Nurr1 by a Dopamine Metabolite. Cell Chemical Biology, 26(5), 674-685.e676. https://doi.org/10.1016/j.chembiol.2019.02.002
Chu, Y., Le, W., Kompoliti, K., Jankovic, J., Mufson, E. J., & Kordower, J. H. (2006). Nurr1 in Parkinson’s disease and related disorders. Journal of Comparative Neurology, 494(3), 495–514. https://doi.org/10.1002/cne.20828
Codina, A., Benoit, G., Gooch, J. T., Neuhaus, D., Perlmann, T., & Schwabe, J. W. (2004). Identification of a novel co-regulator interaction surface on the ligand binding domain of Nurr1 using NMR footprinting. Journal of Biological Chemistry, 279(51), 53338–53345. https://doi.org/10.1074/jbc.M409096200
de Vera, I. M., Giri, P. K., Munoz-Tello, P., Brust, R., Fuhrmann, J., Matta-Camacho, E., et al. (2016). Identification of a binding site for unsaturated fatty acids in the orphan nuclear receptor Nurr1. ACS Chemical Biology, 11(7), 1795–1799. https://doi.org/10.1021/acschembio.6b00037
Decressac, M., Volakakis, N., Bjorklund, A., & Perlmann, T. (2013). NURR1 in Parkinson disease–from pathogenesis to therapeutic potential. Nature Review Neurology, 9(11), 629–636. https://doi.org/10.1038/nrneurol.2013.209
Emsley, P., & Cowtan, K. (2004). Coot: Model-building tools for molecular graphics. Acta Crystallographica Section D Biological Crystallography, 60(Pt 12 Pt 1), 2126–2132. https://doi.org/10.1107/s0907444904019158
Evans, P. (2006). Scaling and assessment of data quality. Acta Crystallographica Section D Biological Crystallography, 62(Pt 1), 72–82. https://doi.org/10.1107/S0907444905036693
Flaig, R., Greschik, H., Peluso-Iltis, C., & Moras, D. (2005). Structural basis for the cell-specific activities of the NGFI-B and the Nurr1 ligand-binding domain. Journal of Biological Chemistry, 280(19), 19250–19258. https://doi.org/10.1074/jbc.M413175200
Freund, C., Schmalz, H. G., Sticht, J., & Kuhne, R. (2008). Proline-rich sequence recognition domains (PRD): Ligands, function and inhibition. Handbook of experimental pharmacology (Vol. 186, pp. 407–429). Springer. https://doi.org/10.1007/978-3-540-72843-6_17
Kagaya, S., Ohkura, N., Tsukada, T., Miyagawa, M., Sugita, Y., Tsujimoto, G., et al. (2005). Prostaglandin A2 acts as a transactivator for NOR1 (NR4A3) within the nuclear receptor superfamily. Biological and Pharmaceutical Bulletin, 28(9), 1603–1607. https://doi.org/10.1248/bpb.28.1603
Kim, K. S., Kim, C. H., Hwang, D. Y., Seo, H., Chung, S., Hong, S. J., et al. (2003). Orphan nuclear receptor Nurr1 directly transactivates the promoter activity of the tyrosine hydroxylase gene in a cell-specific manner. Journal of Neurochemistry, 85(3), 622–634. https://doi.org/10.1046/j.1471-4159.2003.01671.x
Kim, C.-H., Han, B.-S., Moon, J., Kim, D.-J., Shin, J., Rajan, S., et al. (2015). Nuclear receptor Nurr1 agonists enhance its dual functions and improve behavioral deficits in an animal model of Parkinson’s disease. Proceedings of the National Academy of Sciences USA, 112(28), 8756–8761. https://doi.org/10.1073/pnas.1509742112
Lakshmi, S. P., Reddy, A. T., Banno, A., & Reddy, R. C. (2019). Molecular, chemical, and structural characterization of prostaglandin A2 as a novel agonist for Nur77. Biochemical Journal, 476(19), 2757–2767. https://doi.org/10.1042/BCJ20190253
Lee, W., Tonelli, M., & Markley, J. L. (2015). NMRFAM-SPARKY: Enhanced software for biomolecular NMR spectroscopy. Bioinformatics, 31(8), 1325–1327. https://doi.org/10.1093/bioinformatics/btu830
Li, L., Liu, Y., Chen, H. Z., Li, F. W., Wu, J. F., Zhang, H. K., et al. (2015). Impeding the interaction between Nur77 and p38 reduces LPS-induced inflammation. Nature Chemical Biology, 11(5), 339–346. https://doi.org/10.1038/nchembio.1788
Liu, Z., Wang, X., Yu, Y., Li, X., Wang, T., Jiang, H., et al. (2008). A Drosophila model for LRRK2-linked parkinsonism. Proceedings of the National Academy of Sciences USA, 105(7), 2693–2698. https://doi.org/10.1073/pnas.0708452105
Madeira, F., Park, Y. M., Lee, J., Buso, N., Gur, T., Madhusoodanan, N., et al. (2019). The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Research, 47(W1), W636–W641. https://doi.org/10.1093/nar/gkz268
McCoy, A. J., Grosse-Kunstleve, R. W., Adams, P. D., Winn, M. D., Storoni, L. C., & Read, R. J. (2007). Phaser crystallographic software. Journal of Applied Crystallography, 40(Pt 4), 658–674. https://doi.org/10.1107/s0021889807021206
Murshudov, G. N., Skubak, P., Lebedev, A. A., Pannu, N. S., Steiner, R. A., Nicholls, R. A., et al. (2011). REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallographica Section D Biological Crystallography, 67(Pt 4), 355–367. https://doi.org/10.1107/S0907444911001314
Ng, C. H., Mok, S. Z., Koh, C., Ouyang, X., Fivaz, M. L., Tan, E.-K., et al. (2009). Parkin protects against LRRK2 G2019S mutant-induced dopaminergic neurodegeneration in Drosophila. Journal of Neuroscience, 29(36), 11257–11262. https://doi.org/10.1523/JNEUROSCI.2375-09.2009
Ng, C. H., Guan, M. S., Koh, C., Ouyang, X., Yu, F., Tan, E. K., et al. (2012). AMP kinase activation mitigates dopaminergic dysfunction and mitochondrial abnormalities in Drosophila models of Parkinson’s disease. Journal of Neuroscience, 32(41), 14311–14317. https://doi.org/10.1523/JNEUROSCI.0499-12.2012
Rajan, S., Jang, Y., Kim, C. H., Kim, W., Toh, H. T., Jeon, J., et al. (2020). PGE1 and PGA1 bind to Nurr1 and activate its transcriptional function. Nature Chemical Biology, 16(8), 876–886. https://doi.org/10.1038/s41589-020-0553-6
Rastinejad, F., Huang, P., Chandra, V., & Khorasanizadeh, S. (2013). Understanding nuclear receptor form and function using structural biology. Journal of Molecular Endocrinology, 51(3), T1–T21. https://doi.org/10.1530/JME-13-0173
Robert, X., & Gouet, P. (2014). Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Research, 42(W1), W320-324. https://doi.org/10.1093/nar/gku316
Vellieux, F. M. D., & Dijkstra, B. W. (1997). Computation of Bhat’s OMIT maps with different coefficients. Journal of Applied Crystallography, 30(3), 396–399. https://doi.org/10.1107/S0021889896012551
Waku, T., Shiraki, T., Oyama, T., Fujimoto, Y., Maebara, K., Kamiya, N., et al. (2009). Structural insight into PPARgamma activation through covalent modification with endogenous fatty acids. Journal of Molecular Biology, 385(1), 188–199. https://doi.org/10.1016/j.jmb.2008.10.039
Wang, Z., Benoit, G., Liu, J., Prasad, S., Aarnisalo, P., Liu, X., et al. (2003). Structure and function of Nurr1 identifies a class of ligand-independent nuclear receptors. Nature, 423(6939), 555. https://doi.org/10.1038/nature01645
Wang, C., Lu, R., Ouyang, X., Ho, M. W. L., Chia, W., Yu, F., et al. (2007). Drosophila overexpressing parkin R275W mutant exhibits dopaminergic neuron degeneration and mitochondrial abnormalities. The Journal of Neuroscience: The Official Journal of the Society for Neuroscience, 27(32), 8563–8570. https://doi.org/10.1523/JNEUROSCI.0218-07.2007
Wang, W. J., Wang, Y., Chen, H. Z., Xing, Y. Z., Li, F. W., Zhang, Q., et al. (2014). Orphan nuclear receptor TR3 acts in autophagic cell death via mitochondrial signaling pathway. Nature Chemical Biology, 10(2), 133–140. https://doi.org/10.1038/nchembio.1406
Wang, W. J., Wang, Y., Hou, P. P., Li, F. W., Zhou, B., Chen, H. Z., et al. (2015). Induction of autophagic death in cancer cells by agonizing TR3 and attenuating Akt2 activity. Chemical Biology, 22(8), 1040–1051. https://doi.org/10.1016/j.chembiol.2015.06.023
Weikum, E. R., Liu, X., & Ortlund, E. A. (2018). The nuclear receptor superfamily: A structural perspective. Protein Science, 27(11), 1876–1892. https://doi.org/10.1002/pro.3496
Whitworth, A. J., Theodore, D. A., Greene, J. C., Beneš, H., Wes, P. D., & Pallanck, L. J. (2005). Increased glutathione S-transferase activity rescues dopaminergic neuron loss in a Drosophila model of Parkinson’s disease. Proceedings of the National Academy of Sciences, 102(22), 8024–8029. https://doi.org/10.1073/pnas.0501078102
Winn, M. D., Ballard, C. C., Cowtan, K. D., Dodson, E. J., Emsley, P., Evans, P. R., et al. (2011). Overview of the CCP4 suite and current developments. Acta Crystallographica Section D Biological Crystallography, 67(Pt 4), 235–242. https://doi.org/10.1107/s0907444910045749
Zhan, Y. Y., Chen, Y., Zhang, Q., Zhuang, J. J., Tian, M., Chen, H. Z., et al. (2012). The orphan nuclear receptor Nur77 regulates LKB1 localization and activates AMPK. Nature Chemical Biology, 8(11), 897–904. https://doi.org/10.1038/nchembio.1069
Acknowledgements
The authors thank the National Synchrotron Radiation Research Center (NSRRC) and their staff at beamline TLS 13B1 for help with data collection. The NSRRC is a national user facility supported by the National Science Council of Taiwan, ROC; the National Research Program Genomic Medicine supports the Synchrotron Radiation Protein Crystallography Facility at NSRRC. We thank Dr. Aida Serra for help with mass spectrometry data analysis.
Funding
This work was supported by the Ministry of Education Singapore AcRF Tier 2 Grant (MOE2016-T2-2-055) (YHS) and Ministry of Health NMRC-LCG Singapore Parkinson’s disease Translational Clinical Programme (MOH-000207) (LKL).
Author information
Authors and Affiliations
Contributions
HSY conceived the study. SR, HTT, HY, ZW, AHB, TP, JYY performed the research and analyzed the data. SR, HTT, KLL, and HSY wrote the paper and all the authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rajan, S., Toh, H.T., Ye, H. et al. Prostaglandin A2 Interacts with Nurr1 and Ameliorates Behavioral Deficits in Parkinson’s Disease Fly Model. Neuromol Med 24, 469–478 (2022). https://doi.org/10.1007/s12017-022-08712-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12017-022-08712-3